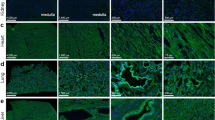
Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which causes COVID-19, utilizes angiotensin-converting enzyme 2 (ACE2) for entry into target cells. ACE2 has been proposed as an interferon-stimulated gene (ISG). Thus, interferon-induced variability in ACE2 expression levels could be important for susceptibility to COVID-19 or its outcomes. Here, we report the discovery of a novel, transcriptionally independent truncated isoform of ACE2, which we designate as deltaACE2 (dACE2). We demonstrate that dACE2, but not ACE2, is an ISG. In The Cancer Genome Atlas, the expression of dACE2 was enriched in squamous tumors of the respiratory, gastrointestinal and urogenital tracts. In vitro, dACE2, which lacks 356 amino-terminal amino acids, was non-functional in binding the SARS-CoV-2 spike protein and as a carboxypeptidase. Our results suggest that the ISG-type induction of dACE2 in IFN-high conditions created by treatments, an inflammatory tumor microenvironment or viral co-infections is unlikely to increase the cellular entry of SARS-CoV-2 and promote infection.
Similar content being viewed by others
Main
Cells expressing ACE2 are potential targets of SARS-CoV-2 infection1,2. Studies based on single-cell RNA sequencing (scRNA-seq) of lung cells have identified type II pneumocytes, ciliated cells and transient secretory cells as the main types of ACE2-expressing cell3,4. Furthermore, ACE2 was proposed to be an ISG, on the basis of its inducible expression in cells treated with interferons (IFNs) or infected by viruses that induce IFN responses, such as influenza4,5. These findings implied that the induction of ACE2 expression in IFN-high conditions could result in an amplified risk of SARS-CoV-2 infection4,5. Concerns could also be raised about possible ACE2-inducing side effects of IFN-based treatments proposed for COVID-19 (refs. 6,7,8,9).
ACE2 plays multiple roles in normal physiological conditions and as part of the host tissue-protective machinery in damaging conditions, including viral infections. As a terminal carboxypeptidase, ACE2 cleaves a single carboxy-terminal residue from peptide hormones such as angiotensin II and des-Arg9-bradykinin. ACE and ACE2 belong to the renin–angiotensin–aldosterone system, which regulates blood pressure and fluid–electrolyte balance; dysfunction of this system contributes to comorbidities in COVID-19 (refs. 10,11). des-Arg9-bradykinin is generated from bradykinin and belongs to the kallikrein–kinin system, which is critical in regulating vascular leakage and pulmonary edema, early signs of severe COVID-19 (refs. 12,13).
High plasma angiotensin II levels were found to be responsible for coronavirus-associated acute respiratory distress syndrome (ARDS), lung damage and high mortality in mouse models14,15 and as a predictor of lethality in avian influenza in humans16,17. In the same conditions, ACE2, which decreases the levels of angiotensin II, was identified as a protective factor. The hijacking of the normal host tissue-protective machinery guarded by ACE2 was suggested as a mechanism through which SARS-CoV-2 could infect more cells4,5. Thus, it is critically important to identify factors affecting ACE2 expression in normal physiological processes and during viral infections and associated pathologies, such as in COVID-19.
Herein, aiming to explore the IFN-inducible expression of ACE2 and its role in SARS-CoV-2 infection, we identified a novel, truncated isoform of ACE2, which we designate as dACE2. We then showed that dACE2, but not ACE2, is induced in various human cell types by IFNs and viruses; this information is important to consider for future therapeutic strategies and understanding COVID-19 susceptibility and outcomes.
Results
dACE2 is a novel inducible isoform of ACE2
To address the extent to which IFNs induce the expression of ACE2 in human cells, we used our existing RNA-seq dataset (NCBI Sequence Read Archive (SRA): PRJNA512015) of a breast cancer cell line T47D infected with Sendai virus (SeV), known to be a strong inducer of IFNs and ISGs18,19,20. IFNs were not expressed in T47D cells at baseline, but SeV strongly induced expression of IFNB1, a type I IFN, and all type III IFNs (IFNL1, 2, 3 and 4). Several well-known ISGs (ISG15, MX1 and IFIT1) were moderately expressed at baseline but were strongly induced by SeV (Supplementary Table 1). ACE2 was not expressed at baseline but was strongly induced by SeV, exclusively as an isoform initiated from a novel first exon in intron 9 of the full-length ACE2 gene (Fig. 1a,b).
a, A UCSC Genome Browser view of the human ACE2 region (chrX: 15,560,521–15,602,580, GRCh38/hg38) showing alternative first exons ACE2-Ex1a, ACE2-Ex1b and Ex1c, a novel first exon that creates a truncated ACE2 isoform designated as dACE2. The combination of various ENCODE epigenetic marks for human cells, with H3K4me1, H3K4me3 and H3K27ac shown as peaks, and a cluster of DNase I hypersensitivity sites shown as bars indicates that dACE2-Ex1c is located within a putative regulatory region that can affect gene expression. The DNase I hypersensitivity site overlying Ex1c is detected in 38 of 95 cell lines tested. b, RNA-seq Sashimi plots depicting splicing patterns defining ACE2 and dACE2 isoforms in SeV/mock-infected T47D cells and uninfected RT-4 cells. The numbers on the Sashimi plots indicate the counts of exon–exon splicing reads. c, ACE2 is a single-span transmembrane protein with a signal peptide (SigP) of 17 aa and four functional domains—peptidase domain (PD, aa 18–615), collectrin-like domain (CLD, aa 616–740), transmembrane domain (TM, aa 741–761) and intracellular domain (ICD, aa 762–805). In dACE2, the signal peptide is not predicted; the peptidase domain starts from aa R357; the first 356 aa are replaced by 10 aa of a unique protein sequence; * and ** mark cleavage sites of the membrane-bound proteases ADAM17 and TMPRSS2, respectively. d,e, Sequence alignments of the 5' UTR (d) and protein sequences encoded by dACE2-Ex1c and part of the downstream exon (e) in select primates. TSS, transcription start site. dACE2 is not predicted to be encoded in any non-primate species. Additional alignment analyses are shown in Supplementary Fig. 1a,b.
RNA-seq analysis in T47D and RT-4 cell lines (Fig. 1b) demonstrated that ACE2 exists as two full-length transcripts initiated from two independent first exons, which we designated as Ex1a and Ex1b (the latter is shared between these transcripts). Additionally, an alternative transcript was initiated from the novel first exon in intron 9, which we designated as Ex1c (Fig. 1b). The combination of ENCODE chromatin modification marks (H3K4me1, H3K4me3 and H3K27ac) and a cluster of DNase I hypersensitivity sites (Fig. 1a) suggests that Ex1c, but not Ex1a and Ex1b, is located within a putative regulatory region that might affect gene expression.
The novel ACE2 isoform is predicted to encode a protein of 459 amino acids (aa), including the first 10 aa encoded by Ex1c. Compared to the full-length ACE2 protein of 805 aa, the truncation eliminates 17 aa of the signal peptide and 339 aa of the N-terminal peptidase domain (Fig. 1c). We designate this novel isoform as dACE2 (NCBI GenBank accession number MT505392). Analysis of 100 vertebrate species with genomic sequences available through the UCSC Genome Browser showed that the putative dACE2 protein could be encoded only in primates (Supplementary Fig. 1a). Comparison of human Ex1c and its proximal promoter in select species showed 96.7–99.6% of sequence identity in primates and 54.0–73.5% in non-primate mammals (Supplementary Fig. 1b). In primates, despite some differences on the messenger RNA level, there was strong conservation within the putative protein encoded by dACE2-Ex1c (Fig. 1d,e).
Several binding motifs for transcription factors relevant for IFN signaling were predicted within the promoter of dACE2-Ex1c (P3; Fig. 2a). In contrast, ISG-type motifs were not predicted in the promoters of ACE2-Ex1a (P1) and ACE2-Ex1b (P2; Fig. 2a). We evaluated the IFN-inducible activity of all three promoters (ACE2-P1, ACE2-P2 and dACE2-P3) by testing their ability to drive expression of the luciferase reporter (Fig. 2b). The reporter constructs were transiently transfected into HepG2 cells, in which the signaling of all IFNs has been reported. Only dACE2-P3 significantly induced luciferase expression in response to 6 h of treatment with IFN-β or IFN-γ (Fig. 2c). The deletion of the first 100 base pairs of the dACE2-P3 promoter resulted in the loss of luciferase activity, suggesting that the predicted ISG-type motifs in the proximal promoter are important for IFN-driven dACE2 expression (Fig. 2c). The promoter of IFIT1, an ISG, was used as a positive control and, as expected, was strongly responsive to treatments with IFNs (Fig. 2d).
a, Promoters of ACE2 (P1 and P2) and dACE2 (P3) were analyzed for binding motifs of transcription factors relevant for IFN signaling. Promoters were defined within the −800/+100 bp windows from the corresponding TSSs. ISRE, interferon-stimulated response element. b, Schematics of luciferase (Luc) reporter constructs. c, Luciferase activity in HepG2 cells transiently co-transfected with the indicated luciferase reporter constructs and Renilla (normalization control) and treated with 1 ng ml−1 of IFN-β or 2 ng ml−1 of IFN-γ for 6 h. d, Luciferase activity driven by the promoter of IFIT1 (an ISG and positive control). Luciferase/Renilla ratios were normalized by corresponding mock-treated samples and are presented as fold change to the negative control (empty promoterless pGL4.21 vector). The P values are for unpaired, two-sided Student’s t-tests. The experiment was conducted in six biological replicates per construct, and the results of one of three independent experiments are shown.
dACE2 is induced by in vitro treatment with IFNs
We confirmed the SeV-induced expression of the full-length dACE2 by PCR with reverse transcription (RT–PCR; Fig. 3a,b) and verified the corresponding PCR products by Sanger sequencing. Using custom-designed assays, we explored ACE2 and dACE2 expression in multiple cell lines at baseline and after SeV infection (Fig. 3c and Supplementary Table 2a). In most cell lines tested, dACE2 but not ACE2 was strongly upregulated by SeV infection (Fig. 3b,c). To directly address whether IFN was responsible for the induced expression of dACE2, we performed expression analysis in primary normal human bronchial epithelial (NHBE) cells21 and human intestinal (colon and ileum) organoid cultures22. In NHBE cells from five healthy donors, the baseline expression levels of dACE2 and ACE2 were comparable, but only dACE2 was significantly induced by treatment with IFN-α or IFN-λ3 (Fig. 3d and Supplementary Table 2b). In contrast, ACE2 was expressed at high levels already at baseline both in colon and ileum organoid cultures, while the expression of dACE2 was very low. Treatments with IFN-β or a cocktail of IFN-λ1–3 significantly induced only expression of dACE2 and not ACE2 (Fig. 3e and Supplementary Table 2c). In both cell models, the expression pattern of dACE2 was similar to that of the known ISGs—MX1 (Fig. 3d and Supplementary Table 2b) and IFIT1 (Fig. 3e and Supplementary Table 2c).
a, A schematic representation of ACE2 and dACE2 transcripts and the positions of the forward (F) and reverse (R) PCR primers to generate full-length dACE2 amplicons. b, An agarose gel showing an RT–PCR product of 1,535 bp corresponding to full-length dACE2 in several cell lines with/without SeV infection. c, A heatmap of ACE2 and dACE2 expression and a positive control ISG (ISG15) measured by TaqMan expression assays in human cell lines infected with SeV for 12 h; the colors represent expression differences as ddCt normalized by endogenous controls (GAPDH and ACTB) and comparing SeV-infected to uninfected samples. d, The expression of ACE2 and dACE2 and a positive control ISG MX1 in primary NHBE cells from five healthy donors; NHBEs were untreated or treated with IFN-α or IFN-λ3 for 24 h. The data are presented as means and s.d. The P values are for paired Student’s t-tests. e, The expression of ACE2, dACE2 and a positive control ISG IFIT1 in colon and ileum organoid cultures from one donor; the organoids were treated with IFN-β or a cocktail of IFN-λ1–3 for 24 h in three biological replicates. The results are presented with means and s.d.; the P values are for unpaired, two-sided Student’s t-tests. The full expression results are presented in Supplementary Table 2b,c.
dACE2 is induced in virally infected human respiratory cells
To investigate whether dACE2 expression is induced by RNA viruses, which are potent inducers of the IFN response, we de novo quantified the expression of ACE2-Ex1a, ACE2-Ex1b and the newly annotated dACE2-Ex1c in several public RNA-seq datasets of virally infected human respiratory epithelial cells. In an RNA-seq dataset of human nasal airway epithelial cells from patients with asthma ex vivo infected with respiratory rhinovirus strains RV-A16 and RV-C15 (NCBI SRA: PRJNA627860), both ACE2 and dACE2 were expressed (Fig. 4a). Compared to that in uninfected cells, dACE2-Ex1c expression was strongly induced by both viruses—by RV-A16 (2.58-fold) and RV-C15 (2.42-fold)—while expression of ACE2-Ex1b was moderately induced only by RV-C15 (1.13-fold; Fig. 4b). Only dACE2 expression strongly correlated with multiple ISGs and IFNs (Fig. 4c). Similarly, in human lung explants infected with a seasonal influenza A/H3N2 strain (NCBI SRA: PRJNA557257), only dACE2 was induced by infection, and its expression correlated with the levels of IFNs and ISGs (Fig. 4d,e).
a–e, Expression patterns of ACE2, represented by Ex1a and Ex1b, and dACE2, represented by Ex1c, in uninfected and rhinovirus (RV)-infected human nasal epithelial cells (a–c) and in uninfected and influenza H3N2–infected cells from human lung explants (d,e). a, RNA-seq Sashimi plots. b,d, First-exon-specific expression levels of ACE2 and dACE2 are presented by box-and-whisker plots as: mean (+), median (center line), box (25th to 75th interquartile range), whiskers (5th–90th percentile range) and outliers (individual data points). FC, fold change compared to mock. The P values are for non-parametric, two-sided Wilcoxon matched-pairs signed rank tests. c,e, Pearson correlation coefficients with P values for two-sided tests for exon-specific expression levels of ACE2 and dACE2 with select ISGs and IFNs in the full datasets.
It was reported that, in contrast to ACE2 expression in human cells, Ace2 was not induced in primary mouse tracheal basal cells in response to in vitro and in vivo IFN stimulation, and on in vivo viral infection4. To explore this further, we analyzed a dataset for human and mouse lung cells infected with the respiratory syncytial virus (NCBI SRA: PRJNA588982). Indeed, we did not observe induction of Ace2 in mouse lung cells (Extended Data Fig. 1a), while dACE2 but not ACE2 was induced in a human pulmonary carcinoma cell line (H292) infected with the respiratory syncytial virus (Extended Data Fig. 1b). These results illustrate that the identification of ACE2 as an ISG4,5 was likely based on the detection of inducible expression of dACE2, since 3'-scRNA-seq would detect both ACE2 and dACE2. The differences between the human and mouse sequences corresponding to dACE2-Ex1c and promoters (Supplementary Fig. 1b) might be responsible for the lack of ISG-type ACE2 expression in mice (Extended Data Fig. 1a). We also analyzed Ace2 expression in an RNA-seq dataset of nasal washes from mock/SARS-CoV-2-infected ferrets and did not observe any dAce2-type transcripts (Extended Data Fig. 1c).
We also tested ACE2 and dACE2 expression in a commonly used cell line, Vero E6, derived from green monkey kidney. ACE2 expression was high at baseline but not inducible by treatments with IFN-β or IFN-λ1 (which induced an ISG control, MX1), while dACE2 was not detected in any conditions (Supplementary Table 2b). In comparison, in the human kidney cells HEK293T, ACE2 expression was also high at baseline and not inducible by IFN treatment, while dACE2 was moderately induced by IFN-β (Supplementary Table 2b). Although dACE2 was not inducible by IFNs in Vero E6, while being moderately expressed in HEK293T cells, this should be further tested in additional cell lines and primary tissues before making conclusions about whether dACE2 could be induced in non-human primates.
dACE2 is enriched in squamous epithelial tumors
We explored the expression patterns of dACE2 in various human tissues. In a dataset of 95 normal human tissues of 27 types, dACE2-Ex1c was detectable in select tissues but at very low levels (≤10 RNA-seq reads), while ACE2-Ex1b expression was common (Extended Data Fig. 2). In the set of normal human tissues from the Genotype-Tissue Expression (GTEx) project, only total gene expression was available for ACE2, with the highest expression observed in the testes and small intestine (Supplementary Fig. 2). We hypothesized that as an ISG, dACE2 might be absent or expressed at low levels in normal tissues, but could be induced by the inflammatory tissue microenvironment. We explored the data from The Cancer Genome Atlas (TCGA), which represents the largest collection of tumors and tumor-adjacent normal tissues, and de novo quantified the expression of ACE2 and dACE2 in all TCGA samples (Supplementary Table 2f–h). Expression of both ACE2 and dACE2 was detectable in many tumor-adjacent normal tissues (Fig. 5a and Extended Data Fig. 3). In the set of 10,185 TCGA tumors of 33 cancer types, ACE2-Ex1a, ACE2-Ex1b and dACE2-Ex1c were expressed in 12.6, 38.0 and 16.8% of tumors, respectively, with ≥5 RNA-seq reads per sample. dACE2 was most expressed in head and neck squamous carcinoma (HNSC) and lung squamous carcinoma (LUSC), which represent oral and bronchial mucosal epithelial surfaces, while ACE2 was most expressed in kidney tumors (Extended Data Fig. 4).
a, A bubble plot showing the mean expression levels (RNA-seq read counts) and proportions of samples with dACE2-Ex1c in TCGA tumor-adjacent normal tissues and 33 tumor types. dACE2-Ex1c is expressed at relatively high levels and in many tumors of the bladder (BLCA), cervix (CESC), esophagus (ESCA), head and neck (HNSC) and lung squamous carcinoma (LUSC). b, RNA-seq counts of ACE2-Ex1a, ACE2-Ex1b and dACE2-Ex1c in tumor-adjacent normal tissues (AN), squamous tumors (SQ) and non-squamous tumors (NSQ). The dACE2 expression is significantly higher in squamous compared to non-squamous tumors of the same tissue origin and corresponding tumor-adjacent normal tissues. Specifically, dACE2-Ex1c is expressed similarly in tumor-adjacent normal tissues adjacent to LUSC and lung adenocarcinoma (LUAD), while it is significantly higher in corresponding tumors and higher in LUSC than in LUAD, due to the clonal origin of these tumors from cells with differential expression of dACE2. The box-and-whisker plots represent mean (+), median (center line), box (25th–75th interquartile range), whiskers (5th–90th percentile range) and outliers (individual data points). The P values are for non-parametric, two-sided Mann–Whitney U tests.
Generally, dACE2 expression was enriched in squamous tumors representing epithelial tracts. Squamous carcinomas of the lung (LUSC) and head and neck (HNSC) represent respiratory tract, esophageal cancer (ESCA)–upper gastrointestinal, and bladder cancer (BLCA) and cervical squamous carcinoma (CESC)–urogenital tract (Fig. 5a). In each tumor type, dACE2 expression was significantly higher in squamous compared to non-squamous tumors and adjacent normal tissues (Fig. 5b). As tumors represent results of clonal expansion of individual cells, this analysis highlighted the differential etiology (squamous versus non-squamous) of ACE2- and dACE2-expressing cells in various tissues.
dACE2 expression is IFN-γ inducible
In addition to the cell-type origin, the observed enrichment of dACE2 expression in some tumors might reflect persistent IFN exposure due to an inflammatory tumor microenvironment or underlying infection. An IFN-γ-induced signature emerged as a prominent feature in airway epithelial cells of patients with COVID-19, and this signature was linked with enrichment of cytotoxic T lymphocytes5. Unlike normal tissues, tumors are extensively infiltrated by immune cells, making the TCGA dataset particularly informative for the analysis of IFN signatures. IFNG was the most commonly expressed IFN gene in TCGA tumors (with 61% of all tumors expressing IFNG at RSEM ≥ 1, mean RSEM = 19.8), while other IFNs were expressed at low levels (mean RSEM ≤ 1.3; Extended Data Fig. 5a). dACE2-Ex1c levels significantly correlated (P ≤ 0.01, r ≥ 0.2) with IFNG expression in 8 out of 32 tumor types tested (Extended Data Fig. 5b), while ACE2-Ex1b showed only moderate and predominantly negative correlations with IFNG expression in some tumor types (Extended Data Fig. 5c). Furthermore, in vitro treatment with IFN-γ significantly induced dACE2, but not ACE2 (Extended Data Fig. 5d). Thus, in addition to type I and type III IFNs (Fig. 3e–g), dACE2 expression might be partly driven by IFN-γ contributed by tumor-infiltrating immune cells or inflamed virally infected tissues.
In the initial analysis of the TCGA-LUSC dataset (n = 501), which represents ACE2- and dACE2-expressing tumors of bronchial origin (Fig. 5b), IFNG expression did not correlate with dACE2 (Extended Data Fig. 5b). To further investigate ACE2 and dACE2 expression in this tumor dataset, we used an unsupervised machine learning approach to assign all LUSC tumors to 6 clusters based on the expression of 270 ISGs23 (Extended Data Fig. 6a,b). A set of ISGs (n = 100) that most strongly contributed to the definition of these clusters was used for correlation analysis. The analysis of a cluster that included 114 LUSC tumors with the highest ISG expression (cluster 5) showed that dACE2 was strongly and significantly (false discovery rate (FDR) P value < 0.05) correlated with the expression of 20 ISGs and ACE2—with 5 ISGs (Extended Data Fig. 6c,d). Thus, ISG-type dACE2 expression could be contributed by various factors, possibly determined by cell- and tissue-specific microenvironments and exposures.
dACE2 is induced by SARS-CoV-2 in vitro
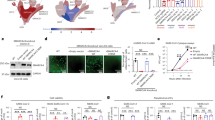
Once we established that dACE2 is an ISG in multiple human cell types under various conditions, we tested whether its expression could also be induced by SARS-CoV-2. There was a noticeable difference in the baseline expression levels of ACE2 and dACE2 in three cell lines tested (Calu3, Caco-2 and T84). Expression of ACE2 and dACE2 was much higher in the lung adenocarcinoma cell line Calu3 compared to both the colon adenocarcinoma cell lines Caco-2 and T84 (Fig. 6a and Supplementary Table 2d). The baseline dACE2 expression in T84 was higher than in Caco-2, in line with the RT–PCR results (Fig. 1b). All cell lines were successfully infected with SARS-CoV-2 (Fig. 6b and Supplementary Table 2d), but ACE2 expression was not affected by infection in any cell line tested (Fig. 6a and Supplementary Table 2d). Induction of dACE2 expression tracked with previously reported SARS-CoV-2 infectivity rates in these cells22. Specifically, dACE2 was most strongly induced in Caco-2 cells, in which over 80% of cells were infected by 24 h, moderately induced in T84 cells (20% of cells were infected) and not induced in Calu3 cells (10% of cells were infected). A similar expression pattern was observed for an ISG, IFIT1, except for in Calu3 cells, in which only IFIT1 was significantly induced (Fig. 6a and Supplementary Table 2d). We performed similar analyses in human colon and ileum organoid cultures derived from three donors. Overall, the expression of dACE2 and IFIT1, but not of ACE2, was significantly induced by SARS-CoV-2 infection (Fig. 6c,d and Supplementary Table 2e).
a–d, Expression of ACE2, dACE2 and a control ISG IFIT1 in the colon cancer cell lines Caco-2 and T84 and the lung cancer cell line Calu3 (all in three to four biological replicates) (a) and colon and ileum organoid cultures from three donors (c), and the SARS-CoV-2 viral loads in the corresponding cells (b,d). The P values are for two-sided Student’s t-tests, unpaired tests for cell lines and paired tests for organoids. The results are presented with means and s.d.
dACE2 is non-functional as a SARS-CoV-2 receptor and peptidase
Despite the strong induction of dACE2 mRNA expression, we were unable to detect endogenous dACE2 in SeV-infected cell lines by western blotting with commercial antibodies for ACE2 (data not shown). However, in the proteome database of mass spectrometry data available for breast, colon and ovarian TCGA tumors24, we detected human-specific peptides matching the 10 aa encoded by dACE2-Ex1c (Extended Data Fig. 7), suggesting that the dACE2 protein could be expressed in some conditions.
Transiently overexpressed dACE2–GFP was detected on the cell surface, although at levels lower than ACE2–GFP (Extended Data Fig. 8). However, the substantial N-terminal truncation by 356 aa in the peptidase domain of the putative dACE2 protein is expected to have important functional consequences compared to the activity of the full-length ACE2 protein of 805 aa. For example, decreased or no binding by the SARS-CoV-2 spike receptor-binding domain (RBD) would be expected for dACE2. Indeed, only cells overexpressing ACE2–GFP but not GFP alone or dACE2–Myc were able to bind and internalize the spike RBD (Fig. 7a–c). Compared to the case for ACE2–GFP alone, a moderately increased ACE2–GFP expression and binding of spike RBD was observed in cells co-expressing dACE2–Myc and ACE2–GFP (Fig. 7d,e and Extended Data Fig. 9a), which could suggest that dACE2 increases infection. However, when we transiently transfected ACE2–GFP with a plasmid for an unrelated transmembrane protein, TMEM129–Myc, we observed a similar pattern (Extended Data Fig. 9b,c). This suggests that the observed effect of dACE2 on the increased ACE2 levels and spike RBD binding might be a non-specific effect due to transient overexpression of multiple plasmids.
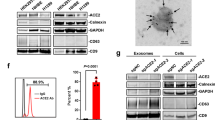
a, Representative confocal images of T24 cells transiently overexpressing dACE2–Myc (white) or ACE2–GFP (green) and treated with the RBD of the SARS-CoV-2 spike protein (red); nuclei (DAPI), blue. Scale bars, 20 µm. b–e, Representative flow cytometry histograms (b,d), and mean fluorescence intensity (MFI) values from three biological replicates (c,e), of spike RBD binding to the surface of ACE2–GFP- but not dACE2–Myc-expressing T24 cells. Gating for cells expressing dACE2–Myc, ACE2–GFP or both proteins is shown in Extended Data Fig. 9a. The results are based on three biological replicates, and represent one of two independent experiments. f, SARS-CoV-2 infectivity rates (%) in a lung cancer cell line, A549, transfected with GFP or dACE2–Myc, or stably expressing ACE2 (ACE2-stable) and transfected with GFP. g, The SARS-CoV-2 viral load as ddCt values compared to mock, corresponding to the plot in f. Additional details are provided in Extended Data Fig. 10. h, A representative western blot with an anti-ACE2 antibody that detects both recombinant ACE2–GFP and dACE2–GFP overexpressed in T24 cells. The amount of the ACE2–GFP lysate was kept constant, while the amount of the dACE2–GFP cell lysate was increased and the difference in the lysate volume was compensated by the empty GFP vector. i, The results of carboxypeptidase assays using variable amounts of lysates of cells (as described in the plot in h), showing that the activity of ACE2 is not affected by increasing amounts of dACE2. The results are based on three biological replicates and are presented with means and s.d. The western blot shows the results of one representative replicate.
We then evaluated the effect of dACE2 expression on SARS-CoV-2 infection in the lung cancer cell line A549. The wild-type A549 cells (transfected with GFP as a transient transfection control) were not infected by SARS-CoV-2, even after transfection with dACE2–Myc, because of low expression of endogenous ACE2 (Extended Data Fig. 10a,b). However, the A549 cells stably expressing recombinant ACE2 (ACE2-stable) were infected (Fig. 7f,g and Extended Data Fig. 10c–h). These results further support the conclusion that, if expressed, dACE2 induced by viruses or IFNs is unlikely to increase SARS-CoV-2 infection.
The N-terminal truncation is also predicted to affect the carboxypeptidase activity of dACE2, which is important for its ability to cleave angiotensin II, des-Arg9-bradykinin and other substrates of ACE2. Indeed, we observed carboxypeptidase activity in lysates of cells transfected with ACE2–GFP but not with dACE2–GFP, and this activity was not affected by the addition of lysates of cells overexpressing dACE2–GFP (Fig. 7h,i).
Discussion
ACE2 was recently proposed to be an ISG because of its induction in IFN-high conditions, raising concerns about its potential role in increasing SARS-CoV-2 infection4,5 and the safety of IFN-based treatments proposed for COVID-19. Our discovery of dACE2, a truncated version of ACE2, demonstrates that it is dACE2 and not ACE2 that is induced by IFNs and viruses, including SARS-CoV-2. Overexpressed recombinant dACE2, however, did not appear to bind SARS-CoV-2 spike RBD or affect the binding of ACE2 in our in vitro experiments, thus suggesting that ISG-type induction of dACE2 would not increase viral entry.
Along with previously reported data3,4, our results indicate that the expression of both ACE2 and dACE2 is limited to specific cell populations and conditions, contributing to low levels of their expression when analyzed by bulk RNA-seq methods. Although scRNA-seq analyses provide more specific information about cell populations that express these transcripts, the commonly used 3'-scRNA-seq methods do not discriminate between ACE2 and dACE2. Thus, dACE2 expression should be considered in expression studies of ACE2 by various methods (RNA-seq, microarrays or targeted expression assays). By analyses in multiple human cell types and tissues, we showed that expression of dACE2, but not ACE2, is inducible by IFNs (type I, II and III) and viruses that induce IFN responses. Suppression of IFN signaling by SARS-CoV-2 has been reported by several studies25,26, possibly explaining only a moderate effect of SARS-CoV-2 infection on dACE2 induction in our experiments. While the levels and the role of type I and III IFNs in COVID-19 remain controversial9,27,28,29, high levels of IFN-γ in the peripheral blood of patients with COVID-19 have been reported5. Thus, in tissues, dACE2 could be induced owing to exposure to IFN-γ-expressing immune infiltrates30,31,32. Specifically, a 3'-scRNA-seq analysis showed ACE2 induction by SARS-CoV-2 infection in ciliated epithelia, where high levels of IFN-γ-producing immune cells were also detected5. Our results strongly suggest that the induction of dACE2 and not ACE2 was detected in these patients.
We explored the extensive TCGA dataset of more than 10,000 tumors in which we de novo quantified dACE2 expression based on RNA-seq data and concluded that IFN-γ-driven ISG signatures can be contributed by tumor-infiltrating immune cells. These conclusions can be extended to inflamed virally infected tissues, for which comparable RNA-seq data are limited by small sample sets, a low percentage of mappable reads due to substantially degraded input RNA, or unavailability of raw data due to patient privacy restrictions. Furthermore, the expression patterns observed in TCGA indicated that dACE2 expression might be intrinsically enriched in squamous epithelial cells, which give rise to corresponding tumors of the respiratory, gastrointestinal and urogenital tracts. We found that in normal primary bronchial respiratory cells, the baseline expression levels of dACE2 were comparable to those of ACE2, and further strongly induced by IFN treatments, suggesting some cell-type-specific role of dACE2, which should be further explored.
The detection of dACE2-specific peptides in some TCGA tumors and the predicted existence of dACE2 protein only in primates argue for its potentially important role. Although dACE2 expression was induced by IFNs and viruses in various human cell lines, dACE2 was not detected in these cell lines by western blotting with a C-terminal ACE2 antibody. The detection of endogenous dACE2 might require the generation of dACE2-specific antibodies. Alternatively, the translation of dACE2 mRNA might be tightly regulated to exist only in specific conditions, as has been found for several mRNAs33. Further studies are required to confirm dACE2 cell surface expression in stable expression systems. However, on the basis of our in vitro data, we conclude that dACE2 does not increase the binding and cellular access of SARS-CoV-2 or serve as a carboxypeptidase. Extrapolation of these findings into biological and COVID-19-related mechanisms should be carried out with caution until confirmed by studies based on endogenous dACE2.
Although possible ISG-type ACE2 induction was considered as a risk for increasing SARS-CoV-2 infection, ACE2 deficiency rather than overexpression is discussed as a greater problem potentially contributing to COVID-19 morbidity11,12,13,34. Functional ACE2 deficiency occurs due to internalization of the SARS-CoV-2–ACE2 complex2,35, which restricts ACE2 from performing its physiological functions, including its role as a carboxypeptidase for angiotensin II and des-Arg9-bradykinin and other peptide hormones. ACE2 deficiency might also be created owing to its regulation at the mRNA level, such as through regulation by microRNAs. The downregulation of ACE2 protein levels by miR-200c-3p has been reported in vitro36. Since miR-200c-3p binds to the 3' UTR shared by ACE2 and dACE2, the ISG-type induction of dACE2 might serve as a decoy for binding miR-200c-3p and possibly other microRNAs and reduce the downregulation of ACE2 protein. Expression of miR-200c-3p is induced through the NF-κB pathway during infection with the pandemic flu strain H5N1 and is associated with acute respiratory distress syndrome36. Signaling through the NF-κB pathway is hyper-activated by SARS-CoV infections37, suggesting that miR-200c-3p could also be upregulated in patients with COVID-19, possibly resulting in decreased levels of ACE2 protein. In these conditions, the ISG-type induction of dACE2 mRNA could be beneficial to preserve ACE2 protein levels. It will be important to examine this potential cross-talk between the ISG-type induction of dACE2 and its role in the regulation of ACE2 protein levels, especially in COVID-19 conditions.
Patients with cancer are considered to be at a higher risk of more severe COVID-19 outcomes compared to the general population38,39 owing to older age, comorbidities and the effects of cancer and cancer treatments. Patients with lung cancer are at a specifically increased risk of severe COVID-19 outcomes38. In our analysis, dACE2 expression was common in tumors and particularly enriched in lung tumors of bronchial origin (LUSC), where the proper function of ACE2 is essential for protection from virus-induced tissue damage. The possible role of dACE2 expression in COVID-19 outcomes, specifically in patients with cancer, should be further explored. ACE inhibitors (ACEIs) and angiotensin-receptor blockers (ARBs) are widely used to control hypertension and treat heart disease and chronic kidney disease10. Some concerns were raised that ACEIs and ARBs could induce ACE2 expression, leading to increased SARS-CoV-2 infection and possibly accounting for COVID-19 severity and high mortality in those who are likely to use these medications—older people and patients with cardiovascular disease. We demonstrated that ACE2 expression is not inducible by IFNs, but it would be important to explore the effects of ACEIs and ARBs on dACE2 expression to properly assess this risk. The effects of other factors, such as smoking, on ACE2 and dACE2 expression should also be considered.
In conclusion, we report the discovery and functional annotation of dACE2, an IFN-inducible isoform of ACE2. The existence of two functionally distinct ACE2 isoforms reconciles several biological properties previously attributed to ACE2, with dACE2 being an ISG, and ACE2 acting as the SARS-CoV-2 entry receptor and carboxypeptidase, without being regulated by IFNs. While our understanding of the functional role of dACE2, a novel ISG, is still unfolding, we believe these insights will clarify our knowledge on ACE2 and provide new research leads in understanding COVID-19 susceptibility, mechanisms and outcomes.
Methods
Cells
All cell lines and primary cells used are listed in Supplementary Table 3. Cell lines were either used within 6 months after purchase or were periodically authenticated by microsatellite fingerprinting (AmpFLSTR Identifiler, Thermo Fisher) by the Cancer Genomics Research Laboratory/NCI. All cell lines were regularly tested for mycoplasma contamination using the MycoAlert Mycoplasma Detection kit (Lonza). The previously described21 NHBE cells were isolated from normal lungs that were not used for transplantation. The lungs were obtained from de-identified donors via a tissue retrieval service (International Institute for the Advancement of Medicine, Edison, NJ) with ethical approval from the Conjoint Health Research Ethics Board of the University of Calgary and the Internal Ethics Board of the International Institute for the Advancement of Medicine. Anonymized human tissue from colon resections was obtained from the University Hospital Heidelberg, in accordance with the recommendations of the University Hospital Heidelberg and written informed consent obtained from all participants in accordance with the Declaration of Helsinki. The protocol (S-443/2017) was approved by the Ethics Commission of the University Hospital Heidelberg. Organoids were generated from these tissues, as previously described22.
Viral infections
Stocks of SeV Cantell strain were purchased from Charles River Laboratories. The cells listed in Supplementary Table 3 were infected in duplicates or triplicates with SeV (7.5 × 105 50% chicken embryo infectious dose per milliliter) for 12 h as previously described18,19,20. SARS-CoV-2 (strain BavPat1) was obtained from C. Drosten at the Charité in Berlin, Germany, and provided via the European Virology Archive.
Infections with SARS-CoV-2 were performed with a multiplicity of infection of 1 in cell lines and 3 × 105 focus-forming units of the virus in organoids on the basis of titers in Vero E6 cells. Infections in colon cancer cell lines (Caco-2 and T84), a lung cancer cell line (Calu3) and colon and ileum organoid cultures were previously described22. Lung cancer cells A549 (wild-type or stably overexpressing human ACE2 (ACE2-stable)) were seeded either in 48-well plates (for RNA) or on iBIDI glass-bottom 8-well chamber slides (for immunofluorescence analysis) at a density of 7.5 × 104 cells per well or chamber. Cells were transduced at 24 h post-seeding with lentiviruses expressing GFP or dACE2–Myc and infected 3 days post-transduction. Culture medium was removed and the virus was added to cells for 1 h at 37 °C. After virus removal, cells were washed 1× with PBS, and medium was added back to the cells. Cells were collected at 24 h post-infection for RNA extraction or were fixed in 4% paraformaldehyde for 20 min at room temperature for infectivity analysis by immunofluorescence staining, as was previously described22. Briefly, cells were washed and permeabilized in 0.5% Triton-X for 15 min at room temperature. A mouse monoclonal antibody against SARS-CoV-2 nucleoprotein (Sino Biologicals) was diluted in PBS and incubated for 1 h at room temperature. Cells were washed in 1× PBS three times and incubated with goat anti–mouse Alexa Fluor 568 (Molecular Probes) and DAPI for 45 min at room temperature. Cells were washed in 1× PBS three times and imaged by epifluorescence on a Nikon Eclipse Ti-S (Nikon) to quantify the number of infected cells relative to the number of nuclei. It was determined that infection rates were 80% in Caco-2 cells, ~20% in T84 cells and ~10% in Calu3 cells and organoids22. Organoids from three donors were infected in three or four biological replicates that were averaged and presented as one value per donor.
PCR, cloning and Sanger sequencing
Complementary DNA was synthesized from 250 ng of total RNA per 20 µl reaction using the RT2 First Strand cDNA kit and random hexamers (Qiagen). PCR for the full-length dACE2 was performed using the primers and conditions listed in Supplementary Table 4. PCR-amplified products were resolved on 1% agarose gel, cut, purified and Sanger-sequenced. After the cDNA sequences were validated, constructs for dACE2 with C-terminal Myc–DDK and GFP tags cloned in the pcDNA3.4 vector were custom-synthesized by Thermo Fisher. ACE2 with a C-terminal GFP tag (RG208442) and a Myc–DDK tag (RC208442) were purchased from Origene. The empty vectors pMax-GFP (Lonza) and pCMV6-AC-Myc-DDK (Origene) were used as controls.
Treatments with IFNs
All IFNs used are listed in Supplementary Table 5. IFN treatment of NHBE cells was previously described21. Briefly, cells were cultured in BEGM with supplements (Lonza), seeded in 6-well plates and utilized at ~70% confluency (typically after 10–11 days with a medium change every 2 days). Cells were left untreated or treated with IFN-α2b (INTRON A, Merck, 100 IU ml−1) or IFN-λ3 (R&D Systems, 100 ng ml−1) for 24 h. Cells were washed with PBS, resuspended in TRIzol (Thermo Fisher) and stored at −80 °C for future RNA isolation. IFN treatment of human colon and ileum organoids was previously described22. Briefly, at ~70% of cell confluence, the medium was replaced with a cocktail of IFN-λ1–3 (100 ng ml−1 of each for a total of 300 ng ml−1) for 24 h. T84 and Caco-2 cells were treated with IFN-γ (2 ng ml−1) for 24 h.
Quantitative RT–PCR
Total RNA was extracted using the RNAeasy kit (Qiagen) from all samples except for NHBEs, for which the Direct-zol mini RNA isolation kit was used (Zymo Research). cDNA was synthesized from the total RNA with the RT2 First Strand kit (Qiagen, for all cell lines except for SARS-CoV-2-infected cells), Superscript VILO IV (Thermo Fisher, for NHBEs) or the iSCRIPT cDNA kit (Bio-Rad, for organoids and SARS-CoV-2-infected cell lines), always with an additional DNase I treatment step. Quantitative RT–PCR assays were performed in technical duplicates in 96- or 384-well plates on a QuantStudio 7 (Life Technologies) or Bio-Rad CFX 96 instrument, with RT2 SYBR Green (Qiagen), POWER SYBR (Thermo Fisher), iTaq SYBR (Bio-Rad) or TaqMan (Thermo Fisher) expression assays (Supplementary Table 4). The expression of target genes was normalized by geometric means of endogenous controls (GAPDH, HPRT1, TBP or ACTB, as indicated in Supplementary Table 2a), and is presented as dCt values relative to endogenous controls (log2 scale). For cell lines, the analyses were based on biological replicates for samples obtained from donors (NHBEs and organoids), and 3–4 biological replicates were averaged and presented per donor.
RNA sequencing (RNA-seq) of T47D and RT-4 cells
Total RNA was extracted from T47D and RT-4 cells by using the RNeasy Mini kit with an on-column DNase digestion (Qiagen) and treated with Ribo-Zero (Illumina). RNA-seq libraries were prepared from high-quality RNA samples (RIN scores >9.0) with the KAPA Stranded RNA-seq kit with RiboErase (Roche). Paired 150-bp reads (21.2–118.8 million reads per sample) were generated with HiSeq 2500 (Illumina) by the Cancer Genomics Laboratory (Division of Cancer Epidemiology and Genetics, National Cancer Institute (DCEG/NCI)). The reads were aligned with STAR alignment tool version 7.1.2a (21) using the GRChg37/hg19 genome assembly and visualized using the Integrative Genomics Viewer. The RNA-seq dataset of a breast cancer cell line, T47D, infected with SeV was deposited as NCBI SRA: PRJNA512015.
RNA-seq analysis of data from NCBI SRA and TCGA
RNA-seq datasets (Supplementary Table 6) were downloaded from NCBI SRA using SRA tools. The FASTQ files were compressed using GZIP and aligned with STAR version 7.1.3a to the human GRChg38/hg38 genome assembly. BAM files with ≤80% of mappable reads were excluded. BAM files were indexed and sliced using SAMtools to include 51.6 kilobases of the human ACE2 genomic region (chrX: 15,556,393–15,608,016, hg38). For non-human RNA-seq datasets, the alignment was performed with the reference genomes mm10 for mice, and MusPutFur1.0 for ferrets. For TCGA STAR-aligned RNA-seq data, BAM slices for the ACE2 region were acquired for 10,898 TCGA samples (10,185 tumors and 713 tumor-adjacent normal tissues) through the NCI Genomics Data Commons portal accessed on 12 May 2020, using workflow https://docs.gdc.cancer.gov/API/Users_Guide/BAM_Slicing/.
Estimation of RNA-seq read counts for ACE2 exons
RNA-seq reads corresponding to ACE2-Ex1a, ACE2-Ex1b and dACE2-Ex1c were counted by processing RNA-seq BAM slices using the R package ASpli version 1.5.1 with default settings. Genomic coordinates were manually curated in the GTF file and the ‘counts’ function was used to generate and export RNA-seq reads for the selected exons in a tab file format. The analysis of exon expression patterns within tissue subtypes was based on log2-transformed raw reads for each exon. The reads were normalized by dividing by the exon length (Supplementary Table 7) and multiplying by the geometric mean of the total reads of the three exons (Ex1a, Ex1b and Ex1c) across all samples as a scaling factor to adjust for variability in sequencing coverage between samples. Correlation analyses of log2[normalized exon expression + 1] of ACE2-Ex1a, ACE2-Ex1b and dACE2-Ex1c with log2[transcripts per million + 1] of IFNs, genes encoding signal transducers and activators of transcription (STATs), interferon regulatory factors (IRFs) and select ISG controls (ISG15 and ISG20) were performed in R using the package dplyr.
Expression values for all IFN genes for all tumors in the TCGA PanCancer Atlas were downloaded as RSEM values from the cBioPortal for Cancer Genomics (https://www.cbioportal.org/). Expression of IFNL4, a most recently discovered IFN (ref. 40), was not available in the TCGA dataset based on the hg19/GRCh37 reference. As IFNG was expressed in most samples compared to other IFN genes (Extended Data Fig. 5a), it was used for further analysis. Correlation analyses of the log2[normalized exon expression + 1] of ACE2-Ex1b and dACE2-Ex1c were performed with log2[RSEM + 1] values for IFNG. Correlation patterns between IFNG and ACE2-Ex1a were similar to those between IFNG and ACE2-Ex1b (data not shown). Correlation analyses were performed with Spearman and Pearson methods and provided similar results. The P values and coefficients presented are for Pearson correlations.
Unsupervised clustering and correlation analyses in TCGA
Gene expression Z-scores in the lung squamous cell carcinoma (TCGA-LUSC, n = 501) dataset were calculated for 270 ISGs from a previously curated list23. ISGs with low expression values (below 10 reads) or expressed in less than 5% of tumors were excluded. The data were used for self-organizing map (SOM) clustering, which is an unsupervised machine learning approach enabling data dimensionality reduction without relying on any assumption about the data structure41,42. The SOM algorithm was iterated 100,000 times with Euclidean distance, linearly decreasing the learning rate from 0.05 to 0.01 using the R package Kohonen. The ISG expression patterns were projected onto a two-dimensional 10 × 10 hexagonal map. Thus, each node in this map is an expression profile representing a subset of the samples. SOM output, trained on the basis of 100,000 iterations, was used to estimate the contribution of each ISG to defining the clusters as a variance weighted according to the size of each node. A total of six clusters were estimated by the kmeans algorithm and used to generate an expression heatmap. Expression Z-scores of the top 100 ISGs ranked on the basis of their contribution to defining the clusters were plotted using the R package pheatmap. Pearson correlation coefficients and corresponding FDR-adjusted P values were calculated between Z-scores for the top 100 ISGs and both ACE2 and dACE2 in cluster 5, which included 114 tumors with the highest ISG expression. The analysis was performed using the R package Hmisc.
In silico analysis of promoter regulatory elements relevant for IFN signaling
Promoters were defined within the −800 bp/+100 bp window from the corresponding TSSs. The window was limited by 800 bp, based on the intronic distance between the TSS of Ex1c and its upstream exon. Promoters of ACE2-Ex1a (P1), ACE2-Ex1b (P2) and dACE2-Ex1c (P3) were analyzed using Nsite tool43 from the online bioinformatics gateway Softberry (http://www.softberry.com) to predict transcription-factor-binding sites. The search was set against the Object-oriented Transcription Factors Database of human and animal transcription-factor-binding sites largely curated according to the functional data from the literature. The parameters were set to allow a maximum of 1 or 2 mismatches with the known motifs. ISG-type motifs were manually curated from ~300 predicted and annotated motifs per promoter.
Luciferase promoter assays
ACE2-P1, ACE2-P2 and dACE2-P3 promoters (here defined as −800/−1 bp from the corresponding TSS), and two variants of the dACE2-P3 promoter with 100-bp deletions harboring predicted ISG-type motifs (Fig. 2b), were custom-synthesized by Thermo Fisher and cloned upstream of the luciferase reporter in a promoterless vector pGL4.21 (Promega) using the Xho1 and HindIII restriction sites. HepG2 cells were seeded in 96-well plates (4 × 103 cells per well) and 24 h after plating transiently co-transfected with the corresponding luciferase constructs together with a normalization control (Renilla pGL4.74 plasmid, Promega, in a 10:1 ratio), in six biological replicates per construct. The medium was changed 6 h after transfection and cells were treated with IFN-β (1 ng ml−1, R&D Systems), IFN-γ (2 ng ml−1, R&D Systems) or medium (mock) starting from 48 h post-transfection. After 6 h of treatment, cells were lysed and analyzed with the GloMax multi-detection system (Promega) and the luciferase levels in each well were normalized to the corresponding Renilla levels. The results were normalized by respective mock-treated controls and are presented as fold change over the negative control (empty pGL4.21 vector).
Mining of proteomics datasets
Mass spectrometry datasets generated for TCGA colon, breast and ovarian tumors (http://www.pepquery.org/) were mined for matches to the 36-aa fragment of dACE2, including the unique 10 aa encoded by dACE2-Ex1c. The analysis was performed with the PepQuery peptide-centric search engine24, using the following parameters: mass spectrometry dataset of a specific cancer type; target event as protein; scoring algorithm as hyperscore and not selecting for unrestricted modification filtering. The identified peptides for each cancer type were exported as CSV files and manually analyzed for further assessment of peptide quality.
Transient transfections
Transient transfections were performed with Lipofectamine 3000 (Thermo Fisher). Unless specified, T24, a bladder cancer cell line in which no baseline expression of ACE2 or dACE2 was detected (Supplementary Table 2a), was used for transfections at 70–90% confluency in 12- or 6-well plates for 24 h.
Western blot
Cells were lysed with RIPA buffer (Sigma) supplemented with protease inhibitor cocktail (Promega) and PhosSTOP (Roche) and placed on ice for 30 min, with vortexing every 10 min. Lysates were pulse-sonicated for 30 s, with 10 s burst-cooling cycles, at 4°C, boiled in reducing sample buffer for 5 min and resolved on 4–12% Bis-Tris Bolt gels and transferred using an iBlot 2 (Thermo Fisher). Blots were blocked in 2.5% milk in 1% TBS–Tween before staining with antibodies (Supplementary Table 5). The signals were detected with HyGLO Quick Spray (Denville Scientific) or SuperSignal West Femto Maximum Sensitivity Substrate (Thermo Fisher) and viewed on a ChemiDoc Touch Imager with Image Lab 5.2 software (Bio-Rad).
Cell surface biotinylation and pulldown with streptavidin beads
T24 cells were transiently transfected with dACE2–Myc, ACE2–Myc or both constructs for 24 h. Cell surface biotinylation and pulldown with streptavidin beads were carried out using the Pierce Cell Surface Biotinylation and Isolation kit (Thermo Fisher). Briefly, the cell surface of T24 cells was biotinylated using EZ-Link sulfo-NHS-SS-biotin. Cells were lysed with RIPA buffer (Sigma) supplemented with protease inhibitor cocktail (Promega) and PhosSTOP (Roche) and placed on ice for 30 min, with vortexing every 10 min. Lysates were pulse-sonicated for 30 s, with 10 s burst-cooling cycles, at 4 °C. Biotinylated proteins were isolated with Neutravidin beads supplied with the kit. Input lysates and pulldown proteins from biotin+ and biotin− fractions were analyzed by western blotting with the C-terminal ACE2 antibody (Abcam) as described above and anti-GAPDH antibody (Abcam).
Confocal microscopy
T24 cells were transiently transfected with ACE2–GFP or dACE2–Myc, or co-transfected with both constructs in 4-well chambered slides (2 × 104 cells per well, LabTek). After 24 h, cells were treated with 2 ng ml−1 of recombinant biotinylated SARS-CoV-2 spike protein RBD (spike protein RBD, Sino Biological) for 1 h at 37 °C. Cells were washed twice with medium and then stained with 5 µg ml−1 streptavidin PE (Thermo Fisher) for 30 min at 37 °C. Cells were then washed twice with PBS and fixed with 4% paraformaldehyde (BD Biosciences) for 30 min. After rinsing twice in PBS and permeabilization buffer (BD Biosciences), cells were incubated with permeabilization buffer for 1 h. Fixed cells were incubated with rabbit anti-FLAG antibody (1:250 dilution, Thermo Fisher) overnight, washed and then stained with anti-rabbit Alexa Fluor 680 (1:500 dilution, Thermo Fisher). Slides were mounted with antifade mounting media with DAPI (Thermo Fisher) and imaged at ×40 magnification on an LSM700 confocal laser scanning microscope (Carl Zeiss) using an inverted oil lens.
Flow cytometry analysis of SARS-CoV-2 spike protein RBD binding
T24 cells were transiently transfected with ACE2–GFP or dACE2–Myc, or co-transfected with both constructs in 12-well plates (1 × 104 cells per well). After 24 h, cells were stained with recombinant biotinylated spike protein RBD as described above and analyzed with multiparametric flow cytometry on a FACS Aria III (BD Biosciences) and FlowJo v10 software (BD Biosciences).
Carboxypeptidase activity assays
T24 cells were plated overnight in T-25 flasks (5 × 105 cells per flask) and transiently transfected with 10 µg of ACE2–GFP, dACE2–GFP or empty GFP vector. At 24 h post-transfection, cells were pelleted and lysed with 400 μl of the lysis buffer provided with the ACE2 activity kit (no. K897, BioVision). Keeping the reaction volumes and the amount of ACE2–GFP lysates constant, we added dACE2–GFP lysate in the ratio of 0.25, 0.5 and 1.0 to ACE2–GFP, with differences in volume compensated by lysates from GFP-expressing cells. The lysate mixtures were processed in triplicates using the kit reagents and according to the protocol. The carboxypeptidase activity was measured as fluorescence (Ex/Em = 365/410–460 nm) using a Promega GlowMax plate reader for two time points between 30 min and 2 h after adding the corresponding substrate mix. A positive control was provided by the kit. Cell lysates were also analyzed by western blots with C-terminal anti-ACE2 antibody (Abcam), which detects both ACE2 and dACE2, with GAPDH as a loading control.
Statistical analysis
Expression of ACE2-Ex1a, ACE2-Ex1b and dACE2-Ex1c between groups of samples was evaluated by two-sided tests: unpaired non-parametric Mann–Whitney U tests, paired Student’s t-tests (for NHBE cells from five donors and organoids for three donors) and unpaired Student’s t-tests (for biological replicates of cell lines and organoids from one donor). Statistical tests for other analyses are indicated in the corresponding sections. FDR adjustment was applied when indicated. P values < 0.05 were considered significant.
Computational resources
We used the NIH Biowulf supercomputing cluster and specific packages for R version 3.6.2.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The sequence for dACE2 was deposited to NCBI GenBank with the accession number MT505392. The RNA-seq dataset of a breast cancer cell line, T47D, infected with SeV was deposited as NCBI SRA: PRJNA512015. The accession numbers for all of the data used in the paper are presented in Supplementary Table 6. Source data are provided with this paper. Requests for any additional data or reagents should be addressed to L.P.-O. (prokuninal@mail.nih.gov).
References
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Li, W. et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 426, 450–454 (2003).
Lukassen, S. et al. SARS-CoV-2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells. EMBO J. 39, e105114 (2020).
Ziegler, C. G. K. et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell 181, 1016–1035 (2020).
Chua, R. L. et al. COVID-19 severity correlates with airway epithelium–immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 38, 970–979 (2020).
Zhou, Q. et al. Interferon-α2b treatment for COVID-19. Front. Immunol. 11, 1061 (2020).
Hung, I. F. et al. Triple combination of interferon beta-1b, lopinavir-ritonavir, and ribavirin in the treatment of patients admitted to hospital with COVID-19: an open-label, randomised, phase 2 trial. Lancet 395, 1695–1704 (2020).
Prokunina-Olsson, L. et al. COVID-19 and emerging viral infections: the case for interferon lambda. J. Exp. Med. 217 (2020).
O’Brien, T. R. et al. Weak induction of interferon expression by SARS-CoV-2 supports clinical trials of interferon lambda to treat early COVID-19. Clin. Infect. Dis. 71, 1410–1412 (2020).
Re, R. N. Mechanisms of disease: local renin-angiotensin-aldosterone systems and the pathogenesis and treatment of cardiovascular disease. Nat. Clin. Pract. Cardiovasc. Med. 1, 42–47 (2004).
Diamond, B. The renin-angiotensin system: an integrated view of lung disease and coagulopathy in COVID-19 and therapeutic implications. J. Exp. Med. 217, jem.20201000 (2020).
van de Veerdonk, F. L. et al. Kallikrein–kinin blockade in patients with COVID-19 to prevent acute respiratory distress syndrome. eLife 9, e57555 (2020).
de Maat, S., de Mast, Q., Danser, A. H. J., van de Veerdonk, F. L. & Maas, C. Impaired breakdown of bradykinin and its metabolites as a possible cause for pulmonary edema in COVID-19 infection. Semin. Thromb. Hemost. https://doi.org/10.1055/s-0040-1712960 (2020).
Imai, Y. et al. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature 436, 112–116 (2005).
Kuba, K. et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat. Med. 11, 875–879 (2005).
Huang, F. et al. Angiotensin II plasma levels are linked to disease severity and predict fatal outcomes in H7N9-infected patients. Nat. Commun. 5, 3595 (2014).
Zou, Z. et al. Angiotensin-converting enzyme 2 protects from lethal avian influenza A H5N1 infections. Nat. Commun. 5, 3594 (2014).
Middlebrooks, C. D. et al. Association of germline variants in the APOBEC3 region with cancer risk and enrichment with APOBEC-signature mutations in tumors. Nat. Genet. 48, 1330–1338 (2016).
Obajemu, A. A. et al. IFN-lambda4 attenuates antiviral responses by enhancing negative regulation of IFN signaling. J. Immunol. 199, 3808–3820 (2017).
Minas, T. Z. et al. IFNL4-ΔG is associated with prostate cancer among men at increased risk of sexually transmitted infections. Commun. Biol. 1, 191 (2018).
Santer, D. M. et al. Differential expression of interferon-lambda receptor 1 splice variants determines the magnitude of the antiviral response induced by interferon-lambda 3 in human immune cells. PLoS Pathog. 16, e1008515 (2020).
Stanifer, M. L. et al. Critical role of type III interferon in controlling SARS-CoV-2 infection in human intestinal epithelial cells. Cell Rep. 32, 107863 (2020).
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Wen, B., Wang, X. & Zhang, B. PepQuery enables fast, accurate, and convenient proteomic validation of novel genomic alterations. Genome Res. 29, 485–493 (2019).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 (2020).
Chu, H. et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. 71, 1400–1409 (2020).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. 5, eabd.1554 (2020).
Broggi, A. et al. Type III interferons disrupt the lung epithelial barrier upon viral recognition. Science 369, 706–712 (2020).
Major, J. et al. Type I and III interferons disrupt lung epithelial repair during recovery from viral infection. Science 369, 712–717 (2020).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Lagunas-Rangel, F. A. & Chavez-Valencia, V. High IL-6/IFN-gamma ratio could be associated with severe disease in COVID-19 patients. J. Med. Virol. 92, jmv.25900 (2020).
Thijsen, S. et al. Elevated nucleoprotein-induced interferon-gamma release in COVID-19 patients detected in a SARS-CoV-2 enzyme-linked immunosorbent spot assay. J. Infect. 81, 452–482 (2020).
Leppek, K., Das, R. & Barna, M. Functional 5' UTR mRNA structures in eukaryotic translation regulation and how to find them. Nat. Rev. Mol. Cell Biol. 19, 158–174 (2018).
Zores, F. & Rebeaud, M. E. COVID and the renin–angiotensin system: are hypertension or its treatments deleterious? Front. Cardiovasc. Med. 7, 71 (2020).
Wan, Y., Shang, J., Graham, R., Baric, R. S. & Li, F. Receptor recognition by the novel coronavirus from Wuhan: an analysis based on decade-long structural studies of SARS coronavirus. J. Virol. 94, e00127-20 (2020).
Liu, Q. et al. miRNA-200c-3p is crucial in acute respiratory distress syndrome. Cell Discov. 3, 17021 (2017).
Hirano, T. & Murakami, M. COVID-19: a new virus, but a familiar receptor and cytokine release syndrome. Immunity 52, 731–733 (2020).
Mehta, V. et al. Case fatality rate of cancer patients with COVID-19 in a New York hospital system. Cancer Discov. 10, 935–941 (2020).
Kuderer, N. M. et al. Clinical impact of COVID-19 on patients with cancer (CCC19): a cohort study. Lancet 395, 1907–1918 (2020).
Prokunina-Olsson, L. et al. A variant upstream of IFNL3 (IL28B) creating a new interferon gene IFNL4 is associated with impaired clearance of hepatitis C virus. Nat. Genet. 45, 164–171 (2013).
Nikkila, J. et al. Analysis and visualization of gene expression data using self-organizing maps. Neural Netw. 15, 953–966 (2002).
Vesanto, J. & Alhoniemi, E. Clustering of the self-organizing map. IEEE Trans. Neural Netw. 11, 586–600 (2000).
Shahmuradov, I. A. & Solovyev, V. V. Nsite, NsiteH and NsiteM computer tools for studying transcription regulatory elements. Bioinformatics 31, 3544–3545 (2015).
Acknowledgements
We thank N. Cole (DCEG/NCI) for help with the acquisition of TCGA sliced BAM files and D. Proud (University of Calgary) for providing NHBE cells. We thank the CGR/DCEG/NCI for help with RNA-seq and authentication of cell lines by Identifiler profiling. The results are partially based on data generated by the TCGA Research Network. The project was supported by the Intramural Research Program of the DCEG/NCI; the NIH grant R21AG064479-01 and a Brain Health Research Institute Pilot Award from Kent State University (H.P.); S.B. and M.L.S. received financial support from the Deutsche Forschungsgemeinschaft: project numbers 240245660 (SFB1129) 415089553 (Heisenberg) and 272983813 (TRR179) to S.B. and project number 416072091 to M.L.S. D.M.S. and D.L.T. received financial support from the Li Ka Shing Institute of Virology.
Author information
Authors and Affiliations
Contributions
O.O.O., A.R.B. and L.P.-O. conceived and designed the study. L.P.-O. supervised the study. O.O.O., A.R.B., M.L.S., D.M.S., S.B. and L.P.-O. designed the experiments. O.O.O., A.R.B., M.L.S., D.M.S., W.Y., A.O., J.M.V., T.J.R., C.K. and P.D. performed the experiments and analyzed the data. J.L.M. analyzed structural features of isoforms. A.R.B. performed computational analyses and quantification of RNA-seq datasets. A.R.B., O.O.O., H.P. and O.F.-V. performed statistical analyses. O.O.O., A.R.B., M.L.S., S.B., D.M.S., D.L.T. and L.P.-O. contributed reagents/materials/analysis tools. O.O.O., A.R.B. and L.P.-O. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks David Levy, Carly Ziegler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ACE2 expression patterns in the mouse and human lung cells infected with the respiratory syncytial virus (RSV) and nasal washes from ferrets infected with SARS-CoV-2.
a, Sashimi plots of the Ace2 region in a lung RNA-seq dataset from mice mock/RSV- infected in triplicates. The expression of dAce2-Ex1c is not observed and Ace2 expression is not affected by the infection. b, Sashimi plots of the ACE2 region in H292, a human lung mucoepidermoid pulmonary carcinoma cell line. Expression of ACE2 from Ex1a and Ex1b is very low, while the dACE2 from Ex1c is expressed at baseline and is further induced by RSV infection. c, Representative IGV plots showing exon and exon-exon junction RNA-seq reads from nasal washes of mock/SARS-CoV-2-infected ferrets. The expression of dAce2-Ex1c is not observed. Note: The mouse and human/ferret ACE2/Ace2 genes are shown in opposite orientations, as presented in IGV. The positions corresponding to the human dACE2-Ex1c are marked in the mouse and ferret genomes. Datasets: PRJNA588982, PRJNA61503.
Extended Data Fig. 2 ACE2 and dACE2 expression in normal human tissues.
RNA-seq read counts for ACE2-Ex1a, ACE2-Ex1b, and dACE2-Ex1c in 27 human tissues. Dataset: PRJEB4337, n = 95.
Extended Data Fig. 3 ACE2 and dACE2 expression in tumor-adjacent normal tissues in TCGA.
Based on RNA-seq read counts, ACE2-Ex1b is detectable in multiple samples of several tissue types. dACE2-Ex1c expression is more restricted and most common in normal tissue adjacent to tumors of head and neck (HNSC), stomach (STAD), lung squamous carcinoma (LUSC), colon (COAD), and esophagus (ESCA).
Extended Data Fig. 4 ACE2 and dACE2 expression across 10,185 tumors of 33 types in TCGA.
Based on RNA-seq read counts, ACE2-Ex1b is most expressed in kidney tumors—kidney renal clear cell carcinoma (KIRC) and kidney renal papillary cell carcinoma (KIRP). Most samples expressing dACE2-Ex1c are squamous tumors of head and neck (HNSC) and the lungs (LUSC). Based on a ≥ 5 reads/sample threshold, ACE2-Ex1a is expressed in 12.6%, ACE2-Ex1b – in 38.0% and dACE2-Ex1c—in 16.8% of all tumors.
Extended Data Fig. 5 dACE2 and ACE2 expression in relation to IFNG expression in TCGA tumors and in cells treated with IFNγ.
a, Expression levels of all IFN genes annotated in TCGA tumors (n = 10,185) were acquired from cBioPortal (https://www.cbioportal.org/); expression of IFNL4 was not annotated in this dataset. At RSEM ≥ 1, only expression of IFNG is common (61% samples), with mean RSEM = 19.8 compared to other IFN genes (mean RSEM ≤ 1.3). b, and c, Pearson correlation coefficients (r) for dACE2 and ACE2 vs. IFNG expression across tumors. dACE2 showed significant positive correlations (r ≥ 0.2) with IFNG in 8 tumor types, while ACE2 showed mainly negative correlations and only one positive correlation in breast cancer (r = 0.15). Expression values for dACE2 and ACE2 were based on log2-normalized exon read counts (Ex1b and Ex1c) and for IFNG—on RSEM values. d, Treatment of cell lines with IFNγ (2 ng/ml for 24 hrs) induced expression of dACE2 but not ACE2 in T84 cells. The results are presented with means + /- SD for three biological replicates; P-values are for the unpaired, two-sided Student’s T-tests.
Extended Data Fig. 6 Unsupervised self-organizing map (SOM) analysis in TCGA-LUSC tumors.
a, Construction of the unsupervised SOM of TCGA-LUSC tumors (n = 501) based on Z-scores calculated for each of the 270 curated ISGs. Each hexagon includes a mean of 5 (range 1-14) tumors with similar ISG expression profiles. Colors denote clusters (1-6) of tumors with similar ISG expression profiles. b, Heatmap of the six SOM-defined clusters, visualized by plotting the expression levels of top 100 ISGs selected by the ranking of the initial set of 270 ISGs based on their contribution to these clusters. Cluster 5 includes 114 tumors with the highest ISG expression, whereas cluster 3 includes 72 tumors with the lowest ISG expression. c, and d, Volcano plots showing FDR-adjusted p-values and Pearson correlation coefficients (r), for two-sided tests, for expression of dACE2 and ACE2 in relation to the expression of the top 100 ISGs within cluster 5. In total, dACE2 was significantly (FDR p-value < 0.05) correlated with the expression of 20 ISGs and ACE2—with 5 ISGs.
Extended Data Fig. 7 Peptides encoded by dACE2-Ex1c are detected by protein sequencing in tumors.
a, Results of peptide query in PepQuery2 proteomics database of mass-spec data in 174 ovarian, 95 colon, and 105 breast tumors in TCGA7. Three peptides – MREAGWDK, EAGWDKGGR, and GGRILMCTK uniquely correspond to the 10 aa encoded by dACE2-Ex1c. The latter peptide results from the splicing of dACE2-Ex1c with its downstream exon. The total number of identified peptides, the number of samples with specific peptides, and corresponding parameters for a peptide-spectrum match (PSM) are shown in table format. b, Representative spectra of two peptides matching with the protein encoded by dACE2-Ex1c. M/z refers to the mass by charge ratio. The b-series and y-series ions showed the correct mapping of residues in the query aa sequence.
Extended Data Fig. 8 Transiently overexpressed ACE2-GFP and dACE2-GFP are detectable on the cell surface.
T24 cells were transiently transfected for 24 hrs with indicated constructs or not transfected (control) and then treated with (+) or without (-) biotin. Western blots with the C-terminal anti-ACE2 antibody detected a similar expression of both proteins in input lysates, but only in the biotin+ fraction after pull-down with streptavidin beads, showing much stronger expression of ACE2 compared to dACE2. GAPDH, a cytoplasmic protein, is undetectable in the pull-down fraction. The experiment was repeated three times, and representative results are shown.
Extended Data Fig. 9 ACE2-GFP levels are non-specifically increased in cells co-transfected with dACE2-Myc or TMEM129-Myc.
a, A representative flow cytometry plot showing gates for the T24 cells co-transfected with dACE2-Myc and ACE2-GFP, corresponding to Fig. 7d, and e. The gates were drawn to identify cells expressing dACE2-Myc, ACE2-GFP, or both proteins. b, and c, Mean fluorescence intensity (MFI) of ACE2-GFP expression in the T24 cells transiently co-transfected in triplicates with ACE2-GFP and dACE2-Myc b, or ACE2-GFP and TMEM129-Myc c, and gated as described in a. TMEM129 is a transmembrane protein, which serves as an independent control, showing that the increase in ACE2-GFP expression might be a non-specific effect due to transfection rather than due to dACE2 co-expression.
Extended Data Fig. 10 Immunofluorescence analysis of SARS-CoV-2 infection in the A549 cells.
a, and b, Quantification of dACE2 expression a, and ACE2 expression. b, in SARS-CoV-2 (MOI = 1) or mock-infected lung cancer cell line A549 transfected with GFP (used as a transient transfection control), dACE2-Myc, or stably expressing ACE2 (ACE2-stable cell line) and transfected with GFP. The results are presented with means for three biological replicates. c-h) Immunofluorescence images in cells corresponding to plots a, and b. Cells were fixed and stained 72 hours after infection—SARS-CoV-2 nuclear protein (white), nuclei—DAPI (blue). Representative images from one of three biological replicates are shown. Corresponding plots for viral load and % of infected cells are presented in Fig. 7f and g. Scale bars, 100 µM.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 3–7, and Figs. 1 and 2
Supplementary Table 2
a, Expression in cell lines, baseline and SeV infection (Fig. 3c). b, Expression in NHBE cells from five donors, IFN treatments (Fig. 3d), and in Vero E6 and HEK293T cells. c, Expression in organoid cultures from one donor, IFN treatments in biological replicates (Fig. 3e). d, Expression in cell lines, SARS-CoV-2 infection, in biological replicates (Fig. 6a,b). e, Expression in organoid cultures from three donors, SARS-CoV-2, infection in biological replicates (Fig. 6c,d). f, De novo quantification of ACE2 and dACE2 expression in all TCGA samples based on RNA-seq data. g, Expression in TCGA tumors. h, Expression in TCGA tumor-adjacent normal tissues.
Source data
Source Data Fig. 1
Unprocessed image of the agarose gel for Fig. 3b.
Source Data Fig. 2
Unprocessed western blots for Fig. 7h.
Source Data Extended Data Fig. 8
Unprocessed western blots for Extended Data Fig. 8.
Rights and permissions
About this article
Cite this article
Onabajo, O.O., Banday, A.R., Stanifer, M.L. et al. Interferons and viruses induce a novel truncated ACE2 isoform and not the full-length SARS-CoV-2 receptor. Nat Genet 52, 1283–1293 (2020). https://doi.org/10.1038/s41588-020-00731-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-00731-9
This article is cited by
-
ACE2 in chronic disease and COVID-19: gene regulation and post-translational modification
Journal of Biomedical Science (2023)
-
Olfactory immune response to SARS-CoV-2
Cellular & Molecular Immunology (2023)
-
Virologic Studies in COVID-Positive Donors
Current Transplantation Reports (2023)
-
Pharmacological disruption of mSWI/SNF complex activity restricts SARS-CoV-2 infection
Nature Genetics (2023)
-
Mouse genome rewriting and tailoring of three important disease loci
Nature (2023)