Abstract
Genome-wide association studies identify genomic variants associated with human traits and diseases. Most trait-associated variants are located within cell-type-specific enhancers, but the molecular mechanisms governing phenotypic variation are less well understood. Here, we show that many enhancer variants associated with red blood cell (RBC) traits map to enhancers that are co-bound by lineage-specific master transcription factors (MTFs) and signaling transcription factors (STFs) responsive to extracellular signals. The majority of enhancer variants reside on STF and not MTF motifs, perturbing DNA binding by various STFs (BMP/TGF-β-directed SMADs or WNT-induced TCFs) and affecting target gene expression. Analyses of engineered human blood cells and expression quantitative trait loci verify that disrupted STF binding leads to altered gene expression. Our results propose that the majority of the RBC-trait-associated variants that reside on transcription-factor-binding sequences fall in STF target sequences, suggesting that the phenotypic variation of RBC traits could stem from altered responsiveness to extracellular stimuli.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The massively parallel sequencing data associated with this manuscript have been uploaded to GEO under the accession numbers GSE74483 and GSE104574 and are currently open to the public. The web links for the publicly available databases used in this study are: UniPROBE, http://thebrain.bwh.harvard.edu/uniprobe/; CIS-BP, http://cisbp.ccbr.utoronto.ca/; FHS, https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000007.v30.p11; RegulomeDB, https://regulomedb.org/regulome-search/; HEMMER, http://hmmer.org/; EMBOSS Needle, https://www.ebi.ac.uk/Tools/psa/emboss_needle/; dbSNP, https://www.ncbi.nlm.nih.gov/snp/?cmd=search. Links to all of the PBM datasets used are available in Supplementary Table 7. Source data are provided with this paper.
Code availability
The custom codes used in this study are available at https://bitbucket.org/abrahamb/workspace/projects/TSC. The code and data files for the PBM analyses are available at https://github.com/BulykLab/RBCSNPs_2020.
References
Evans, D. M., Frazer, I. H. & Martin, N. G. Genetic and environmental causes of variation in basal levels of blood cells. Twin Res. 2, 250–257 (1999).
Guindo, A., Fairhurst, R. M., Doumbo, O. K., Wellems, T. E. & Diallo, D. A. X-linked G6PD deficiency protects hemizygous males but not heterozygous females against severe malaria. PLoS Med. 4, e66 (2007).
Lin, J. P. et al. Evidence for linkage of red blood cell size and count: genome-wide scans in the Framingham Heart Study. Am. J. Hematol. 82, 605–610 (2007).
Lo, K. S. et al. Genetic association analysis highlights new loci that modulate hematological trait variation in Caucasians and African Americans. Hum. Genet. 129, 307–317 (2011).
Tishkoff, S. A. et al. Haplotype diversity and linkage disequilibrium at human G6PD: recent origin of alleles that confer malarial resistance. Science 293, 455–462 (2001).
Whitfield, J. B. & Martin, N. G. Genetic and environmental influences on the size and number of cells in the blood. Genet. Epidemiol. 2, 133–144 (1985).
Koury, M. J. Abnormal erythropoiesis and the pathophysiology of chronic anemia. Blood Rev. 28, 49–66 (2014).
Edwards, S. L., Beesley, J., French, J. D. & Dunning, A. M. Beyond GWASs: illuminating the dark road from association to function. Am. J. Hum. Genet. 93, 779–797 (2013).
Guo, M. H. et al. Comprehensive population-based genome sequencing provides insight into hematopoietic regulatory mechanisms. Proc. Natl Acad. Sci. USA 114, E327–E336 (2017).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Melnikov, A. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nat. Biotechnol. 30, 271–277 (2012).
Nandakumar, S. K., Ulirsch, J. C. & Sankaran, V. G. Advances in understanding erythropoiesis: evolving perspectives. Br. J. Haematol. 173, 206–218 (2016).
Patwardhan, R. P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nat. Biotechnol. 30, 265–270 (2012).
Polfus, L. M. et al. Whole-exome sequencing identifies loci associated with blood cell traits and reveals a role for alternative GFI1B splice variants in human hematopoiesis. Am. J. Hum. Genet. 99, 785 (2016).
Ulirsch, J. C. et al. Systematic functional dissection of common genetic variation affecting red blood cell traits. Cell 165, 1530–1545 (2016).
van der Harst, P. et al. Seventy-five genetic loci influencing the human red blood cell. Nature 492, 369–375 (2012).
Ganesh, S. K. et al. Multiple loci influence erythrocyte phenotypes in the CHARGE Consortium. Nat. Genet. 41, 1191–1198 (2009).
van Rooij, F. J. et al. Genome-wide trans-ethnic meta-analysis identifies seven genetic loci influencing erythrocyte traits and a role for RBPMS in erythropoiesis. Am. J. Hum. Genet. 100, 51–63 (2017).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429 (2016).
Chami, N. et al. Exome genotyping identifies pleiotropic variants associated with red blood cell traits. Am. J. Hum. Genet. 99, 8–21 (2016).
Pankratz, N. et al. Genomewide association study for susceptibility genes contributing to familial Parkinson disease. Hum. Genet. 124, 593–605 (2009).
Levo, M. et al. Unraveling determinants of transcription factor binding outside the core binding site. Genome Res. 25, 1018–1029 (2015).
Ulirsch, J. C. et al. Interrogation of human hematopoiesis at single-cell and single-variant resolution. Nat. Genet. 51, 683–693 (2019).
Dent, P. et al. Stress and radiation-induced activation of multiple intracellular signaling pathways. Radiat. Res. 159, 283–300 (2003).
Gaki, G. S. & Papavassiliou, A. G. Oxidative stress-induced signaling pathways implicated in the pathogenesis of Parkinson’s disease. Neuromolecular Med. 16, 217–230 (2014).
Uchida, K. et al. Activation of stress signaling pathways by the end product of lipid peroxidation. 4-hydroxy-2-nonenal is a potential inducer of intracellular peroxide production. J. Biol. Chem. 274, 2234–2242 (1999).
Mullen, A. C. et al. Master transcription factors determine cell-type-specific responses to TGF-β signaling. Cell 147, 565–576 (2011).
Trompouki, E. et al. Lineage regulators direct BMP and Wnt pathways to cell-specific programs during differentiation and regeneration. Cell 147, 577–589 (2011).
Sankaran, V. G. et al. Human fetal hemoglobin expression is regulated by the developmental stage-specific repressor BCL11A. Science 322, 1839–1842 (2008).
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Lenox, L. E., Perry, J. M. & Paulson, R. F. BMP4 and Madh5 regulate the erythroid response to acute anemia. Blood 105, 2741–2748 (2005).
Lenox, L. E., Shi, L., Hegde, S. & Paulson, R. F. Extramedullary erythropoiesis in the adult liver requires BMP-4/Smad5-dependent signaling. Exp. Hematol. 37, 549–558 (2009).
McReynolds, L. J., Tucker, J., Mullins, M. C. & Evans, T. Regulation of hematopoiesis by the BMP signaling pathway in adult zebrafish. Exp. Hematol. 36, 1604–1615 (2008).
Porayette, P. & Paulson, R. F. BMP4/Smad5 dependent stress erythropoiesis is required for the expansion of erythroid progenitors during fetal development. Dev. Biol. 317, 24–35 (2008).
Hnisz, D. et al. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol. Cell 58, 362–370 (2015).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Fisher, R. C. & Scott, E. W. Role of PU.1 in hematopoiesis. Stem Cells 16, 25–37 (1998).
Li, Y., Luo, H., Liu, T., Zacksenhaus, E. & Ben-David, Y. The ets transcription factor Fli-1 in development, cancer and disease. Oncogene 34, 2022–2031 (2015).
Shivdasani, R. A. & Orkin, S. H. Erythropoiesis and globin gene expression in mice lacking the transcription factor NF-E2. Proc. Natl Acad. Sci. USA 92, 8690–8694 (1995).
Siatecka, M. & Bieker, J. J. The multifunctional role of EKLF/KLF1 during erythropoiesis. Blood 118, 2044–2054 (2011).
Nakao, A. et al. TGF-β receptor-mediated signalling through Smad2, Smad3 and Smad4. EMBO J. 16, 5353–5362 (1997).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Kurita, R. et al. Establishment of immortalized human erythroid progenitor cell lines able to produce enucleated red blood cells. PLoS ONE 8, e59890 (2013).
Zhang, F. & Lupski, J. R. Non-coding genetic variants in human disease. Hum. Mol. Genet. 24, R102–R110 (2015).
Visscher, P. M. et al. 10 years of GWAS discovery: biology, function, and translation. Am. J. Hum. Genet. 101, 5–22 (2017).
Cohen, A. J. et al. Hotspots of aberrant enhancer activity punctuate the colorectal cancer epigenome. Nat. Commun. 8, 14400 (2017).
Corradin, O. et al. Combinatorial effects of multiple enhancer variants in linkage disequilibrium dictate levels of gene expression to confer susceptibility to common traits. Genome Res. 24, 1–13 (2014).
Morrow, J. J. et al. Positively selected enhancer elements endow osteosarcoma cells with metastatic competence. Nat. Med. 24, 176–185 (2018).
Scacheri, C. A. & Scacheri, P. C. Mutations in the noncoding genome. Curr. Opin. Pediatr. 27, 659–664 (2015).
The CHARGE Consortium Hematology Working Group. Meta-analysis of rare and common exome chip variants identifies S1PR4 and other loci influencing blood cell traits. Nat. Genet. 48, 867–876 (2016).
Chen, Z. et al. Genome-wide association analysis of red blood cell traits in African Americans: the COGENT Network. Hum. Mol. Genet. 22, 2529–2538 (2013).
Li, C. et al. Genome-wide association study meta-analysis of long-term average blood pressure in East Asians. Circ. Cardiovasc. Genet. 10, e001527 (2017).
Paul, D. S. et al. Maps of open chromatin highlight cell type-restricted patterns of regulatory sequence variation at hematological trait loci. Genome Res. 23, 1130–1141 (2013).
Paul, D. S. et al. Maps of open chromatin guide the functional follow-up of genome-wide association signals: application to hematological traits. PLoS Genet. 7, e1002139 (2011).
Amos, C. I. et al. The oncoarray consortium: a network for understanding the genetic architecture of common cancers. Cancer Epidemiol. Biomark. Prev. 26, 126–135 (2017).
Fachal, L. et al. Fine-mapping of 150 breast cancer risk regions identifies 191 likely target genes. Nat. Genet. 52, 56–73 (2020).
Fritsche, L. G. et al. Association of polygenic risk scores for multiple cancers in a phenome-wide study: results from the Michigan Genomics Initiative. Am. J. Hum. Genet. 102, 1048–1061 (2018).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Lin, J. R. et al. Integrated post-GWAS analysis sheds new light on the disease mechanisms of schizophrenia. Genetics 204, 1587–1600 (2016).
Vicente, C. T. et al. Long-range modulation of PAG1 expression by 8q21 allergy risk variants. Am. J. Hum. Genet. 97, 329–336 (2015).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Liu, N. et al. Direct promoter repression by BCL11A controls the fetal to adult hemoglobin switch. Cell 173, 430–442 (2018).
Hume, M. A., Barrera, L. A., Gisselbrecht, S. S. & Bulyk, M. L. UniPROBE, update 2015: new tools and content for the online database of protein-binding microarray data on protein–DNA interactions. Nucleic Acids Res. 43, D117–D122 (2015).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Barrera, L. A. et al. Survey of variation in human transcription factors reveals prevalent DNA binding changes. Science 351, 1450–1454 (2016).
Mariani, L., Weinand, K., Vedenko, A., Barrera, L. A. & Bulyk, M. L. Identification of human lineage-specific transcriptional coregulators enabled by a glossary of binding modules and tunable genomic backgrounds. Cell Syst. 5, 187–201 (2017).
Peterson, K. A. et al. Neural-specific Sox2 input and differential Gli-binding affinity provide context and positional information in Shh-directed neural patterning. Genes Dev. 26, 2802–2816 (2012).
Joehanes, R. et al. Integrated genome-wide analysis of expression quantitative trait loci aids interpretation of genomic association studies. Genome Biol. 18, 16 (2017).
Tran, F. H. & Zheng, J. J. Modulating the wnt signaling pathway with small molecules. Protein Sci. 26, 650–661 (2017).
Caron, B., Luo, Y. & Rausell, A. NCBoost classifies pathogenic non-coding variants in Mendelian diseases through supervised learning on purifying selection signals in humans. Genome Biol. 20, 32 (2019).
Sankaran, V. G., Orkin, S. H. & Walkley, C. R. Rb intrinsically promotes erythropoiesis by coupling cell cycle exit with mitochondrial biogenesis. Genes Dev. 22, 463–475 (2008).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Vinjamur, D. S. & Bauer, D. E. Growing and genetically manipulating human umbilical cord blood-derived erythroid progenitor (HUDEP) cell lines. Methods Mol. Biol. 1698, 275–284 (2018).
Canver, M. C. et al. Integrated design, execution, and analysis of arrayed and pooled CRISPR genome-editing experiments. Nat. Protoc. 13, 946–986 (2018).
Gregory, T. et al. GATA-1 and erythropoietin cooperate to promote erythroid cell survival by regulating bcl-xL expression. Blood 94, 87–96 (1999).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Chelala, C., Khan, A. & Lemoine, N. R. SNPnexus: a web database for functional annotation of newly discovered and public domain single nucleotide polymorphisms. Bioinformatics 25, 655–661 (2009).
Dayem Ullah, A. Z., Lemoine, N. R. & Chelala, C. SNPnexus: a web server for functional annotation of novel and publicly known genetic variants (2012 update). Nucleic Acids Res. 40, W65–W70 (2012).
Dayem Ullah, A. Z., Lemoine, N. R. & Chelala, C. A practical guide for the functional annotation of genetic variations using SNPnexus. Brief. Bioinform. 14, 437–447 (2013).
Leslie, R., O’Donnell, C. J. & Johnson, A. D. GRASP: analysis of genotype-phenotype results from 1390 genome-wide association studies and corresponding open access database. Bioinformatics 30, i185–i194 (2014).
Splansky, G. L. et al. The third generation cohort of the National Heart, Lung, and Blood Institute’s Framingham Heart Study: design, recruitment, and initial examination. Am. J. Epidemiol. 165, 1328–1335 (2007).
Acknowledgements
We are grateful to S. Orkin, A. Wagers and C. Santoriello for the critical reading and editing of our manuscript. We thank the HHMI high-throughput sequencing facility at the Children’s Hospital Boston for generating the genome-wide raw sequencing data. We acknowledge V. G. Sankaran and S. Nandakumar for providing the luciferase plasmids containing non-coding alleles of the RBM38 gene and the enhancer mutant for the RBM38 gene in K562 cells. This work was supported by the following grants to L.I.Z.—R01 HL04880, P015PO1HL32262-32, 5P30 DK49216, 5R01 DK53298, 5U01 HL10001-05, R24 DK092760, 1R24OD017870-01. Additional support came from the funding by the Max Planck Society, The Fritz Thyssen Stiftung (Az 10.17.1.026MN), a Marie Curie Career Integration Grant (631432 Bloody Signals), the Deutsche Forschungsgemeinschaft DFG under Germany’s Excellence Strategy (CIBSS-EXC-2189-Project-ID-390939984) and the Deutsche Forschungsgemeinschaft, Research Training Group 322977937/GRK2344 ‘MeInBio –BioInMe’ to E.T., the Hope Funds for Cancer Research Grillo-Marxuach Family Fellowship and the American Lebanese Syrian Associated Charities to B.J.A. R.A.Y. is supported by NIH grants GM123511, CA213333 and CA155258. K.H.K. is supported by an A*STAR National Science Scholarship. M.L.B. is supported by the NIH grant R21 HG010200. S.K.G. is supported by R01HL139672, R01HL122684 and R01HL086694. D.E.B. was supported by the NHLBI (P01HL32262 and DP2HL137300). The FHS is funded by National Institutes of Health contract N01-HC-25195. The eQTL work for this investigation was funded by the Division of Intramural Research, National Heart, Lung, and Blood Institute, National Institutes of Health, Bethesda, MD (D. Levy, Principal Investigator). The analytical component of this project was funded by the Division of Intramural Research, National Heart, Lung, and Blood Institute, and the Center for Information Technology, National Institutes of Health, Bethesda, MD. The views expressed in this manuscript are those of the authors and do not necessarily represent the views of the National Heart, Lung, and Blood Institute; the National Institutes of Health; or the US Department of Health and Human Services.
Author information
Authors and Affiliations
Contributions
A.C. and E.T. designed and performed the experiments. B.J.A., L.M.C., K.H.K., W.M., S.Y., B.H., A.L., M.S. and S.N.G. performed bioinformatics and data analysis. T.V.B., A.S., D.S.V. and A.G. helped design strategies for key experiments, including CRISPR-mediated mutations. R.J. and M.-L.Y. carried out the FHS data analysis. K.H., V.C., S.B. and S. Tseng performed supervised experiments. S. Takahashi provided the PU.1-knockdown and PU.1-overexpressing K562 cell lines. S.K.N. provided the luciferase plasmids containing non-coding alleles of the RBM38 gene and the enhancer mutant for the RBM38 gene in K562 cells. Y.Z., A.B.C., S.K.G., J.R., D.E.B., P.S.A., S.J.C., M.L.B. and R.A.Y. provided insights on the analysis and interpretation of data. A.C., E.T., B.J.A., L.M.C., K.H.K., A.S. and L.I.Z. wrote and revised the manuscript. L.I.Z. supervised the study. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.I.Z. is a founder and stockholder of Fate Therapeutics, CAMP4 Therapeutics, Amagma Therapeutics, and Scholar Rock. He is a consultant for Celularity and Cellarity. R.A.Y. is a founder and shareholder of Syros Pharmaceuticals, Camp4 Therapeutics, Omega Therapeutics and Dewpoint Therapeutics. B.J.A. is a shareholder in Syros Pharmaceuticals. M.L.B. is a co-inventor on patents on PBM technology. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work.
Extended data
Extended Data Fig. 1 Human CD34+ cells commit to an erythroid fate around day 3 (D3) of differentiation.
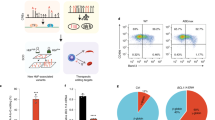
a, Representative FACS plots for the erythroid markers CD71 and CD235a on CD34+ cells after induction of erythroid differentiation at hour 6 (H6), day 2 (D2), day 3 (D3), day 4 (D4), day 5 (D5) and day 7 (D7). b-c, Heatmaps depicting correlation of peaks from H3K27ac ChIP-seq or ATAC-seq obtained from distinct differentiation time-points. d-e, Gene tracks showing H3K27ac ChIP-seq signal (d) or ATAC-seq signal (e) at FLI1 and at the β-globin locus control region at different differentiation stages. D0 = progenitor CD34+ cells before induction of differentiation; H6 = 6 hours after differentiation; and D1 through D5 = 1, 2, 3, 4 and 5 days after differentiation. f, Heatmap depicting correlation of gene expression profiles of all protein-coding RNAs from D0 through D8 of erythroid differentiation. D0 = progenitor CD34+ cells before induction of differentiation; H2 and H6 = 2 and 6 hours after differentiation; and D1 through D8 = 1, 2, 3, 4, 5, 6, 7 and 8 days after differentiation. g, Signal heatmaps comparing ChIP-seq read densities of SMAD1, GATA2, and GATA1 at SMAD1 peaks identified at D0 (upper panel) and D5 (lower panel). Signal intensities centered around +/− 2.5 kb shown. h, Ingenuity pathway analysis (IPA) for GATA2+SMAD1 bound genes at D0, H6, D3 and D4 and GATA1+SMAD1 bound genes at D3, D4, D5 identifying differentiation stage-specific biological properties.
Extended Data Fig. 2 Comparative TF motif enrichment and H3K27ac signal density analysis surrounding GATA+SMAD1 versus GATA-only regions.
Bar charts depicting the enrichment of transcription factor motif hits at regions co-bound by GATA+SMAD1 (left) versus by GATA only (right) at H6 (a) and D5 (b). Length of the bar indicates the fraction of peaks containing a given motif hit, and the number associated with the bar represents the corresponding -log10(p-value) obtained from the one-tailed hyper-geometric test to assess the significance of motif enrichment. For both (a) and (b), top and bottom of the ranked lists are shown. c, (left panel) Region heatmaps depicting signal of ChIP-seq reads for D0 SMAD1, GATA2 and H3K27ac at 33,470 GATA2 bound peaks at D0. Peaks are ranked by the SMAD1 intensity across the row. Each plot represents signal intensities around +/− 2 kb of the peak center. (right panel) Region heatmaps depicting signal of ChIP-seq reads for D5 SMAD1, GATA1 and H3K27ac at 10,855 GATA1 bound peaks at D5. Peaks are ranked by the SMAD1 intensity across the row. Each plot represents signal intensities around +/− 2 kb of the peak center.
Extended Data Fig. 3 TSCs are a small subset of overall enhancers as defined by SMAD1 and GATA co-occupancy.
a, Venn diagram representing genomic regions co-occupied by different STFs - SMAD1, SMAD2 and TCF7L2 in progenitor CD34+ cells upon stimulation with BMP4, TGFβ and WNT signaling, respectively. The genomic regions bound by all three factors are 4549. The other numbers refer to the total number of peaks bound by each factor combination, as indicated. Regions are considered occupied if they pass a significant coverage cutoff. b, Table representing different strategies to define enhancers using H3K27ac ChIP-seq and/or ATAC-seq. The proportion of enhancers that can be classified as TSCs (GATA+SMAD1 co-bound) at progenitor (D0) or erythroid (D5) stages are as indicated.
Extended Data Fig. 4 Dissecting a TSC near the LHFPL2 gene by CRISPR-Cas9.
a, sgRNAs (shown in brown) targeting specific sequences near PU.1 (pink), GATA (red) and SMAD1 (green) motif-hits within the TSCs. b, Genomic sequences of the specific CRISPR-edited clones are compared against wild-type genomic sequence. Clones 14.13, 16.03 and 15.16 appear to target multiple motifs (that is PU.1 and partial GATA; partial PU.1 and partial GATA; SMAD1 and partial GATA). c, qPCR results depicting relative expression of LHFPL2 (black bar) and two other flanking genes (SCAMP1, grey bar and AP3B1, white bar) are shown in different CRISPR clones compared to the WT K562 cells, as indicated. Mean ± SEM shown. (n = 3; 3 biologically independent experiments). Two-sided student t-tests used.
Extended Data Fig. 5 Approach for interrogating enhancer-associated RBC-trait SNPs showing that SNPs targeting STF motif-hits are localized within TSCs.
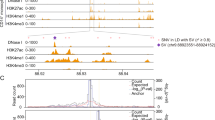
a, Schematic diagram of the strategy used to identify SNPs that may alter activity of transcriptional enhancers during human erythroid differentiation. Human CD34+ cells from mobilized peripheral blood were differentiated towards erythrocytes. Genomic experiments were performed at D0, H6, D3, D4 and D5. 1270 lead RBC-trait SNPs and additional SNPs that are in linkage disequilibrium with lead SNPs, with LD score r2 ≥ 0.6 (total number of SNPs = 29,069), were first overlapped with genomic regions that are defined as non-exonic enhancer (represented as violet tracks) and open chromatin peaks (represented as grey tracks) in our study. SNPs that fall within such regions (indicated with red arrows) were used to carry out motif hit analysis, and were overlapped either with TSCs, or overall enhancers or GATA-only enhancers. b, Gene tracks showing RBC-trait SNPs that are located within stage-specific TSCs are shown. The binding of GATA2, GATA1, SMAD1, PU.1 and KLF1 and the peaks of H3K27ac and ATAC-seq are shown in progenitor and differentiated stages. Black lines indicate the positions of representative SNPs. The potential STF motifs that these SNPs could perturb (for example GLI, EGR) are as indicated. For each representative SNP that resides in a TSC, the other associated SNPs in significant LD that fall within H3K27ac/ATAC-positive enhancers are also indicated with grey dashed lines.
Extended Data Fig. 6 Analysis of PBM 8-mer data identifies several RBC trait-associated SNPs that perturb STF-DNA binding.
a, Schematic representation of the strategy to identify SNPs that alter STF binding utilizing protein binding microarrays. b, Bar charts for the GATA average PBM dataset for rs737092. The p-value (0.5469) is computed using two-sided Wilcoxon signed-rank test. c, Additional examples of SNPs showing perturbed TF binding of indicated STFs from PBM analysis (left) and corresponding distribution of expression values of the most significantly altered nearby gene in homozygous and heterozygous individuals obtained from FHS eQTL analysis (right). For PBM analysis, the P-values are computed using two-sided Wilcoxon signed-rank tests. Individual genotypes and the cis-eQTL gene/exon obtained from the FHS dataset are as indicated. Two-sided test with linear model for EffectAlleleDosage used for eQTL analysis with P-values adjusted using the Benjamini-Hochberg procedure. The lower and upper bounds of the box correspond to the 25th and 75th percentiles, respectively. The upper whisker extends from the upper bound of the box to the largest value no further than 1.5 * inter-quartile range (IQR) away (the IQR is defined as the distance between the 25th and 75th percentiles). The lower whisker extends from the lower bound of the box to the smallest value no further than 1.5 * IQR away.
Extended Data Fig. 7 RBC-trait SNPs perturb STF-DNA binding.
a, TSC containing rs9467664 at SMAD motif near HIST1H4A. b, HIST1H4A expression during CD34+ differentiation. c, Western blot showing the expression of FLAG-SMAD1. TBP is loading control. (n = 3; 3 biologically independent experiments). d, Representative gel-shift assay with A or T allele of rs9467664. (n = 3; 3 biologically independent experiments). e, HIST1H4A eQTL analysis for rs9467664: boxplots represent median HIST1H4A64 expression as the thickest line, the first and third quartile as the box, and 1.5 times the interquartile range as whiskers. Two-sided test with linear model for EffectAlleleDosage used: effect estimate (β)=0.1562; T-statistics=31.0243, R2 = 0.15536; log10(P-value)=-193.41, log10(Benjamin–Hochberg’s FDR) = -190.1. f, Schematic representation of K562 clone with altered sequence around rs737092. g, Alteration of TF binding in K562 mutants from f. Mean ± SEM shown. (n = 3; 3 biologically independent experiments). Two-sided student t-tests used. h, Expression alteration of RBM38 in K562 mutants from f, under BMP and BIO treatment. Mean ± SEM shown. (n = 3; 3 biologically independent experiments). Two-sided student t-tests used. i, Luciferase assays for alternative alleles of rs737092. Boxplots represent median as the thickest line, the first and third quartile as the box, and 1.5 times the interquartile range as whiskers. Two-sided student t-tests used. j, Western blot comparing SMAD1 expression in control and shRNA treated CD34+ cells with indicated doses. TBP is loading control. (n = 3; 3 biologically independent experiments). k, RBM38 expression upon SMAD1 loss. Mean ± SEM shown. (n = 3; 3 biologically independent experiments). Two-sided student t-tests used.
Extended Data Fig. 8 A model proposing how human genetic variation within TSCs induces RBC trait phenotypes.
A combination of STFs and MTFs drives optimal gene expression via the TSC. The normal signal-induced expression of a red blood cell gene is perturbed due to a SNP that either eliminates an existing STF binding event or creates a new STF binding site in a critical signaling center. This can lead to a lack of response to an episodic signaling event, initiated by an exogenous stressor, and eventually manifest as phenotypic variability.
Supplementary information
Supplementary Information
Supplementary Notes 1–5 and Fig. 1
Supplementary Tables
Supplementary Tables 1–16
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Source genomic regions and genes for GREAT analysis.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data for PBM analysis.
Source Data Fig. 7
Statistical source data for FHS analysis.
Source Data Extended Data Fig. 2
Statistical source data for comparative motif analysis.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data for PBM analysis.
Source Data Extended Data Fig. 7
Unprocessed western blots/gels.
Rights and permissions
About this article
Cite this article
Choudhuri, A., Trompouki, E., Abraham, B.J. et al. Common variants in signaling transcription-factor-binding sites drive phenotypic variability in red blood cell traits. Nat Genet 52, 1333–1345 (2020). https://doi.org/10.1038/s41588-020-00738-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-00738-2
This article is cited by
-
Soft computing techniques for biomedical data analysis: open issues and challenges
Artificial Intelligence Review (2023)
-
Integrating transcription factor occupancy with transcriptome-wide association analysis identifies susceptibility genes in human cancers
Nature Communications (2022)
-
Revealing modifier variations characterizations for elucidating the genetic basis of human phenotypic variations
Human Genetics (2022)