Abstract
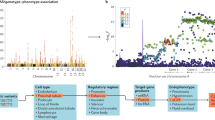
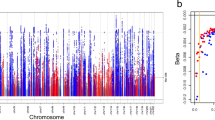
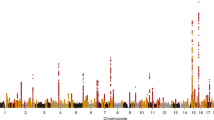
Chronic kidney disease (CKD) is responsible for a public health burden with multi-systemic complications. Through trans-ancestry meta-analysis of genome-wide association studies of estimated glomerular filtration rate (eGFR) and independent replication (n = 1,046,070), we identified 264 associated loci (166 new). Of these, 147 were likely to be relevant for kidney function on the basis of associations with the alternative kidney function marker blood urea nitrogen (n = 416,178). Pathway and enrichment analyses, including mouse models with renal phenotypes, support the kidney as the main target organ. A genetic risk score for lower eGFR was associated with clinically diagnosed CKD in 452,264 independent individuals. Colocalization analyses of associations with eGFR among 783,978 European-ancestry individuals and gene expression across 46 human tissues, including tubulo-interstitial and glomerular kidney compartments, identified 17 genes differentially expressed in kidney. Fine-mapping highlighted missense driver variants in 11 genes and kidney-specific regulatory variants. These results provide a comprehensive priority list of molecular targets for translational research.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Genome-wide summary statistics for this study have been made publicly available at http://ckdgen.imbi.uni-freiburg.de.
References
Eckardt, K. U. et al. Evolving importance of kidney disease: from subspecialty to global health burden. Lancet 382, 158–169 (2013).
Jha, V. et al. Chronic kidney disease: global dimension and perspectives. Lancet 382, 260–272 (2013).
Ene-Iordache, B. et al. Chronic kidney disease and cardiovascular risk in six regions of the world (ISN-KDDC): a cross-sectional study. Lancet Glob. Health 4, e307–e319 (2016).
Go, A. S., Chertow, G. M., Fan, D., McCulloch, C. E. & Hsu, C. Y. Chronic kidney disease and the risks of death, cardiovascular events, and hospitalization. N. Engl. J. Med. 351, 1296–1305 (2004).
GBD 2016 Causes of Death Collaborators. Global, regional, and national age-sex specific mortality for 264 causes of death, 1980–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet 390, 1151-1210 (2017).
Inrig, J. K. et al. The landscape of clinical trials in nephrology: a systematic review of ClinicalTrials.gov. Am. J. Kidney Dis. 63, 771–780 (2014).
Levin, A. et al. Global kidney health 2017 and beyond: a roadmap for closing gaps in care, research, and policy. Lancet 390, 1888–1917 (2017).
Wuttke, M. & Kottgen, A. Insights into kidney diseases from genome-wide association studies. Nat. Rev. Nephrol. 12, 549–562 (2016).
Gorski, M. et al. 1000 Genomes-based meta-analysis identifies 10 novel loci for kidney function. Sci. Rep. 7, 45040 (2017).
Pattaro, C. et al. Genetic associations at 53 loci highlight cell types and biological pathways relevant for kidney function. Nat. Commun. 7, 10023 (2016).
Chasman, D. I. et al. Integration of genome-wide association studies with biological knowledge identifies six novel genes related to kidney function. Hum. Mol. Genet. 21, 5329–5343 (2012).
Pattaro, C. et al. Genome-wide association and functional follow-up reveals new loci for kidney function. PLoS Genet. 8, e1002584 (2012).
Kottgen, A. et al. New loci associated with kidney function and chronic kidney disease. Nat. Genet. 42, 376–384 (2010).
Chambers, J. C. et al. Genetic loci influencing kidney function and chronic kidney disease. Nat. Genet. 42, 373–375 (2010).
Kottgen, A. et al. Multiple loci associated with indices of renal function and chronic kidney disease. Nat. Genet. 41, 712–717 (2009).
Kanai, M. et al. Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat. Genet. 50, 390–400 (2018).
Okada, Y. et al. Meta-analysis identifies multiple loci associated with kidney function-related traits in east Asian populations. Nat. Genet. 44, 904–909 (2012).
Hishida, A. et al. Genome-wide association study of renal function traits: results from the Japan Multi-institutional Collaborative Cohort study. Am. J. Nephrol. 47, 304–316 (2018).
Lee, J. et al. Genome-wide association analysis identifies multiple loci associated with kidney disease-related traits in Korean populations. PLoS One 13, e0194044 (2018).
Mahajan, A. et al. Trans-ethnic fine mapping highlights kidney-function genes linked to salt sensitivity. Am. J. Hum. Genet. 99, 636–646 (2016).
Devuyst, O. & Pattaro, C. The UMOD locus: insights into the pathogenesis and prognosis of kidney disease. J. Am. Soc. Nephrol. 29, 713–726 (2018).
Yeo, N. C. et al. Shroom3 contributes to the maintenance of the glomerular filtration barrier integrity. Genome Res. 25, 57–65 (2015).
Gaziano, J. M. et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol. 70, 214–223 (2016).
Benner, C. et al. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am. J. Hum. Genet. 101, 539–551 (2017).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Li, M. et al. SOS2 and ACP1 loci identified through large-scale exome chip analysis regulate kidney development and function. J. Am. Soc. Nephrol. 28, 981–994 (2017).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Jing, J. et al. Combination of mouse models and genomewide association studies highlights novel genes associated with human kidney function. Kidney Int. 90, 764–773 (2016).
Wakefield, J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
Dong, C. et al. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum. Mol. Genet. 24, 2125–2137 (2015).
Tsuda, M. et al. Targeted disruption of the multidrug and toxin extrusion 1 (Mate1) gene in mice reduces renal secretion of metformin. Mol. Pharm. 75, 1280–1286 (2009).
Sieber, K. B. et al. Integrated functional genomic analysis enables annotation of kidney genome-wide association study loci. J. Am. Soc. Nephrol. 30, 421–441 (2019).
Olden, M. et al. Common variants in UMOD associate with urinary uromodulin levels: a meta-analysis. J. Am. Soc. Nephrol. 25, 1869–1882 (2014).
Moreau, M. E. et al. The kallikrein–kinin system: current and future pharmacological targets. J. Pharm. Sci. 99, 6–38 (2005).
Battle, A., Brown, C. D., Engelhardt, B. E. & Montgomery, S. B. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Gamazon, E. R. et al. Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation. Nat. Genet. 50, 956–967 (2018).
Eckardt, K. U. et al. Autosomal dominant tubulointerstitial kidney disease: diagnosis, classification, and management—a KDIGO consensus report. Kidney Int. 88, 676–683 (2015).
Gillies, C. E. et al. An eQTL landscape of kidney tissue in human nephrotic syndrome. Am. J. Hum. Genet. 103, 232–244 (2018).
Dudley, A. J., Bleasby, K. & Brown, C. D. The organic cation transporter OCT2 mediates the uptake of β-adrenoceptor antagonists across the apical membrane of renal LLC-PK1 cell monolayers. Br. J. Pharm. 131, 71–79 (2000).
Filipski, K. K., Mathijssen, R. H., Mikkelsen, T. S., Schinkel, A. H. & Sparreboom, A. Contribution of organic cation transporter 2 (OCT2) to cisplatin-induced nephrotoxicity. Clin. Pharm. Ther. 86, 396–402 (2009).
Motohashi, H. & Inui, K. Organic cation transporter OCTs (SLC22) and MATEs (SLC47) in the human kidney. AAPS J. 15, 581–588 (2013).
Popejoy, A. B. & Fullerton, S. M. Genomics is failing on diversity. Nature 538, 161–164 (2016).
Humm, A., Huber, R. & Mann, K. The amino acid sequences of human and pig l-arginine:glycine amidinotransferase. FEBS Lett. 339, 101–107 (1994).
Olives, B. et al. Molecular characterization of a new urea transporter in the human kidney. FEBS Lett. 386, 156–160 (1996).
Phan, N. N. et al. Voltage-gated calcium channels: novel targets for cancer therapy. Oncol. Lett. 14, 2059–2074 (2017).
Thi Do, D., Phan, N. N., Wang, C. Y., Sun, Z. & Lin, Y. C. Novel regulations of MEF2-A, MEF2-D, and CACNA1S in the functional incompetence of adipose-derived mesenchymal stem cells by induced indoxyl sulfate in chronic kidney disease. Cytotechnology 68, 2589–2604 (2016).
Parsa, A. et al. Common variants in Mendelian kidney disease genes and their association with renal function. J. Am. Soc. Nephrol. 24, 2105–2117 (2013).
Xie, W. et al. Genetic variants associated with glycine metabolism and their role in insulin sensitivity and type 2 diabetes. Diabetes 62, 2141–2150 (2013).
Raffler, J. et al. Genome-wide association study with targeted and non-targeted NMR metabolomics identifies 15 novel loci of urinary human metabolic individuality. PLoS Genet. 11, e1005487 (2015).
Janicki, N. et al. Increased occurence of valporoic acid-induced hyperammonemia in carriers of T1405N polymorphism in carbamoyl phosphate synthetase 1 gene. ISRN Neurol. 2013, 261497 (2013).
Seppala, A. et al. Genome-wide association study on dimethylarginines reveals novel AGXT2 variants associated with heart rate variability but not with overall mortality. Eur. Heart J. 35, 524–531 (2014).
Sveinbjornsson, G. et al. Rare mutations associating with serum creatinine and chronic kidney disease. Hum. Mol. Genet. 23, 6935–6943 (2014).
Pescio, L. G. et al. Changes in ceramide metabolism are essential in Madin–Darby canine kidney cell differentiation. J. Lipid Res. 58, 1428–1438 (2017).
Imgrund, S. et al. Adult ceramide synthase 2 (CERS2)-deficient mice exhibit myelin sheath defects, cerebellar degeneration, and hepatocarcinomas. J. Biol. Chem. 284, 33549–33560 (2009).
Shiffman, D. et al. A gene variant in CERS2 is associated with rate of increase in albuminuria in patients with diabetes from ONTARGET and TRANSCEND. PLoS One 9, e106631 (2014).
Yoshioka, K. et al. IgA nephropathy in patients with congenital C9 deficiency. Kidney Int. 42, 1253–1258 (1992).
Fuchsberger, C., Taliun, D., Pramstaller, P. P. & Pattaro, C. GWAtoolbox: an R package for fast quality control and handling of genome-wide association studies meta-analysis data. Bioinformatics 28, 444–445 (2012).
Coresh, J. et al. Decline in estimated glomerular filtration rate and subsequent risk of end-stage renal disease and mortality. J. Am. Med. Assoc. 311, 2518–2531 (2014).
Levey, A. S. et al. A new equation to estimate glomerular filtration rate. Ann. Intern. Med. 150, 604–612 (2009).
Pattaro, C. et al. Estimating the glomerular filtration rate in the general population using different equations: effects on classification and association. Nephron Clin. Pract. 123, 102–111 (2013).
Schwartz, G. J. et al. Improved equations estimating GFR in children with chronic kidney disease using an immunonephelometric determination of cystatin C. Kidney Int. 82, 445–453 (2012).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Higgins, J. P. & Thompson, S. G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Magi, R. et al. Trans-ethnic meta-regression of genome-wide association studies accounting for ancestry increases power for discovery and improves fine-mapping resolution. Hum. Mol. Genet. 26, 3639–3650 (2017).
Hadfield, J. MCMC methods for multi-response generalized linear mixed models: the MCMC glmm R package. J. Stat. Softw. 33, 1–22 (2010).
Pattaro, C. et al. The Cooperative Health Research in South Tyrol (CHRIS) study: rationale, objectives, and preliminary results. J. Transl. Med. 13, 348 (2015).
Noce, D. et al. Sequential recruitment of study participants may inflate genetic heritability estimates. Hum. Genet. 136, 743–757 (2017).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Kottgen, A. et al. Genome-wide association analyses identify 18 new loci associated with serum urate concentrations. Nat. Genet. 45, 145–154 (2013).
Dastani, Z. et al. Novel loci for adiponectin levels and their influence on type 2 diabetes and metabolic traits: a multi-ethnic meta-analysis of 45,891 individuals. PLoS Genet. 8, e1002607 (2012).
Canela-Xandri, O., Rawlik, K. & Tenesa, A. An atlas of genetic associations in UK Biobank. Nat. Genet. 50, 1593–1599 (2018).
Fehrmann, R. S. et al. Gene expression analysis identifies global gene dosage sensitivity in cancer. Nat. Genet. 47, 115–125 (2015).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Frey, B. J. & Dueck, D. Clustering by passing messages between data points. Science 315, 972–976 (2007).
Hoppmann, A. S., Schlosser, P., Backofen, R., Lausch, E. & Kottgen, A. GenToS: use of orthologous gene information to prioritize signals from human GWAS. PLoS One 11, e0162466 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Wakefield, J. Bayes factors for genome-wide association studies: comparison with P-values. Genet. Epidemiol. 33, 79–86 (2009).
Arnold, M., Raffler, J., Pfeufer, A., Suhre, K. & Kastenmuller, G. SNiPA: an interactive, genetic variant-centered annotation browser. Bioinformatics 31, 1334–1336 (2015).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Zeller, T. et al. Genetics and beyond—the transcriptome of human monocytes and disease susceptibility. PLoS One 5, e10693 (2010).
Fehrmann, R. S. et al. Trans-eQTLs reveal that independent genetic variants associated with a complex phenotype converge on intermediate genes, with a major role for the HLA. PLoS Genet. 7, e1002197 (2011).
Westra, H. J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Joehanes, R. et al. Integrated genome-wide analysis of expression quantitative trait loci aids interpretation of genomic association studies. Genome Biol. 18, 16 (2017).
Kirsten, H. et al. Dissecting the genetics of the human transcriptome identifies novel trait-related trans-eQTLs and corroborates the regulatory relevance of non-protein coding loci. Hum. Mol. Genet. 24, 4746–4763 (2015).
Beutner, F. et al. Rationale and design of the Leipzig (LIFE) Heart Study: phenotyping and cardiovascular characteristics of patients with coronary artery disease. PLoS One 6, e29070 (2011).
Loeffler, M. et al. The LIFE-Adult-Study: objectives and design of a population-based cohort study with 10,000 deeply phenotyped adults in Germany. BMC Public Health 15, 691 (2015).
Eckardt, K. U. et al. The German Chronic Kidney Disease (GCKD) study: design and methods. Nephrol. Dial. Transpl. 27, 1454–1460 (2012).
Acknowledgements
We thank D. Di Domizio (Eurac Research) and J. Knaus (University of Freiburg) for IT assistance and T. Johnson (GlaxoSmithKline) for sharing his code and discussion on credible set fine-mapping and colocalization analysis. This research has been conducted using the UK Biobank resource under application number 20272. Study-specific acknowledgements and funding sources are listed in the Supplementary Information.
Author information
Authors and Affiliations
Consortia
Contributions
Manuscript writing group: M. Wuttke, Y.L., M. Li, K.B.S., M.F., M. Gorski, A. Tin, L. Wang, H. Kirsten, T.A., K. Ho, I.H., M. Scholz, A. Teumer, A. Köttgen, C.P. Design of the study: C.A.B., C.F., M. Gorski, A. Köttgen, A.P.M., C.P., A. Teumer, A. Tin, M. Wuttke. Management of an individual contributing study: T.S.A., E.d.A., S. Akilesh, S.J.B., G.B., M. Bochud, M. Boehnke, E.B., M.H.d.B., H.B., A.S.B., C.A.B., A.C., R.J.C., J.C.C., D.I.C., C.-Y.C., K.C., R.C., M. Ciullo, J.C., D.C., R.M.v.D., J. Danesh, O.D., C.M.v.D., K.-U.E., G.E., P.E., M.K.E., J.F.F., O.H.F., B.I.F., Y.F., R.T.G., H.G., P.G., J.M.G., V. Giedraitis, C.G., F.G., A.D.G., V. Gudnason, T.B.H., P.v.d.H., C.A.H., C.H., C.-K.H., A.A.H., K. Ho, A.M.H., M.A.I., O.S.I., E.I., V.W.J., J.B.J., B.J., C.M.K., C.-C.K., W. Kiess, M.E.K., W. Koenig, J.S.K., H. Kramer, F.K., B.K.K., M. Kubo, J.K., M. Kähönen, A. Körner, A. Köttgen, T.L., Y.L., S.-C.L., M. Loeffler, R.J.L., S.L., M.A.L., P.K.M., N.G.M., D.M., K. Matsuda, O.M., A. Metspalu, E.K.M., Y.M., K.L.M., G.W.M., A.P.M., R.d.M., W.M., G.N.N., J.O’C., M.L.O’D., A.J.O., M.O.-M., W.H.O., A.P., C.P., S.A.P., B.W.P., T. Perls, M. Perola, M. Pirastu, O.P., B.P., P.P.P., M.A.P., B.M.P., T.J.R., O.T.R., D.F.R., R. Rettig, M.R., P.M.R., D.J.R., P.R., I.R., C.S., V.S., K.-U.S., H. Schmidt, R.S., M. Scholz, B.S., X.S., H. Snieder, N. Soranzo, C.N.S., K. Stefansson, K. Strauch, M. Stumvoll, G.S., P.O.S., E.-S.T., B.O.T., Y.-C.T., J. Thiery, A. Tin, D.T., J. Tremblay, I.T., A. Tönjes, P.V., A.P.d.V., U.V., G.W., L. Wallentin, Y.X.W., D.M.W., W.B.W., H.W., J.B.W., S.H.W., J.G.W., C. Wong, T.-Y.W., M. Wuttke, L.X., Q.Y., M.Y., W.Z., A.B.Z. Statistical methods and analysis: T.S.A., M.A., P.A., M.L.B., G.B., M. Boissel, T.S.B., M. Brumat, C.A.B., M. Canouil, R.J.C., J.-F.C., D.I.C., Miao-Li Chee, X.C., Y.C., A.Y.C., M. Cocca, M.P.C., J.P.C., T.C., A. Dehghan, G.D., A. Demirkan, J. Divers, R.D., D.R.V.E., T.L.E., M.F.F., J.F.F., B.I.F., S.F.-W., C.F., S.G., A.G., M. Gorski, D.F.G., M. Gögele, T.H., P.H., P.v.d.H., I.M.H., J.N.H., E.H., A.H., K. Horn, S.-J.H., J.J., P.K.J., N.S.J., B.J., Y.K., M. Kanai, C.-C.K., H. Kirsten, M.E.K., A. Krajcoviechova, H. Kramer, M. Kuokkanen, A. Köttgen, B.K., L.A.L., C.D.L., M. Li, Y.L., Jianjun Liu, Jun Liu, L.-P.L., A. Mahajan, J. Marten, J. Martins, K. Miliku, P.P.M., N.M., A.P.M., P.J.v.d.M., W.M., M.A.N., M.N., B.N., D.N., I.M.N., R.N., T.N., Y.O., C.P., S.A.P., N.P., M.H.P., B.P.P., L.M.R., M.R., K.M.R., F. Rivadeneira, F. Rizzi, R. Rueedi, K.A.R., Y. Saba, E.S., M. Scholz, C.-A.S., S. Sedaghat, Y. Shi, K.B.S., X.S., A.V.S., C.N.S., H.M.S., G.S., S. Szymczak, S.M.T., B.O.T., A. Teumer, C.H.T., H.T., G.T., J. Tremblay, N.V., V.V., S. Vogelezang, C. Wang, L. Wang, J.F.W., M.K.W., M. Wuttke, Y.X., Q.Y., L.M.Y.-A., W.Z. Bioinformatics: T.S.A., S. Akilesh, P.A., D.B., S.B., A.S.B., C.A.B., E.C., R.J.C., X.C., A.Y.C., M. Cocca, M.P.C., T.C., E.W.D., F.D., A. Dehghan, J. Divers, R.D., G.E., A.F., H.G., S.G., A.G., S.D.G., M. Gorski, P.H., I.M.H., E.H., A.H., K. Horn, J.J., N.S.J., C.-C.K., H. Kirsten, M.E.K., A. Krajcoviechova, A. Köttgen, C.D.L., B.L., M. Li, Y.L., Jianjun Liu, L.-P.L., J. Marten, J. Martins, Y.M., P.P.M., K.L.M., D.O.M.-K., P.J.v.d.M., R.M., W.M., R.N., T.N., S.A.P., N.P., G.P., A.I.P., M.H.P., B.P.P., F. Rizzi, R. Rueedi, Y. Saba, E.S., M. Scholz, C.-A.S., S. Sedaghat, C.M.S., K.B.S., A.V.S., C.N.S., S. Szymczak, H.T., J. Tremblay, C. Wang, J.F.W., M. Wuttke, Y.X., L.M.Y.-A., Z.Y., W.Z. Interpretation of results: T.S.A., E.d.A., C.A.B., C.-Y.C., K.D., J. Divers, R.D., K.E., M.F.F., J.F.F., B.I.F., S.G., C.G., A.G., M. Gorski, P.H., P.v.d.H., H.T., I.M.H., K. Ho, K. Horn, W.H., S.-J.H., B.J., H. Kirsten, W. Koenig, A. Krajcoviechova, A. Köttgen, M. Laakso, C.D.L., M. Li, Y.L., P.K.M., J. Marten, K. Miliku, K.L.M., A.P.M., N.D.P., C.P., S.A.P., B.P.P., D.F.R., M.R., P.M.R., M. Scholz, S. Sedaghat, K.B.S., C.N.S., P.O.S., B.O.T., A. Teumer, A. Tin, J. Tremblay, I.T., A.G.U., N.V., V.V., S. Vogelezang, L. Wallentin, H.W., M. Wuttke, Y.X., M.Y., L.M.Y.-A. Genotyping: N.A., D.B., R.B., A.S.B., C.A.B., A.C., H.C., D.I.C., C.-Y.C., E.W.D., A. Demirkan, R.D., C.M.v.D., G.E., M.K.E., M.F.F., A.F., Y.F., C.F., R.T.G., H.G., S.D.G., P.H., P.v.d.H., H.T., C.H., C.-K.H., W.H., E.I., C.-C.K., M.E.K., W. Koenig, J.S.K., P.K., A.T.K., A. Krajcoviechova, F.K., M. Kubo, M. Kähönen, A. Körner, L.A.L., T.L., L.-P.L., P.K.M., T.M., O.M., Y.M., K.L.M., N.M., G.W.M., D.O.M.-K., A.P.M., J.C.M., W.M., M.A.N., M.O.-M., S.P., N.D.P., B.W.P., M. Perola, D.J.P., M.H.P., O.T.R., D.F.R., F. Rivadeneira, F. Rizzi, J.I.R., D.R., V.S., E.S., B.H.S., C.N.S., S.M.T., K.T., A. Teumer, D.T., J. Tremblay, A.G.U., S. Vaccargiu, U.V., M. Waldenberger, C. Wang, L. Wang, Y.X.W., J.G.W., M.K.W., A.B.Z., J.Ä. Critical review of manuscript: T.S.A., S. Akilesh, P.A., E.d.A., S.J.B., N.B., M.L.B., G.B., M.H.d.B., E.P.B., T.S.B., H.B., A.S.B., C.A.B., H.C., D.I.C., X.C., Y.C., A.Y.C., M. Ciullo, J.C., R.M.v.D., G.D., O.D., J. Divers, R.D., K.-U.E., D.R.V.E., T.L.E., P.E., K.E., M.K.E., M.F.F., J.F.F., O.H.F., A.F., B.I.F., Y.F., C.F., H.G., S.G., C.G., A.G., S.D.G., M. Gorski, D.F.G., P.H., T.B.H., P.v.d.H., C.A.H., C.H., I.M.H., J.N.H., C.-K.H., K. Ho, A.H., W.H., N.H.-K., S.-J.H., O.S.I., E.I., V.W.J., J.J., J.B.J., P.K.J., B.J., M. Kastarinen, S.M.K., M.E.K., W. Koenig, A.T.K., H. Kramer, F.K., B.K.K., M. Kuokkanen, M. Kähönen, A. Körner, A. Köttgen, B.K., M. Laakso, L.A.L., C.D.L., J.J.-M.L., T.L., M. Li, Y.L., W.L., L.L., C.M.L., M. Loeffler, R.J.L., L.-P.L., P.K.M., A. Mahajan, J. Marten, N.G.M., D.M., C.M., T.M., O.M., E.K.M., K. Miliku, K.L.M., G.W.M., D.O.M.-K., R.d.M., W.M., G.N.N., M.A.N., M.N., K.N., B.N., I.M.N., R.N., T.N., M.L.O’D., A.J.O., M.O.-M., N.D.P., R.P., A.P., C.P., S.A.P., B.W.P., M. Perola, O.P., M.H.P., B.P.P., B.M.P., T.J.R., L.M.R., O.T.R., R. Rettig, M.R., K.M.R., P.M.R., F. Rivadeneira, D.J.R., P.R., I.R., D.R., C.S., V.S., K.-U.S., M. Scholz, C.-A.S., N. Schupf, B.S., S. Sedaghat, K.B.S., X.S., A.V.S., H. Snieder, C.N.S., K. Strauch, G.S., P.O.S., S.M.T., N.Y.Q.T., B.O.T., A. Teumer, H.T., A. Tin, J. Tremblay, I.T., A. Tönjes, A.G.U., N.V., V.V., S. Vogelezang, A.P.d.V., U.V., M. Waldenberger, L. Wallentin, D.M.W., H.W., J.B.W., S.H.W., J.G.W., M. Wuttke, Q.Y., Z.Y., A.B.Z. Subject recruitment: S. Afaq, E.P.B., H.B., C.A.B., A.C., H.C., J.C.C., Miao-Ling Chee, K.C., R.C., M. Ciullo, D.C., K.D., M.K.E., V.H.X.F., B.I.F., R.T.G., V. Gudnason, C.A.H., W.H., N.H.-K., O.S.I., M.I., V.W.J., J.B.J., B.J., C.M.K., M. Kastarinen, J.S.K., A. Krajcoviechova, F.K., M. Kubo, M. Kähönen, A. Köttgen, M. Laakso, J.J.-M.L., T.L., W.L., L.L., N.G.M.,. K. Matsuda, C.M., A. Metspalu, R.d.M., W.M., K.N., M.L.O’D., I.O., A.J.O., S.P., C.P., S.A.P., B.W.P., M. Perola, O.P., B.P., D.J.P., T. Poulain, M.A.P., T.J.R., O.T.R., M.R., P.M.R., P.R., I.R., D.R., V.S., R.S., B.H.S., P.O.S., N.Y.Q.T., A. Teren, Y.-C.T., J. Tremblay, I.T., A. Tönjes, S. Vaccargiu, S. Vogelezang, P.V., A.P.d.V., G.W., L. Wallentin, H.W., J.B.W., S.H.W., J.G.W., A.B.Z., J.Ä.
Corresponding authors
Ethics declarations
Competing interests
W. Koenig reports modest consultation fees for advisory board meetings from Amgen, DalCor, Kowa, Novartis, Pfizer and Sanofi and modest personal fees for lectures from Amgen, AstraZeneca, Novartis, Pfizer and Sanofi, all outside the scope of the submitted work. W.M. is employed with Synlab Services and holds shares of Synlab Holding Deutschland. D.O.M.-K. is a part-time research consultant at Metabolon. M.A.N. is supported by a consulting contract between Data Tecnica International and the National Institute on Aging (NIA), National Institutes of Health (NIH) and consults for Illumina, the Michael J. Fox Foundation and University of California Healthcare. O.H.F. works in ErasmusAGE, a center for aging research across the life course funded by Nestlé Nutrition (Nestec); Metagenics; and AXA. K.B.S., L.Y.-A., D.M.W. and M.A.L. are full-time employees of GlaxoSmithKline. M.L.O’D. received grant support from GlaxoSmithKline, MSD, Eisai, AstraZeneca, MedCo and Janssen. H.W. received grants and non-financial support from GlaxoSmithKline, during the conduct of the study; grants from Sanofi-Aventis, Eli Lilly, the National Institute of Health, Omthera Pharmaceuticals, Pfizer New Zealand, Elsai Inc. and Dalcor Pharma UK; honoraria and non-financial support from AstraZeneca; and is on advisory boards for Sirtex and Acetilion and received personal fees from CSL Behring and American Regent outside the scope of the submitted work. L. Wallentin received institutional grants from GlaxoSmithKline, AstraZeneca, BMS, Boehringer-Ingelheim, Pfizer, MSD and Roche Diagnostics. D.F.R. and A.I.P. are employees of MSD. M. Scholz received consultancy of and grant support from Merck Serono not related to this project. B.M.P. serves on the DSMB of a clinical trial funded by the manufacturer (Zoll LifeCor) and on the steering committee of the Yale Open Data Access Project funded by Johnson & Johnson. J. Danesh is a member of the Novartis Cardiovascular and Metabolic Advisory Board and received grant support from Novartis. A.S.B. received grants from MSD, Pfizer, Novartis, Biogen and Bioverativ and personal fees from Novartis. V.S. has participated in a conference trip sponsored by Novo Nordisk and received a honorarium from the same source for participating in an advisory board meeting. A. Köttgen received grant support from Gruenenthal. All other authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–3 and Supplementary Figures 1 and 3–9
Rights and permissions
About this article
Cite this article
Wuttke, M., Li, Y., Li, M. et al. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nat Genet 51, 957–972 (2019). https://doi.org/10.1038/s41588-019-0407-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0407-x
This article is cited by
-
The causal relationship between COVID-19 and estimated glomerular filtration rate: a bidirectional Mendelian randomization study
BMC Nephrology (2024)
-
Rare copy-number variants as modulators of common disease susceptibility
Genome Medicine (2024)
-
Multivariate canonical correlation analysis identifies additional genetic variants for chronic kidney disease
npj Systems Biology and Applications (2024)
-
Unraveling the epigenetic code: human kidney DNA methylation and chromatin dynamics in renal disease development
Nature Communications (2024)
-
Predicting proximal tubule failed repair drivers through regularized regression analysis of single cell multiomic sequencing
Nature Communications (2024)