Abstract
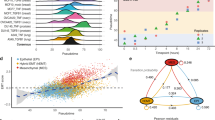
Integrating single-cell trajectory analysis with pooled genetic screening could reveal the genetic architecture that guides cellular decisions in development and disease. We applied this paradigm to probe the genetic circuitry that controls epithelial-to-mesenchymal transition (EMT). We used single-cell RNA sequencing to profile epithelial cells undergoing a spontaneous spatially determined EMT in the presence or absence of transforming growth factor-β. Pseudospatial trajectory analysis identified continuous waves of gene regulation as opposed to discrete ‘partial’ stages of EMT. KRAS was connected to the exit from the epithelial state and the acquisition of a fully mesenchymal phenotype. A pooled single-cell CRISPR-Cas9 screen identified EMT-associated receptors and transcription factors, including regulators of KRAS, whose loss impeded progress along the EMT. Inhibiting the KRAS effector MEK and its upstream activators EGFR and MET demonstrates that interruption of key signaling events reveals regulatory ‘checkpoints’ in the EMT continuum that mimic discrete stages, and reconciles opposing views of the program that controls EMT.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data are available on GEO under accession number GSE114687. Data will also be provided via the Github repository described in ‘Code availability’.
Code availability
Code can be found on Github at https://github.com/cole-trapnell-lab/pseudospace.
References
Lamouille, S., Xu, J. & Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 15, 178–196 (2014).
Nieto, M. A. Epithelial plasticity: a common theme in embryonic and cancer cells. Science 342, 1234850 (2013).
Sauka-Spengler, T. & Bronner-Fraser, M. A gene regulatory network orchestrates neural crest formation. Nat. Rev. Mol. Cell Biol. 9, 557–568 (2008).
Li, M. et al. Epithelial-mesenchymal transition: an emerging target in tissue fibrosis. Exp. Biol. Med. 241, 1–13 (2016).
Nieto, M. A., Angela Nieto, M., Huang, R. Y.-J., Jackson, R. A. & Thiery, J. P. EMT: 2016. Cell 166, 21–45 (2016).
Kalluri, R. & Weinberg, R. A. The basics of epithelial-mesenchymal transition. J. Clin. Invest. 119, 1420–1428 (2009).
Zhang, J. et al. TGF-β-induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops. Sci. Signal. 7, ra91 (2014).
Lu, M., Jolly, M. K., Levine, H., Onuchic, J. N. & Ben-Jacob, E. MicroRNA-based regulation of epithelial–hybrid–mesenchymal fate determination. Proc. Natl Acad. Sci. USA 110, 18144–18149 (2013).
Hong, T. et al. An Ovol2-Zeb1 mutual inhibitory circuit governs bidirectional and multi-step transition between epithelial and mesenchymal states. PLoS Comput. Biol. 11, e1004569 (2015).
Krishnaswamy, S., Zivanovic, N., Sharma, R., Pe’er, D. & Bodenmiller, B. Learning edge rewiring in EMT from single cell data. Preprint at bioRxiv https://doi.org/10.1101/155028 (2017).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729.e27 (2018).
Grande, M. T. et al. Snail1-induced partial epithelial-to-mesenchymal transition drives renal fibrosis in mice and can be targeted to reverse established disease. Nat. Med. 21, 989–997 (2015).
Lovisa, S. et al. Epithelial-to-mesenchymal transition induces cell cycle arrest and parenchymal damage in renal fibrosis. Nat. Med. 21, 998–1009 (2015).
Aiello, N. M. et al. EMT Subtype Influences Epithelial Plasticity and Mode of Cell Migration. Dev. Cell 45, 681–695.e4 (2018).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624.e24 (2017).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Scialdone, A. et al. Resolving early mesoderm diversification through single-cell expression profiling. Nature 535, 289–293 (2016).
Sarrió, D. et al. Epithelial-mesenchymal transition in breast cancer relates to the basal-like phenotype. Cancer Res. 68, 989–997 (2008).
Rodriguez, L. G., Wu, X. & Guan, J.-L. Wound-healing assay. Methods Mol. Biol. 294, 23–29 (2005).
Vuoriluoto, K. et al. Vimentin regulates EMT induction by Slug and oncogenic H-Ras and migration by governing Axl expression in breast cancer. Oncogene 30, 1436–1448 (2011).
Joost, S. et al. Single-cell transcriptomics reveals that differentiation and spatial signatures shape epidermal and hair follicle heterogeneity. Cell Syst. 3, 221–237.e9 (2016).
Schliekelman, M. J. et al. Molecular portraits of epithelial, mesenchymal, and hybrid States in lung adenocarcinoma and their relevance to survival. Cancer Res. 75, 1789–1800 (2015).
George, J. T., Jolly, M. K., Xu, J., Somarelli, J. & Levine, H. Survival outcomes in cancer patients predicted by a partial EMT gene expression scoring metric. Cancer Res. 77, 6415–6428 (2017).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25, 25–29 (2000).
The Gene Ontology Consortium & The Gene Ontology Consortium. The gene ontology resource: 20 years and still going strong. Nucleic Acids Res. 47, D330–D338 (2019)..
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Tchernitsa, O. I. et al. Transcriptional basis of KRAS oncogene-mediated cellular transformation in ovarian epithelial cells. Oncogene 23, 4536–4555 (2004).
Toivola, D. M., Tao, G.-Z., Habtezion, A., Liao, J. & Omary, M. B. Cellular integrity plus: organelle-related and protein-targeting functions of intermediate filaments. Trends Cell Biol. 15, 608–617 (2005).
Huang, R. Y.-J., Guilford, P. & Thiery, J. P. Early events in cell adhesion and polarity during epithelial-mesenchymal transition. J. Cell Sci. 125, 4417–4422 (2012).
Feng, Y.-X. et al. Epithelial-to-mesenchymal transition activates PERK-eIF2α and sensitizes cells to endoplasmic reticulum stress. Cancer Discov. 4, 702–715 (2014).
Miettinen, P. J., Ebner, R., Lopez, A. R. & Derynck, R. TGF-beta induced transdifferentiation of mammary epithelial cells to mesenchymal cells: involvement of type I receptors. J. Cell Biol. 127, 2021–2036 (1994).
CaulIn, C., Scholl, F. G., Frontelo, P., Gamallo, C. & Quintanilla, M. Chronic exposure of cultured transformed mouse epidermal cells to transforming growth factor-/1 induces an epithelial-mesenchymal transdifferentiation and a spindle tumoral phenotype. Cell Growth Differ. 6, 1027–1036 (1995).
Cacchiarelli, D. et al. Aligning single-cell developmental and reprogramming trajectories identifies molecular determinants of myogenic reprogramming outcome. Cell Syst. 7, 258–268.e3 (2018).
Alpert, A., Moore, L. S., Dubovik, T. & Shen-Orr, S. S. Alignment of single-cell trajectories to compare cellular expression dynamics. Nat. Methods 15, 267–270 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Vintsyuk, T. K. Speech discrimination by dynamic programming. Cybern. Syst. Anal. 4, 52–57 (1968).
Rabiner, L. & Juang, B. H. Fundamentals of Speech Recognition (PTR Prentice Hall, 1993).
Masszi, A. et al. Integrity of cell-cell contacts is a critical regulator of TGF-β1-induced epithelial-to-myofibroblast transition. Am. J. Pathol. 165, 1955–1967 (2004).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Shaul, Y. D. et al. Dihydropyrimidine accumulation is required for the epithelial-mesenchymal transition. Cell 158, 1094–1109 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Kuo, P.-L., Shen, K.-H., Hung, S.-H. & Hsu, Y.-L. CXCL1/GROα increases cell migration and invasion of prostate cancer by decreasing fibulin-1 expression through NF-κB/HDAC1 epigenetic regulation. Carcinogenesis 33, 2477–2487 (2012).
Al-Alwan, L. A. et al. Differential roles of CXCL2 and CXCL3 and their receptors in regulating normal and asthmatic airway smooth muscle cell migration. J. Immunol. 191, 2731–2741 (2013).
Tian, X.-J., Zhang, H. & Xing, J. Coupled reversible and irreversible bistable switches underlying TGFβ-induced epithelial to mesenchymal transition. Biophys. J. 105, 1079–1089 (2013).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882.e21 (2016).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-Seq. Cell 167, 1883–1896.e15 (2016).
Xie, S., Duan, J., Li, B., Zhou, P. & Hon, G. C. Multiplexed engineering and analysis of combinatorial enhancer activity in single cells. Mol. Cell 66, 285–299.e5 (2017).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 5, 271–274 (2018).
Clark, E. A. & Hynes, R. O. Ras activation is necessary for integrin-mediated activation of extracellular signal-regulated kinase 2 and cytosolic phospholipase A2but not for cytoskeletal organization. J. Biol. Chem. 271, 14814–14818 (1996).
Citri, A. & Yarden, Y. EGF–ERBB signalling: towards the systems level. Nat. Rev. Mol. Cell Biol. 7, 505–516 (2006).
Peschard, P. & Park, M. From Tpr-Met to Met, tumorigenesis and tubes. Oncogene 26, 1276–1285 (2007).
Ornitz, D. M. & Itoh, N. The fibroblast growth factor signaling pathway. Wiley Interdiscip. Rev. Dev. Biol. 4, 215–266 (2015).
Xu, J., Lamouille, S. & Derynck, R. TGF-β-induced epithelial to mesenchymal transition. Cell Res. 19, 156–172 (2009).
Ahmad, I., Iwata, T. & Leung, H. Y. Mechanisms of FGFR-mediated carcinogenesis. Biochim. Biophys. Acta 1823, 850–860 (2012).
Reed, N. I. et al. The v 1 integrin plays a critical in vivo role in tissue fibrosis. Sci. Transl. Med. 7, 288ra79–288ra79 (2015).
Kalluri, R. & Neilson, E. G. Epithelial-mesenchymal transition and its implications for fibrosis. J. Clin. Invest. 112, 1776–1784 (2003).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170, 17–33 (2017).
Karnoub, A. E. & Weinberg, R. A. Ras oncogenes: split personalities. Nat. Rev. Mol. Cell Biol. 9, 517–531 (2008).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Acknowledgements
We thank all members of the Trapnell and Shendure laboratories for helpful discussions during the course of this study and feedback on our manuscript, particularly S. Srivatsan, L. Saunders, H. Pliner and J. Packer. We thank N.M. Cruz for feedback on our manuscript. J.L.M.F. thanks S.V. McFaline-Cruz for support. J.L.M.F. was supported by NIH grants T32HL007828 and T32HG000035. A.J.H. was supported by an NSF Graduate Research Fellowship. J.S. and C.T. are supported by NIH grant no. U54DK107979 and the Paul G. Allen Frontiers Group. C.T. is supported by NIH grant nos. DP2HD088158, RC2DK114777 and R01HL118342 and is partly supported by an Alfred P. Sloan Foundation Research Fellowship. J.S. is supported by NIH grant nos. DP1HG007811 and R01HG006283 and is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.L.M.F., J.S. and C.T. devised the project. J.L.M.F., A.J.H., J.S. and C.T. designed experiments. J.L.M.F., A.J.H. and D.J. performed experiments. D.J. and X.Q. provided substantial technical and computational support, respectively. J.L.M.F and A.J.H. performed analyses. J.L.M.F. and C.T. wrote the manuscript with the support of the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–32 and Table 1
Supplementary Table 2
Differential expression analysis between cell fractions of MCF10A cells undergoing spontaneous EMT.
Supplementary Table 3
Pseudospatial differential expression analysis of spontaneous EMT.
Supplementary Table 4
Geneset analysis of genes differentially expressed across pseudospace in cells undergoing spontaneous EMT.
Supplementary Table 5
Differential expression analysis between cell fractions of HuMEC cells undergoing spontaneous EMT.
Supplementary Table 6
Differential expression analysis between aligned spontaneous and TGF-β-driven MCF10A EMT trajectories.
Supplementary Table 7
Geneset analysis of genes differentially expressed between aligned spontaneous and TGF-β-driven MCF10A EMT trajectories.
Supplementary Table 8
Pseudospatial differential expression analysis between knockout and non-targeting control MCF10A cells undergoing spontaneous EMT.
Supplementary Table 9
Pseudospatial differential expression analysis between knockout and non-targeting control MCF10A cells undergoing TGF-β-driven EMT.
Supplementary Table 10
Sequences of oligonucleotides used in this study.
Rights and permissions
About this article
Cite this article
McFaline-Figueroa, J.L., Hill, A.J., Qiu, X. et al. A pooled single-cell genetic screen identifies regulatory checkpoints in the continuum of the epithelial-to-mesenchymal transition. Nat Genet 51, 1389–1398 (2019). https://doi.org/10.1038/s41588-019-0489-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0489-5
This article is cited by
-
Trajectory inference across multiple conditions with condiments
Nature Communications (2024)
-
Recent advances and applications of CRISPR-Cas9 in cancer immunotherapy
Molecular Cancer (2023)
-
The epithelial–mesenchymal plasticity landscape: principles of design and mechanisms of regulation
Nature Reviews Genetics (2023)
-
A new Bayesian factor analysis method improves detection of genes and biological processes affected by perturbations in single-cell CRISPR screening
Nature Methods (2023)
-
Harnessing epithelial-mesenchymal plasticity to boost cancer immunotherapy
Cellular & Molecular Immunology (2023)