Abstract
Childhood brain tumors have suspected prenatal origins. To identify vulnerable developmental states, we generated a single-cell transcriptome atlas of >65,000 cells from embryonal pons and forebrain, two major tumor locations. We derived signatures for 191 distinct cell populations and defined the regional cellular diversity and differentiation dynamics. Projection of bulk tumor transcriptomes onto this dataset shows that WNT medulloblastomas match the rhombic lip-derived mossy fiber neuronal lineage and embryonal tumors with multilayered rosettes fully recapitulate a neuronal lineage, while group 2a/b atypical teratoid/rhabdoid tumors may originate outside the neuroectoderm. Importantly, single-cell tumor profiles reveal highly defined cell hierarchies that mirror transcriptional programs of the corresponding normal lineages. Our findings identify impaired differentiation of specific neural progenitors as a common mechanism underlying these pediatric cancers and provide a rational framework for future modeling and therapeutic interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Bulk and single-cell RNA-sequencing data for normal human and patient tumor samples have been deposited in the European Genome-phenome Archive under accession number EGAS00001003368. Single-cell RNA-sequencing data for normal mouse samples have been deposited in NCBI GEO under accession number GSE133531. Bulk RNA-sequencing data for human tumor-derived cell lines have been previously deposited in NCBI GEO and are available under accession number GSE117446.
Code availability
Our R package for analysis and visualization of single-cell RNA-seq data, cytobox, which was used to generate the figures presented here, is available on GitHub at https://github.com/fungenomics/cytobox under a GPL v.3.0 license.
References
Kieran, M. W., Walker, D., Frappaz, D. & Prados, M. Brain tumors: from childhood through adolescence into adulthood. J. Clin. Oncol. 28, 4783–4789 (2010).
Fontebasso, A. M. et al. Epigenetic dysregulation: a novel pathway of oncogenesis in pediatric brain tumors. Acta Neuropathol. 128, 615–627 (2014).
Jacob, K. et al. Genetic aberrations leading to MAPK pathway activation mediate oncogene-induced senescence in sporadic pilocytic astrocytomas. Clin. Cancer Res. 17, 4650–4660 (2011).
Johann, P. D. et al. Atypical teratoid/rhabdoid tumors are comprised of three epigenetic subgroups with distinct enhancer landscapes. Cancer Cell 29, 379–393 (2016).
Northcott, P. A. et al. The whole-genome landscape of medulloblastoma subtypes. Nature 547, 311–317 (2017).
Sturm, D. et al. Paediatric and adult glioblastoma: multiform (epi)genomic culprits emerge. Nat. Rev. Cancer 14, 92–107 (2014).
Sturm, D. et al. New brain tumor entities emerge from molecular classification of CNS-PNETs. Cell 164, 1060–1072 (2016).
Li, M. et al. Frequent amplification of a chr19q13.41 microRNA polycistron in aggressive primitive neuroectodermal brain tumors. Cancer Cell 16, 533–546 (2009).
Kleinman, C. L. et al. Fusion of TTYH1 with the C19MC microRNA cluster drives expression of a brain-specific DNMT3B isoform in the embryonal brain tumor ETMR. Nat. Genet. 46, 39–44 (2014).
Kool, M. et al. Molecular subgroups of medulloblastoma: an international meta-analysis of transcriptome, genetic aberrations, and clinical data of WNT, SHH, Group 3, and Group 4 medulloblastomas. Acta Neuropathol. 123, 473–484 (2012).
Northcott, P. A., Korshunov, A., Pfister, S. M. & Taylor, M. D. The clinical implications of medulloblastoma subgroups. Nat. Rev. Neurol. 8, 340–351 (2012).
Gibson, P. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010).
Fontebasso, A. M. et al. Recurrent somatic mutations in ACVR1 in pediatric midline high-grade astrocytoma. Nat. Genet. 46, 462–466 (2014).
Khuong-Quang, D. A. et al. K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol. 124, 439–447 (2012).
Wu, G. et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet. 44, 251–253 (2012).
Mackay, A. et al. Integrated molecular meta-analysis of 1,000 pediatric high-grade and diffuse intrinsic pontine glioma. Cancer Cell 32, 520–537 (2017).
Sturm, D. et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22, 425–437 (2012).
Wu, G. et al. The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat. Genet. 46, 444–450 (2014).
Schwartzentruber, J. et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482, 226–231 (2012).
Versteege, I. et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394, 203–206 (1998).
Torchia, J. et al. Integrated (epi)-genomic analyses identify subgroup-specific therapeutic targets in CNS rhabdoid tumors. Cancer Cell 30, 891–908 (2016).
Chun, H. E. et al. Genome-wide profiles of extra-cranial malignant rhabdoid tumors reveal heterogeneity and dysregulated developmental pathways. Cancer Cell 29, 394–406 (2016).
Fan, X. et al. Spatial transcriptomic survey of human embryonic cerebral cortex by single-cell RNA-seq analysis. Cell Res. 28, 730–745 (2018).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Mayer, C. et al. Developmental diversification of cortical inhibitory interneurons. Nature 555, 457–462 (2018).
Mi, D. et al. Early emergence of cortical interneuron diversity in the mouse embryo. Science 360, 81–85 (2018).
Yuzwa, S. A. et al. Developmental emergence of adult neural stem cells as revealed by single-cell transcriptional profiling. Cell Rep. 21, 3970–3986 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Vladoiu, M. C. et al. Childhood cerebellar tumours mirror conserved fetal transcriptional programs. Nature 572, 67–73 (2019).
Lin, C. Y. et al. Active medulloblastoma enhancers reveal subgroup-specific cellular origins. Nature 530, 57–62 (2016).
Clifford, S. C. et al. Wnt/Wingless pathway activation and chromosome 6 loss characterize a distinct molecular sub-group of medulloblastomas associated with a favorable prognosis. Cell Cycle 5, 2666–2670 (2006).
Valdora, F. et al. Epigenetic silencing of DKK3 in medulloblastoma. Int. J. Mol. Sci. 14, 7492–7505 (2013).
Korshunov, A. et al. Embryonal tumor with abundant neuropil and true rosettes (ETANTR), ependymoblastoma, and medulloepithelioma share molecular similarity and comprise a single clinicopathological entity. Acta Neuropathol. 128, 279–289 (2014).
Neumann, J. E. et al. A mouse model for embryonal tumors with multilayered rosettes uncovers the therapeutic potential of Sonic-hedgehog inhibitors. Nat. Med. 23, 1191–1202 (2017).
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Birey, F. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Han, Z. Y. et al. The occurrence of intracranial rhabdoid tumours in mice depends on temporal control of Smarcb1 inactivation. Nat. Commun. 7, 10421 (2016).
Lu, J. Q., Wilson, B. A., Yong, V. W., Pugh, J. & Mehta, V. Immune cell infiltrates in atypical teratoid/rhabdoid tumors. Can. J. Neurol. Sci. 39, 605–612 (2012).
Funato, K., Major, T., Lewis, P. W., Allis, C. D. & Tabar, V. Use of human embryonic stem cells to model pediatric gliomas with H3.3K27M histone mutation. Science 346, 1529–1533 (2014).
Pathania, M. et al. H3.3K27M cooperates with Trp53 loss and PDGFRA gain in mouse embryonic neural progenitor cells to induce invasive high-grade gliomas. Cancer Cell 32, 684–700.e9 (2017).
Monje, M. et al. Hedgehog-responsive candidate cell of origin for diffuse intrinsic pontine glioma. Proc. Natl Acad. Sci. USA 108, 4453–4458 (2011).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Krug, B. et al. Pervasive H3K27 acetylation leads to ERV expression and a therapeutic vulnerability in H3K27M gliomas. Cancer Cell 35, 782–797.e8 (2019).
Harutyunyan, A. S. et al. H3K27M induces defective chromatin spread of PRC2-mediated repressive H3K27me2/me3 and is essential for glioma tumorigenesis. Nat. Commun. 10, 1262 (2019).
Vitte, J., Gao, F., Coppola, G., Judkins, A. R. & Giovannini, M. Timing of Smarcb1 and Nf2 inactivation determines schwannoma versus rhabdoid tumor development. Nat. Commun. 8, 300 (2017).
Nguyen, Q. H. et al. Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity. Nat. Commun. 9, 2028 (2018).
Nagy, C. et al. Single-nucleus RNA sequencing shows convergent evidence from different cell types for altered synaptic plasticity in major depressive disorder. Preprint at bioRxiv https://doi.org/10.1101/384479 (2019).
Morrissy, A. S. et al. Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529, 351–357 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Hu, H. et al. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 47, D33–D38 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Kinsella, R. J. et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database (Oxford) 2011, bar030 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453 (2015).
Storm, R. et al. The bHLH transcription factor Olig3 marks the dorsal neuroepithelium of the hindbrain and is essential for the development of brainstem nuclei. Development 136, 295–305 (2009).
Ray, R. S. & Dymecki, S. M. Rautenlippe Redux—toward a unified view of the precerebellar rhombic lip. Curr. Opin. Cell Biol. 21, 741–747 (2009).
Landsberg, R. L. et al. Hindbrain rhombic lip is comprised of discrete progenitor cell populations allocated by Pax6. Neuron 48, 933–947 (2005).
Wang, V. Y., Rose, M. F. & Zoghbi, H. Y. Math1 expression redefines the rhombic lip derivatives and reveals novel lineages within the brainstem and cerebellum. Neuron 48, 31–43 (2005).
Dipietrantonio, H. J. & Dymecki, S. M. Zic1 levels regulate mossy fiber neuron position and axon laterality choice in the ventral brain stem. Neuroscience 162, 560–573 (2009).
Morales, D. & Hatten, M. E. Molecular markers of neuronal progenitors in the embryonic cerebellar anlage. J. Neurosci. 26, 12226–12236 (2006).
Li, S., Qiu, F., Xu, A., Price, S. M. & Xiang, M. Barhl1 regulates migration and survival of cerebellar granule cells by controlling expression of the neurotrophin-3 gene. J. Neurosci. 24, 3104–3114 (2004).
Hidalgo-Sanchez, M., Backer, S., Puelles, L. & Bloch-Gallego, E. Origin and plasticity of the subdivisions of the inferior olivary complex. Dev. Biol. 371, 215–226 (2012).
Acknowledgements
This work was supported by funding from: a Large-Scale Applied Research Project grant from Genome Quebec, Genome Canada, the Government of Canada and the Ministère de l'Économie, de la Science et de l’Innovation du Québec, with the support of the Ontario Research Fund through funding provided by the Government of Ontario to N.J., M.D.T., C.L.K., P.B.D., G.B., J.R. and L.G.; the Canadian Institutes for Health Research (CIHR grant nos. PJT-156086, to C.L.K., and MOP-286756 and FDN-154307, to N.J.); the US National Institutes of Health (NIH grant nos. P01-CA196539, to N.J., R01CA148699 and R01CA159859, to M.D.T.); the Canadian Cancer Society (CCSRI grant no. 705182), NSERC (grant no. RGPIN-2016-04911) and the Fonds de Recherche du Québec en Santé (FRQS) salary award to C.L.K.; National Sciences and Engineering Research Council (grant no. NSERC-448167-2013) and FRQS (grant no. 25348) to G.B.; CFI Leaders Opportunity Fund (grant nos. 32557, to J.R., and 33902, to C.L.K.), Genome Canada Science Technology Innovation Centre, Compute Canada Resource Allocation Project (grant no. WST-164-AB) and Genome Canada Genome Innovation Node (grant no. 244819) to J.R.; and the Fondation Charles-Bruneau. Data analyses were enabled by computer and storage resources provided by Compute Canada and Calcul Québec. N.J. is a member of the Penny Cole Laboratory and the recipient of a Chercheur Boursier, Chaire de Recherche Award from the FRQS. This work was performed within the context of the International Childhood Astrocytoma Integrated Genomic and Epigenomic (ICHANGE) consortium, and the Stand Up to Cancer (SU2C) Canada Cancer Stem Cell Dream Team Research Funding (grant no. SU2C-AACR-DT-19-15, to M.D.T. and N.J.) and SU2C St. Baldrick’s Pediatric Dream Team Translational Research Grant (no. SU2C-AACR-DT1113, to M.D.T.), with funding from Genome Canada and Genome Quebec. Stand Up to Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research. M.D.T. is supported by The Pediatric Brain Tumour Foundation, The Canadian Institutes of Health Research, The Cure Search Foundation, b.r.a.i.n.child, Meagan’s Walk, SWIFTY Foundation, Genome Canada, Genome BC, Genome Quebec, the Ontario Research Fund, Worldwide Cancer Research, V-Foundation for Cancer Research, Cancer Research UK Brain Tumour Award, Canadian Cancer Society Research Institute Impact grant and the Garron Family Chair in Childhood Cancer Research at the Hospital for Sick Children and the University of Toronto. S.J. is supported by a fellowship from CIHR. A.B.-C. is supported by a fellowship from FRQS and TD/LDI. N.D.J. is a recipient of a fellowship from FRQS and RMGA. M.K.M. is funded by a CIHR Banting postdoctoral fellowship. We thank K. Mann, S. Spira and J. Di Noia for critical reading of the manuscript, and S. Krumholtz for graphical editing of figures. We are especially grateful for the generous philanthropic donations of the Fondation Charles-Bruneau, and the Kat D-DIPG, Poppies for Irina and We Love You Connie Foundations.
Author information
Authors and Affiliations
Contributions
S.J. and C.L.K. designed and coordinated analysis of single-cell data. S.J. and A.B.-C. performed the majority of the scRNA-seq data analyses and visualizations. J.M. and G.B. contributed to the CNA analysis. Y.H. contributed to the algorithm for marker gene discovery. F.M.G.C., M.C., A.B.-C. and S.J. analyzed transcription factor activity in the scRNA-seq data. N.D.J., S.H. and S.J. contributed to the analysis of the bulk RNA-seq data and the data availability submission. M.V. contributed to timed mating and tissue isolation in developing mouse embryos. D.F., M.V. and L.K.D. contributed to primary tissue isolation, preparation and production of scRNA-seq libraries. B.K. performed all experiments in cellular models. L.G., S.J., W.T.F. and K.K.M. contributed to literature review and cell cluster annotations. L.G. provided expert advice on identification of developing pre-cerebellar populations. M.K.M. and L.G.M. contributed to the clinical annotation of tumor samples. P.-E.L. and G.T. provided bulk adult human brain RNA-seq samples. M.R., B.P. and A.A. provided human fetal brain samples. B.E., A.G.W., J.A., J.-P.F., R.W.R.D., V.L., L.C., S.A., M.G.F., H.S. and P.B.D. contributed to the collection and processing of tumor samples. J.R. contributed to optimizing the scRNA-seq protocol and provided equipment and expert advice. D.F., C.N. and G.T. contributed to the optimization of snRNA-seq protocols. S.C. helped design computational analyses. S.J., C.L.K. and N.J. wrote the manuscript with input from M.D.T., W.T.F., L.G. and K.K.M. C.L.K., N.J. and M.D.T. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Overview of the single-cell transcriptomic atlas of the developing brain.
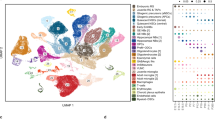
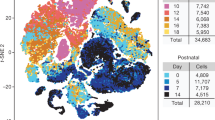
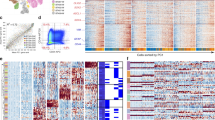
a, Overview of the approach. PCW, post-conception weeks. WNT MB, WNT-subtype medulloblastoma; ETMR, embryonal tumors with multilayered rosettes; ATRT, atypical teratoid/rhabdoid tumors; pHGG, pediatric high-grade gliomas; HGNET, high-grade neuroepithelial tumor; LGG, low-grade gliomas. b, Schematics of mouse brain regions included in dissections; figures adapted from the Allen Brain Atlas. At E12.5 and E15.5, the hindbrain (E12.5) and pons (E15.5) dissections included all of the rhombomere 1 structures with the exception of the cerebellar hemisphere, and all of the structures in rhombomeres 2-11. The forebrain dissections included parts of the dorsal pallium, central subpallium, subpallium, and septopallidal transition area. At P0, P3, and P6, the pons dissections included all of the rhombomere 1 structures with the exception of the prepontine hindbrain, and all of rhombomeres 2-11 with the exception of the roof plate structures in rhombomeres 1 to 6. The forebrain dissections included parts of the alar and roof plates of the telencephalon (including the dorsal pallium and medial pallium), and parts of the thalamus in prosomere 2, the prethalamus in prosomere 3, the preoptic alar plate, and the alar parts of the peduncular and terminal hypothalamus (original figures: © 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas. Available from: developingmouse.brain-map.org). c, t-SNE embeddings of individual mouse hindbrain/pons samples, colored by cluster. Number of cells in each sample is indicated in parentheses at bottom left; see Supplementary Table 2a for description of clusters. d, t-SNE embeddings for mouse forebrain samples, as in (c). e, Labeled t-SNE embedding of the joint mouse forebrain (n = 33,641 cells; Supplementary Table 2b). f, Proportion of cells from each major cell class in the forebrain over the timecourse. g-h, Overview of single-cell human fetal brainstem dataset. g, Labeled t-SNE plots for each sample. Number of cells in each sample is indicated in parentheses at bottom left; see Supplementary Table 2a for description of clusters. h, Proportion of cells from each major cell class in human samples.
Extended Data Fig. 2 Quality control and cell type labeling strategies in scRNA-seq atlas of the developing brain.
a, Distribution of quality control statistics for the E12.5 mouse forebrain. UMIs, unique molecular identifiers. Number of cells in each cluster is indicated in parentheses; clusters with > 100 cells are shown. Violins are colored by cluster identity, and generated as in Fig. 7. b, Illustration of quantification of cell-type specific gene sets (Supplementary Table 1a) to assign broad cell class. E12.5 mouse forebrain is shown. Number of cells in each cluster is indicated in parentheses. c-d, Gene expression distribution for selected cell type-specific canonical markers (Supplementary Table 1b) in clusters of the joint mouse pons (c) and forebrain (d). Number of cells in each cluster is indicated in Supplementary Table 2b, c. Violins are colored by cluster identity and generated as in Fig. 7, with all violins scaled to the same width. e, Heat maps of Spearman correlations of gene expression between clusters in the mouse dataset in this study (columns), and representative populations from a published atlas of the mouse central nervous system by Zeisel et al, 2018, Cell33 (rows). For populations within the Zeisel et al dataset, a representative cluster was selected from each developmental compartment (see Supplementary Note for details). Color annotation on columns corresponds to cluster identity. Number of cells in each cluster is indicated in Supplementary Table 2a.
Extended Data Fig. 3 Patterning and differentiation dynamics during forebrain development.
a, Re-embedding of mouse forebrain progenitor populations from embryonic time points (n = 7,673 cells). Cells are colored by cluster assignment in the re-embedded t-SNE space. b, t-SNE embedding colored by expression of top discriminant gene markers for each cluster, identified using a random forest-based approach (Supplementary Note). c, In situ hybridization of selected discriminant marker genes, from the Allen Brain Atlas (© 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas. Available from: developingmouse.brain-map.org) d, Visualization of forebrain cells from E12-P0 by t-SNE (n = 25,668 cells). Top row, cell clusters are highlighted by age (left panels), or inferred pseudotime for the cortical excitatory neuron trajectory (right). Bottom row: expression of representative gene markers. Expression of each gene was normalized to a [0, 1] scale for visualization. e, Transcription factor activity along the inferred cortical excitatory neuron trajectory (Supplementary Table 3). f-g, Differentiation dynamics in the ventral forebrain inhibitory lineage as in (d-e). h, Cells in the joint forebrain atlas, as in Extended Data Fig. 1e, colored by inferred pseudotime of astro-ependymal and oligodendrocyte (n = 1,354 cells) lineages (n = 4,496 cells). i, Expression of gene markers representative of astro-ependymal (top) and oligodendrocyte (bottom) differentiation, shown in cells from the respective lineages.
Extended Data Fig. 4 Mapping of bulk transcriptomes onto developmental populations.
Best matching signatures using ssGSEA for all samples within each tumor type. For ATRT tumors, populations from a recently published timecourse of the developing mouse cerebellum38 spanning E10-P14 were also included in the projections; cerebellar signatures are denoted by ‘CB’. HGNET-BCOR, high-grade neuroepithelial tumor with BCOR alteration; EBT, embryonal brain tumor; HGG-IDH, IDH-mutant high-grade gliomas; HGG-WT, High-grade gliomas wild-type for histone and IDH1/2 mutations; HF, signature from published scRNA-seq human fetal brain dataset24 containing human cerebral cortex specimens spanning 5-37 PCW. Bars are colored by cluster from which signatures were derived.
Extended Data Fig. 5 Identification of pontine mossy fiber neurons and lower rhombic lip precursors, and analysis of WNT medulloblastoma scRNA-seq.
a, Mossy fiber neuron cluster (n = 198 cells) highlighted in the t-SNE embedding of the P0 mouse pons. b, Left: expression of Olig3, a molecular marker of the lower rhombic lip (LRL), the progenitor domain that gives rise to pre-cerebellar neuron populations including mossy fiber (MF) and climbing fiber (CF) neurons12,71,72. Right: expression of Atoh1, which identifies the MF lineage in the LRL72,73, and is required for their development74. c, Violin plots quantifying expression of genes used to determine cluster identity in the MF neuron population (n = 198 cells). Left: Pax6, Zic1 and Olig3, markers of LRL progenitors that give rise to MF neurons, identified by lineage tracing and loss of function experiments12,72,73,75. Pax6 regulates cell fate allocation in the LRL73, and Zic1 regulates MF neuron positioning and projections in the developing pons75. Middle: Atoh1, a marker of MF lineage in the LRL72,73,74. Pcsk9, a marker of the pontine nucleus, a prominent structure formed exclusively by MF neurons76. Barhl1 is required for the formation of MF nuclei, and is expressed in RL-derivatives except for the inferior olivary nucleus (ION, the structure formed by CF neurons, and the source of climbing fibers to the cerebellum)77. Right: Genes marking the climbing fiber neuron lineage72, which also originates from LRL precursors, are absent in the MF population, resolving the cluster identity (Ptf1a, Neurog1/Ngn1 and Ascl1/Mash1). Foxd3 is a marker of the mature ION78. Brn3a, which marks the ION throughout its maturation78, is undetected. Violin plots are generated as in Fig. 7. d-e, PCA of re-clustered pontine progenitors as in Fig. 2a, with cluster containing LRL precursors highlighted (d) (n = 393 cells), or cells colored by expression of selected gene markers for LRL precursors (e). Expression of each gene was normalized to a [0, 1] scale for visualization. f, In situ hybridization of selected markers in the E13.5 mouse from the Allen Brain Atlas illustrating expression patterns in the LRL. g, Expression of ZIC1, CTNNB1 and OTX2, mossy fiber neuron marker genes (BARHL1, PCSK9), and climbing fiber neuron marker genes (BRN3A, ASCL1) in the t-SNE embedding of the WNT-MB-1 patient tumor sample (n = 3,975 cells). Expression of each gene was normalized to a [0, 1] scale for visualization. Similar expression patterns were observed in the other two patient samples. h, Inferred pseudotime reconstruction from the malignant cells, represented in the t-SNE embedding of the WNT-MB-1 patient tumor sample. i-j, Characterization of two patient WNT medulloblastoma scRNA-seq samples as in Fig. 4. Top left: t-SNE and clustering, with non-malignant clusters labeled, and number of cells indicated in parentheses. Top right: expression of marker genes of malignant tumor clusters. Bottom: cells in malignant tumor clusters colored by pseudotime inferred through trajectory analysis.
Extended Data Fig. 6 Profiling of TTYH1 expression and characterization of patient ETMR scRNA-seq and snRNA-seq samples.
a, Heat maps of TTYH1 expression across developing mouse and human brain samples in this study. Expression was normalized to a [0, 1] scale within each sample for visualization. Number of cells in each cluster is indicated in Supplementary Table 2a. b, Expression of TTYH1 in the developing human brain in datasets from three published scRNA-seq studies which profiled 11 human cerebral cortex specimens spanning 5-37 PCW24 (left, n = 4,261 cells); progenitor and neuron cell populations from 12 and 13 PCW human neocortex specimens44 (top right, n = 226 cells); and human pluripotent stem-cell derived forebrain organoids45 (bottom right, n = 11,838 cells). RG, radial glia; oRG, outer radial glia; vRG, ventricular radial glia; IPC, intermediate progenitor cells; IN, inhibitory neuron; EN, excitatory neuron. Boxplots: center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range; points, outliers. c, Deconvolution (CIBERSORT) analysis of bulk ETMR samples (n = 14), using a panel of signatures from the cortical neuronal lineage. d-f, t-SNE embedding of ETMR1 tumor sample (n = 5,427 cells), with cells colored by inferred pseudotime trajectory (d), by best matching cell type when tumor cells were projected onto the developmental atlas using ssGSEA (e), or by expression of selected marker and diagnostic genes (f). Expression of each gene was normalized to a [0, 1] scale for visualization. g-h, Characterization of two additional patient ETMR samples profiled using single-nuclei RNA-sequencing as in Fig. 6. Top left: t-SNE embedding with cells colored by clustering, and number of cells indicated in parentheses. Bottom left: inferred pseudotime. Right: bubble plots of neuronal lineage markers in tumor clusters.
Extended Data Fig. 7 Characterization and mapping of ATRT patient samples.
a, Deconvolution analysis (CIBERSORT) of bulk ATRT patient samples (n = 11), using mouse developmental populations. b, Top 25 leading edge genes driving ssGSEA enrichment of F-e15 Dorsal RGC signature in bulk ATRT samples (n = 11), and other tumor types of focus (ETMR: n = 14, WNT-MB: n = 10, HGG: n = 12). Genes which are specific to the leading edge of ATRT are indicated with boxes; all other genes appear in the leading edge for this signature in other tumors. Boxplots: center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range. c, Best matching developmental populations for bulk tumors by ssGSEA, when the true lineage of origin (glial populations for HGG, and neuronal populations for WNT MB and ETMR) is removed, indicating that most tumors map non-specifically to RGCs in the absence of the lineage of origin. d-e, scRNA-seq profiling of two additional patient ATRT samples as in Fig. 7. Left: t-SNE visualization and clustering, with non-malignant clusters labeled, and number of cells indicated in parentheses. Right panels: mean expression of inferred ATRT subtype, microglia, and cytotoxic T-cell gene signatures, and expression of VIM, represented in t-SNE embedding (top) and violin plots generated as in Fig. 7 (bottom). Expression of each gene set was normalized to a [0, 1] scale for visualization in t-SNE embeddings. f-g, snRNA-seq profiling of two additional patient ATRT samples as in (d-e).
Extended Data Fig. 8 Differentiation potential is impaired in H3K27M cells.
a-b, Characterization of the 19 PCW human brainstem astrocytes (n = 258 cells), a predominant best match to H3K27M HGG. a, By PCA, the first principal component separates the two populations. b, Heat map of expression of genes most strongly positively and negatively correlated with PC1. c, Western blot of K27M-mutant H3 protein and total H3 protein confirms presence of mutation and knock-out in each replicate of K27M and KO lines respectively. d-g, Analysis of bulk RNA-seq data for DIPG cell lines (n = 2 independent experiments per condition, biological replicates). d, PCA plot. SCM, stem cell media; DM, differentiation media. e, Volcano plots of differential expression analysis between cells in DM vs. SCM for K27M lines (top) and KO lines (bottom). Red color highlights differentially expressed genes present in the human brainstem astrocyte 2 gene signature (left), and any brainstem or pontine astrocyte gene signature (right). P-values (two-sided Wald test) were adjusted using the Benjamini-Hochberg correction. f, Boxplots of log2 fold change of expression for genes in selected developmental signatures, between cells in DM vs. SCM for K27M lines (red) and KO (blue). Statistical significance was assessed using a two-tailed Student’s t-test (p-values: Hindbrain astrocyte: 1.46x10-13; Human astrocyte: 6.85x10-5; OPC/Oligodendrocyte: 0.14; Excit. Neuron: 0.12; ns: not significant). Boxplots: center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range. g, Volcano plot of differential expression analysis between K27M and K27M-KO cell lines in DM; differential expression analysis was performed as described above. h, Representative morphology of GFAP + cells among cell lines at 60X magnification. Experiment was repeated, and images are shown, for n = 2 biologically independent replicates per condition. i, Bubbleplot of projection of K27M-KO cell lines onto developmental atlas using ssGSEA, shown for the neuroectodermal cell types. The color of the bubbles indicates the change in ssGSEA score for each signature between cell lines in SCM and DM, while the size of the bubbles indicates the ssGSEA score in DM. Cell types are stratified into two rows based on direction of change of the score upon differentiation. No bubbles are shown for clusters with non-specific gene signatures.
Supplementary information
Supplementary Information
Supplementary Note and Figs. 1 and 2
Supplementary Table 1–6
Supplementary Tables 1–6
Rights and permissions
About this article
Cite this article
Jessa, S., Blanchet-Cohen, A., Krug, B. et al. Stalled developmental programs at the root of pediatric brain tumors. Nat Genet 51, 1702–1713 (2019). https://doi.org/10.1038/s41588-019-0531-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0531-7
This article is cited by
-
Chronic hypoxia remodels the tumor microenvironment to support glioma stem cell growth
Acta Neuropathologica Communications (2024)
-
Common molecular features of H3K27M DMGs and PFA ependymomas map to hindbrain developmental pathways
Acta Neuropathologica Communications (2023)
-
Pediatric glioma histone H3.3 K27M/G34R mutations drive abnormalities in PML nuclear bodies
Genome Biology (2023)
-
The tumor suppressor CREBBP and the oncogene MYCN cooperate to induce malignant brain tumors in mice
Oncogenesis (2023)
-
(B)On(e)-cohistones and the epigenetic alterations at the root of bone cancer
Cell Death & Differentiation (2023)