Abstract
Increased production of fetal hemoglobin (HbF) can ameliorate the severity of sickle cell disease and β-thalassemia1. BCL11A represses the genes encoding HbF and regulates human hemoglobin switching through variation in its expression during development2,3,4,5,6,7. However, the mechanisms underlying the developmental expression of BCL11A remain mysterious. Here we show that BCL11A is regulated at the level of messenger RNA (mRNA) translation during human hematopoietic development. Despite decreased BCL11A protein synthesis earlier in development, BCL11A mRNA continues to be associated with ribosomes. Through unbiased genomic and proteomic analyses, we demonstrate that the RNA-binding protein LIN28B, which is developmentally expressed in a pattern reciprocal to that of BCL11A, directly interacts with ribosomes and BCL11A mRNA. Furthermore, we show that BCL11A mRNA translation is suppressed by LIN28B through direct interactions, independently of its role in regulating let-7 microRNAs, and that BCL11A is the major target of LIN28B-mediated HbF induction. Our results reveal a previously unappreciated mechanism underlying human hemoglobin switching that illuminates new therapeutic opportunities.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The massively parallel sequencing data associated with this manuscript are available in the Gene Expression Omnibus database under accession code GSE118359.
The original mass spectra and the protein sequence database used for searches have been deposited in the public proteomics repository MassIVE (http://massive.ucsd.edu) and are accessible at accession MSV000084443. Source data for Figs. 1–4 and Extended Data Figs. 1, 3–5 and 10 are available online.
Code availability
Custom computer code for reproduction of sequencing-based analyses is available at https://github.com/sankaranlab/translation-regulation-bcl11a.
References
Sankaran, V. G. & Orkin, S. H. The switch from fetal to adult hemoglobin. Cold Spring Harb. Perspect. Med. 3, a011643 (2013).
Sankaran, V. G. et al. Human fetal hemoglobin expression is regulated by the developmental stage-specific repressor BCL11A. Science 322, 1839–1842 (2008).
Sankaran, V. G. et al. Developmental and species-divergent globin switching are driven by BCL11A. Nature 460, 1093–1097 (2009).
Basak, A. et al. BCL11A deletions result in fetal hemoglobin persistence and neurodevelopmental alterations. J. Clin. Invest. 125, 2363–2368 (2015).
Liu, N. et al. Direct promoter repression by BCL11A controls the fetal to adult hemoglobin switch. Cell 173, 430–442 e17 (2018).
Xu, J. et al. Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6. Genes Dev. 24, 783–798 (2010).
Martyn, G. E. et al. Natural regulatory mutations elevate the fetal globin gene via disruption of BCL11A or ZBTB7A binding. Nat. Genet. 50, 498–503 (2018).
Sankaran, V. G. & Weiss, M. J. Anemia: progress in molecular mechanisms and therapies. Nat. Med. 21, 221–230 (2015).
Menzel, S. et al. A QTL influencing F cell production maps to a gene encoding a zinc-finger protein on chromosome 2p15. Nat. Genet. 39, 1197–1199 (2007).
Uda, M. et al. Genome-wide association study shows BCL11A associated with persistent fetal hemoglobin and amelioration of the phenotype of beta-thalassemia. Proc. Natl Acad. Sci. USA 105, 1620–1625 (2008).
Yen, H. C., Xu, Q., Chou, D. M., Zhao, Z. & Elledge, S. J. Global protein stability profiling in mammalian cells. Science 322, 918–923 (2008).
Ludwig, L. S. et al. Altered translation of GATA1 in Diamond–Blackfan anemia. Nat. Med. 20, 748–753 (2014).
Dieterich, D. C., Link, A. J., Graumann, J., Tirrell, D. A. & Schuman, E. M. Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc. Natl Acad. Sci. USA 103, 9482–9487 (2006).
Sankaran, V. G. et al. Exome sequencing identifies GATA1 mutations resulting in Diamond–Blackfan anemia. J. Clin. Invest. 122, 2439–2443 (2012).
Khajuria, R. K. et al. Ribosome levels selectively regulate translation and lineage commitment in human hematopoiesis. Cell 173, 90–103 e19 (2018).
Darnell, J. C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Ingolia, N. T. Ribosome footprint profiling of translation throughout the genome. Cell 165, 22–33 (2016).
Das Sharma, S. et al. Widespread alterations in translation elongation in the brain of juvenile Fmr1 knockout mice. Cell Rep. 26, 3313–3322 e5 (2019).
Richter, J. D. & Coller, J. Pausing on polyribosomes: make way for elongation in translational control. Cell 163, 292–300 (2015).
Chen, E., Sharma, M. R., Shi, X., Agrawal, R. K. & Joseph, S. Fragile X mental retardation protein regulates translation by binding directly to the ribosome. Mol. Cell 54, 407–417 (2014).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Munschauer, M. et al. The NORAD lncRNA assembles a topoisomerase complex critical for genome stability. Nature 561, 132–136 (2018).
Graf, R. et al. Identification of LIN28B-bound mRNAs reveals features of target recognition and regulation. RNA Biol. 10, 1146–1159 (2013).
Lee, Y. T. et al. LIN28B-mediated expression of fetal hemoglobin and production of fetal-like erythrocytes from adult human erythroblasts ex vivo. Blood 122, 1034–1041 (2013).
Yuan, J., Nguyen, C. K., Liu, X., Kanellopoulou, C. & Muljo, S. A. Lin28b reprograms adult bone marrow hematopoietic progenitors to mediate fetal-like lymphopoiesis. Science 335, 1195–1200 (2012).
Rowe, R. G. et al. Developmental regulation of myeloerythroid progenitor function by the Lin28b–let-7–Hmga2 axis. J. Exp. Med. 213, 1497–1512 (2016).
Copley, M. R. et al. The Lin28b–let-7–Hmga2 axis determines the higher self-renewal potential of fetal haematopoietic stem cells. Nat. Cell Biol. 15, 916–925 (2013).
Bronevetsky, Y., Burt, T. D. & McCune, J. M. Lin28b regulates fetal regulatory T cell differentiation through modulation of TGF-beta signaling. J. Immunol. 197, 4344–4350 (2016).
Newman, M. A., Thomson, J. M. & Hammond, S. M. Lin-28 interaction with the Let-7 precursor loop mediates regulated microRNA processing. RNA 14, 1539–1549 (2008).
Viswanathan, S. R., Daley, G. Q. & Gregory, R. I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008).
Wilbert, M. L. et al. LIN28 binds messenger RNAs at GGAGA motifs and regulates splicing factor abundance. Mol. Cell 48, 195–206 (2012).
Hafner, M. et al. Identification of mRNAs bound and regulated by human LIN28 proteins and molecular requirements for RNA recognition. RNA 19, 613–626 (2013).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Cho, J. et al. LIN28A is a suppressor of ER-associated translation in embryonic stem cells. Cell 151, 765–777 (2012).
Nam, Y., Chen, C., Gregory, R. I., Chou, J. J. & Sliz, P. Molecular basis for interaction of let-7 microRNAs with Lin28. Cell 147, 1080–1091 (2011).
Giani, F. C. et al. Targeted application of human genetic variation can improve red blood cell production from stem cells. Cell Stem Cell 18, 73–78 (2016).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Pimentel, H., Bray, N. L., Puente, S., Melsted, P. & Pachter, L. Differential analysis of RNA-seq incorporating quantification uncertainty. Nat. Methods 14, 687–690 (2017).
Hahne, F. & Ivanek, R. Visualizing genomic data using Gviz and Bioconductor. Methods Mol. Biol. 1418, 335–351 (2016).
Ulirsch, J. C. et al. Altered chromatin occupancy of master regulators underlies evolutionary divergence in the transcriptional landscape of erythroid differentiation. PLoS Genet. 10, e1004890 (2014).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Xue, S. et al. RNA regulons in Hox 5′ UTRs confer ribosome specificity to gene regulation. Nature 517, 33–38 (2015).
Koontz, L. TCA precipitation. Methods Enzymol. 541, 3–10 (2014).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, W., Wang, W., Uren, P. J., Penalva, L. O. F. & Smith, A. D. Riborex: fast and flexible identification of differential translation from Ribo-seq data. Bioinformatics 33, 1735–1737 (2017).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Klass, D. M. et al. Quantitative proteomic analysis reveals concurrent RNA-protein interactions and identifies new RNA-binding proteins in Saccharomyces cerevisiae. Genome Res. 23, 1028–1038 (2013).
Simsek, D. et al. The mammalian ribo-interactome reveals ribosome functional diversity and heterogeneity. Cell 169, 1051–1065 e18 (2017).
Acknowledgements
We are grateful to D. Nathan, G. Daley, L. Zon and members of the Sankaran laboratory for valuable guidance and suggestions. We are grateful to D. Tenen for assistance with RNA immunoprecipitation and H. Keshishian for assistance with mass spectra data access. This work was supported by National Institutes of Health grants nos. U01 HL117720, R01 DK103794 and R33 HL120791 (to V.G.S.) and P01 DK32094 (to N.M.), a gift from the Lodish Family to Boston Children’s Hospital (to V.G.S.) and the New York Stem Cell Foundation (to V.G.S.). V.G.S. is a NYSCF–Robertson Investigator.
Author information
Authors and Affiliations
Contributions
A.B. and V.G.S. conceived and designed the study. A.B., M.M. and K.E.M. performed experiments and analyzed data. C.A.L. and J.C.U. performed analyses. C.R.H., M.S., J.L., Y.W., Y.H., X.W., L.G., C.M.R., X.A., H.A.C., N.M., S.A.C., J.-J.C., S.H.O. and E.S.L. provided experimental assistance, reagents and advice. V.G.S. supervised all experimental and analytic aspects of this work. N.M. and V.G.S. acquired funding. A.B., M.M. and V.G.S. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 BCL11A protein and mRNA expression in fetal, newborn and adult erythroid cells.
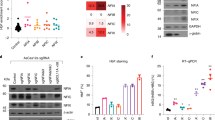
a, Representative flow cytometry plots showing CD71 and CD235a surface expression in newborn (left) and adult (right) at differentiation day 7. Mean ± s.d shown. (n = 3; 3 biologically independent experiments). b, Representative westerns showing BCL11A expression, with GAPDH as control, in newborn (left) and adult (right) at differentiation day 7 (5 independent experiments). c, BCL11A mRNA expression (normalized to GAPDH), in newborn and adult at differentiation day 7 (n = 3; 3 biologically independent experiments). Mean ± s.d shown. Two-sided Student t-test used. N.S., not significant; P = 0.7153. d, Representative westerns showing BCL11A expression, with GAPDH used as control, at differentiation days 4, 7, & 10 (5 independent experiments). e, BCL11A mRNA expression (normalized to GAPDH) in newborn and adult (n = 3; 3 independent experiments) at differentiation days 4, 7, and 10. Mean ± s.d shown. Two-sided Student t-test used. N.S., not significant; P = 0.4395 (d4), P = 0.3051 (d7), P = 0.3672 (d10). f, XL isoform of BCL11A mRNA with 4 exons and qRT–PCR primers. FP, forward primer; RP, reverse primer. g, BCL11A mRNA expression (normalized to GAPDH), in newborn and adult (n = 3; 3 independent experiments) with 2 independent primer sets at differentiation day 7. Mean ± s.d shown. Two-sided Student t-test used. N.S., not significant, P = 0.2365 (pair 1), P = 0.4099 (pair 2). h, Stacked bar graphs showing fetal (HbF, red) and adult (HbA, grey) hemoglobin abundance (by HPLC) in newborn and adult on differentiation day 16. i, Representative westerns showing BCL11A expression with GAPDH as control at differentiation days 4, 7, 10, and 12 in fetal and adult (3 independent experiments). Blots have been cropped and the corresponding full blots are available in the Source Data files.
Extended Data Fig. 2 Assessment of BCL11A and other mRNAs expressed in newborn and adult erythroid cells using RNA sequencing.
a, Scatter plots of gene expression, determined from RNA-sequencing reads, expressed as log2 TPM (transcripts per million reads) in adult and newborn primary erythroid cells. ‘r’ represents the Pearson correlation coefficient. b, Expression of BCL11A, represented as log2 TPM, between newborn (n = 2) and adult (n = 2) erythroid cells. Error bars show s.d. c, BCL11A mRNA structure and splicing is comparable between developmental stages. Sashimi plots depicting exon-exon spanning reads are shown for annotated isoforms of BCL11A. d, Representation of known BCL11A isoforms. e, Relative abundances of BCL11A isoforms in newborn (n = 2) and adult (n = 2). Error bars show s.d. No transcript was differentially expressed at P < 0.01 between the newborn and adult cells.
Extended Data Fig. 3 BCL11A protein expression is regulated via translation by polysome-associated LIN28B.
a, Western blots for BCL11A in the input and flow-through (FT) fractions of the immunoprecipitate in newborn (left) and adult (right) erythroid cells (2 independent experiments). b, Western blots for BCL11A and GATA1 in the total immunoprecipitate (IP) in newborn (left) and adult (right) erythroid cells after L-azidohomoalanine (L-AHA) labeling for 6 hours at day 7 of differentiation, followed by immunoprecipitation with BCL11A and GATA1 antibodies. c, Quantification of adult-BCL11A (blue) and newborn-BCL11A (purple) mRNAs across the different sucrose gradient fractions are shown as a percentage of the gradient. Cells were differentiated until day 7. Blots have been cropped and the corresponding full blots are available in the Source Data files.
Extended Data Fig. 4 LIN28B association with polysomes and expression in newborn and adult cells.
Representative LIN28B occupancy across polysome fractions in newborn erythroid cells at day 7 of differentiation. LIN28B abundance is probed by western blot, using RPL5 and RPS20 as controls. Please note two gaps in the western blot between sequential polysome fractions that were placed to avoid overloading of proteins (2 independent experiments). Blots have been cropped and the corresponding full blots are available in the Source Data files.
Extended Data Fig. 5 LIN28B partially dissociates from polysomes after RNase A treatment.
a, LIN28B occupancy across polysome fractions in erythroid cells either untreated (blue) or treated with RNase A (red). Experiment repeated 3 times. b, LIN28B abundance probed by western blot, using RPL5 and RPS20 as controls in the untreated sucrose gradient fractions. c, LIN28B abundance in polysome fractions with lysates digested with RNase A. Blots have been cropped and the corresponding full blots are available in the Source Data files.
Extended Data Fig. 6 LIN28B expression in newborn and adult cells.
a, Volcano plot of differentially expressed genes between adult (n = 2) and newborn (n = 2) erythroid cells. Each dot is a gene with the value of the β coefficient (x-axis) from the sleuth linear model and the corresponding measure of statistical significance (y-axis). LIN28B is the most over-expressed gene in newborn cells compared to adult. Statistical test: generalized linear model from sleuth. b, Expression of LIN28B, represented as log2 TPM, between newborn and adult erythroid cells. Error bars show s.d. ***P < 0.001. Statistical test: generalized linear model from sleuth.
Extended Data Fig. 7 Effects of high-level LIN28B expression in adult erythroid cells.
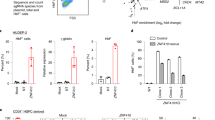
a, BCL11A mRNA levels (normalized to GAPDH expression), upon high-level LIN28B expression (GFP-high) in adult erythroid cells, assessed on differentiation day 7 (n = 3; 3 biologically independent experiments). Mean is plotted and error bars show s.d. Two-sided Student t-test used. **P < 0.01. b, Relative γ-globin expression as a percentage of total globins (γ- and β-globins), upon high-level LIN28B expression (GFP-high) in adult erythroid cells, on differentiation day 15 (n=3; 3 independent experiments). Error bars show s.d. ***P < 0.001. c, Representative flow cytometry plots showing CD71 and CD235a surface expression in control (left) and physiological level LIN28B expressing (right) adult erythroid cells at day 7 of differentiation. Data represents mean ± s.d., n = 3; 3 biologically independent experiments.
Extended Data Fig. 8 LIN28B associates with BCL11A mRNA.
a, RNA-immunoprecipitation (RNA-IP) in newborn erythroid cells at day 7 of differentiation, with antibodies against LIN28B (4196, Cell Signaling) or control IgG. Detection of BCL11A, HMGA1, GATA1, ALAS2, LDB1, KLF1, and LMO2 mRNAs (n = 3; 3 independent experiments). Mean is plotted and error bars show s.d. Two-sided Student t-test used. **P < 0.01; N.S., not significant, P = 0.6395 (GATA1), P = 0.8782 (ALAS2), P = 0.8999 (LDB1), P = 0.9571 (KLF1), P = 0.9550 (LMO2). b, Detection of BCL11A, HMGA1, GATA1, ALAS2, LDB1, KLF1, and LMO2 mRNAs (n = 3; 2 independent experiments) in the input for LIN28B antibody (4196, Cell Signaling). Mean is plotted and error bars show s.d. Two-sided Student t-test used. N.S., not significant, P = 0.3867 (BCL11A), P = 0.8050 (HMGA1), P = 0.6420 (GATA1), P = 0.8656 (ALAS2), P = 0.6413 (LDB1), P = 0.3257 (KLF1), P = 0.7152 (LMO2). c, RNA-IP in newborn erythroid cells with antibodies against LIN28B (A303-588A, Bethyl Labs) or control rabbit IgG. Detection of BCL11A (left), HMGA1 (center), and GATA1 (right) mRNAs (n = 3; 2 independent experiments). Mean is plotted and error bars show s.d. Two-sided Student t-test used. *P < 0.05; ***P < 0.001; N.S., not significant, P = 0.5393. d, Detection of BCL11A (left), HMGA1 (center) and GATA1 (right) mRNAs (n = 3; 3 independent experiments) in the input for LIN28B antibody (A303-588A, Bethyl Labs). Mean is plotted and error bars show s.d. Two-sided Student t-test used. N.S., not significant, P = 0.5261 (BCL11A), P = 0.4871 (HMGA1), P = 0.9464 (GATA1).
Extended Data Fig. 9 CLIP-seq of LIN28B in newborn erythroid cells identifies genome-wide binding peaks.
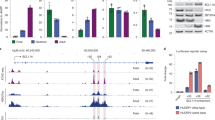
a, Genomic annotation of LIN28B binding sites. Peaks passing a 1% IDR for the consensus between replicates and a 5% false-discovery rate for each replicate are shown. b, 4-mer motifs associated with LIN28B binding sites. The dashed lines represent a threshold of 60%. 17 4-mers are present at >60% in both exons and UTRs, including GGAG, GAAG, and AAGA. c, Fold enrichment of genomic distance covered by LIN28B binding peaks compared to the genome-wide proportions. d, Rank-order enrichment of LIN28B binding peaks genome-wide. Each peak’s enrichment, measured by its -log10 q-value is plotted against its rank genome-wide for both replicates (n = 2). Statistical test: macs2 peak calling algorithm. e, Total coverage and mutation proportion for each replicate of the LIN28B/BCL11A binding site. f, Volcano plots showing the log-fold change with Benjamini-Hochberg adjusted P-values for LIN28B target genes compared between newborn (n = 2) and adult (n = 2) proerythroblasts. Statistical test: generalized linear model from sleuth.
Extended Data Fig. 10 Suppression of γ-globin by BCL11A expression in newborn erythroid cells.
a, γ-globin levels upon BCL11A expression in newborn erythroid cells on differentiation day 12 (n = 3; 3 biologically independent experiments) in control and BCL11A expressing cells. Mean is plotted and error bars show s.d. Two-sided student t-test used. ****P < 0.0001. b, Representative western blots showing BCL11A expression from lentiviral construct in newborn erythroid cells at day 12 of differentiation. GAPDH is used as a loading control (3 independent experiments). c, Representative flow cytometry plots showing CD71 and CD235a surface expression in control (left) and BCL11A expressing (right) newborn erythroid cells at days 8, 10 and 12 of differentiation (3 biologically independent experiments). Blots have been cropped and the corresponding full blots are available in the Source Data files.
Supplementary information
Supplementary Information
Supplementary Fig. 1, Tables 3 and 4, and Note
41588_2019_568_MOESM3_ESM.xlsx
Supplementary Tables 1 and 2. Data relevant to 18 S rRNA RAP–mass spectrometry and list of proteins with relative enrichment to 18 S rRNA and their relative newborn versus adult enrichment.
Source data
Fig. 1
Unprocessed immunoblots.
Fig. 2
Unprocessed immunoblots.
Fig. 3
Unprocessed immunoblots.
Fig. 4
Unprocessed immunoblots.
Extended Data Fig. 1
Unprocessed immunoblots.
Extended Data Fig. 3
Unprocessed immunoblots.
Extended Data Fig. 4
Unprocessed immunoblots.
Extended Data Fig. 5
Unprocessed immunoblots.
Extended Data Fig. 10
Unprocessed immunoblots.
Rights and permissions
About this article
Cite this article
Basak, A., Munschauer, M., Lareau, C.A. et al. Control of human hemoglobin switching by LIN28B-mediated regulation of BCL11A translation. Nat Genet 52, 138–145 (2020). https://doi.org/10.1038/s41588-019-0568-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0568-7
This article is cited by
-
A genetic disorder reveals a hematopoietic stem cell regulatory network co-opted in leukemia
Nature Immunology (2023)
-
HIC2 controls developmental hemoglobin switching by repressing BCL11A transcription
Nature Genetics (2022)
-
Translational control of stem cell function
Nature Reviews Molecular Cell Biology (2021)
-
A unified model of human hemoglobin switching through single-cell genome editing
Nature Communications (2021)
-
Identification of RNA-binding proteins that partner with Lin28a to regulate Dnmt3a expression
Scientific Reports (2021)