Abstract
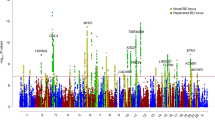
Bipolar disorder is a highly heritable psychiatric disorder. We performed a genome-wide association study (GWAS) including 20,352 cases and 31,358 controls of European descent, with follow-up analysis of 822 variants with P < 1 × 10−4 in an additional 9,412 cases and 137,760 controls. Eight of the 19 variants that were genome-wide significant (P < 5 × 10−8) in the discovery GWAS were not genome-wide significant in the combined analysis, consistent with small effect sizes and limited power but also with genetic heterogeneity. In the combined analysis, 30 loci were genome-wide significant, including 20 newly identified loci. The significant loci contain genes encoding ion channels, neurotransmitter transporters and synaptic components. Pathway analysis revealed nine significantly enriched gene sets, including regulation of insulin secretion and endocannabinoid signaling. Bipolar I disorder is strongly genetically correlated with schizophrenia, driven by psychosis, whereas bipolar II disorder is more strongly correlated with major depressive disorder. These findings address key clinical questions and provide potential biological mechanisms for bipolar disorder.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The PGC’s policy is to make genome-wide summary results public. Summary statistics for our meta-analysis are available through the PGC (https://www.med.unc.edu/pgc/results-and-downloads). Data are accessible with collaborative analysis proposals through the Bipolar Disorder Working Group of the PGC (https://med.unc.edu/pgc).
References
Ferrari, A. J. et al. The prevalence and burden of bipolar disorder: findings from the global burden of disease study 2013. Bipolar Disord. 18, 440–450 (2016).
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
Edvardsen, J. et al. Heritability of bipolar spectrum disorders. Unity or heterogeneity? J. Affect. Disord. 106, 229–240 (2008).
McGuffin, P. et al. The heritability of bipolar affective disorder and the genetic relationship to unipolar depression. Arch. Gen. Psychiatry 60, 497–502 (2003).
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders 5th edn (American Psychiatric Association Publishing, 2013).
World Health Organization. The ICD-10 Classification of Mental and Behavioural Disorders: Clinical Descriptions and Diagnostic Guidelines (World Health Organization, 1992).
Craddock, N. & Owen, M. J. The Kraepelinian dichotomy—going, going… but still not gone. Br. J. Psychiatry 196, 92–95 (2010).
Lee, S. H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Sklar, P. et al. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat. Genet. 43, 977–983 (2011).
Baum, A. E. et al. A genome-wide association study implicates diacylglycerol kinase eta (DGKH) and several other genes in the etiology of bipolar disorder. Mol. Psychiatry 13, 197–207 (2008).
Charney, A. W. et al. Evidence for genetic heterogeneity between clinical subtypes of bipolar disorder. Transl. Psychiatry 7, e993 (2017).
Chen, D. T. et al. Genome-wide association study meta-analysis of European and Asian-ancestry samples identifies three novel loci associated with bipolar disorder. Mol. Psychiatry 18, 195–205 (2013).
Cichon, S. et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am. J. Hum. Genet. 88, 372–381 (2011).
Ferreira, M. A. R. et al. Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat. Genet. 40, 1056–1058 (2008).
Green, E. K. et al. Association at SYNE1 in both bipolar disorder and recurrent major depression. Mol. Psychiatry 18, 614–617 (2013).
Green, E. K. et al. Replication of bipolar disorder susceptibility alleles and identification of two novel genome-wide significant associations in a new bipolar disorder case-control sample. Mol. Psychiatry 18, 1302–1307 (2013).
Hou, L. et al. Genome-wide association study of 40,000 individuals identifies two novel loci associated with bipolar disorder. Hum. Mol. Genet. 25, 3383–3394 (2016).
Mühleisen, T. W. et al. Genome-wide association study reveals two new risk loci for bipolar disorder. Nat. Commun. 5, 3339 (2014).
Schulze, T. G. et al. Two variants in Ankyrin 3 (ANK3) are independent genetic risk factors for bipolar disorder. Mol. Psychiatry 14, 487–491 (2009).
Scott, L. J. et al. Genome-wide association and meta-analysis of bipolar disorder in individuals of European ancestry. Proc. Natl Acad. Sci. USA 106, 7501–7506 (2009).
Sklar, P. et al. Whole-genome association study of bipolar disorder. Mol. Psychiatry 13, 558–569 (2008).
Smith, E. N. et al. Genome-wide association study of bipolar disorder in European American and African American individuals. Mol. Psychiatry 14, 755–763 (2009).
Burton, P. R. et al. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Gratten, J., Wray, N. R., Keller, M. C. & Visscher, P. M. Large-scale genomics unveils the genetic architecture of psychiatric disorders. Nat. Neurosci. 17, 782–790 (2014).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2017).
Palmer, C. & Pe’er, I. Statistical correction of the winner’s curse explains replication variability in quantitative trait genome-wide association studies. PLoS Genet. 13, e1006916 (2017).
Zhong, H. & Prentice, R. L. Bias-reduced estimators and confidence intervals for odds ratios in genome-wide association studies. Biostatistics 9, 621–634 (2008).
Holland, D. et al. Beyond SNP heritability: polygenicity and discoverability estimated for multiple phenotypes with a univariate gaussian mixture model. Preprint at biorXiv https://doi.org/10.1101/133132 (2018).
Gao, X., Starmer, J. & Martin, E. R. A multiple testing correction method for genetic association studies using correlated single nucleotide polymorphisms. Genet. Epidemiol. 32, 361–369 (2008).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Okbay, A. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 48, 624–633 (2016).
Cross-Disorder Group of the Psychiatric Genomics Consortium et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Duncan, L. et al. Significant locus and metabolic genetic correlations revealed in genome-wide association study of anorexia nervosa. Am. J. Psychiatry 174, 850–858 (2017).
Otowa, T. et al. Meta-analysis of genome-wide association studies of anxiety disorders. Mol. Psychiatry 21, 1391–1399 (2016).
Gale, C. R. et al. Pleiotropy between neuroticism and physical and mental health: findings from 108 038 men and women in UK Biobank. Transl. Psychiatry 6, e791 (2016).
Rietveld, C. A. et al. GWAS of 126,559 individuals identifies genetic variants associated with educational attainment. Science 340, 1467–1471 (2013).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Benyamin, B. et al. Childhood intelligence is heritable, highly polygenic and associated with FNBP1L. Mol. Psychiatry 19, 253–258 (2014).
Sniekers, S. et al. Genome-wide association meta-analysis of 78,308 individuals identifies new loci and genes influencing human intelligence. Nat. Genet. 49, 1107–1112 (2017).
MacArthur, J. et al. The new NHGRI-EBI catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 45, D896–D901 (2017).
Ikeda, M. et al. A genome-wide association study identifies two novel susceptibility loci and trans population polygenicity associated with bipolar disorder. Mol. Psychiatry 23, 639–647 (2018).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Pavlides, J. M. W. et al. Predicting gene targets from integrative analyses of summary data from GWAS and eQTL studies for 28 human complex traits. Genome Med. 8, 84 (2016).
Qi, T. et al. Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat. Commun. 9, 2282 (2018).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Bipolar Disorder and Schizophrenia Working Group of the Psychiatric Genomics Consortium. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715.e16 (2018).
Lee, S. H., Goddard, M. E., Wray, N. R. & Visscher, P. M. A better coefficient of determination for genetic profile analysis. Genet. Epidemiol. 36, 214–224 (2012).
Simonsen, C. et al. Neurocognitive dysfunction in bipolar and schizophrenia spectrum disorders depends on history of psychosis rather than diagnostic group. Schizophr. Bull. 37, 73–83 (2011).
Goes, F. S. et al. Genome-wide association of mood-incongruent psychotic bipolar disorder. Transl. Psychiatry 2, e180 (2012).
Allardyce, J. et al. Association between schizophrenia-related polygenic liability and the occurrence and level of mood-incongruent psychotic symptoms in bipolar disorder. JAMA Psychiatry 75, 28–35 (2018).
Palmer, C. & Pe’er, I. Statistical correction of the winner’s curse explains replication variability in quantitative trait genome-wide association studies. PLoS Genet. 13, e1006916 (2017).
Goes, F. S. et al. Genome-wide association study of schizophrenia in Ashkenazi Jews. Am. J. Med. Genet. B 168, 649–659 (2015).
Gaspar, H. A. & Breen, G. Drug enrichment and discovery from schizophrenia genome-wide association results: an analysis and visualisation approach. Sci. Rep. 7, 12460 (2017).
Camandola, S. & Mattson, M. P. Aberrant subcellular neuronal calcium regulation in aging and alzheimer’s disease. Biochim. Biophys. Acta Mol. Cell Res. 1813, 965–973 (2011).
Mertens, J. et al. Differential responses to lithium in hyperexcitable neurons from patients with bipolar disorder. Nature 527, 95–99 (2015).
Yang, C.-P. et al. Comprehensive integrative analyses identify GLT8D1 and CSNK2B as schizophrenia risk genes. Nat. Commun. 9, 838 (2018).
Lee, S.-H., Zabolotny, J. M., Huang, H., Lee, H. & Kim, Y.-B. Insulin in the nervous system and the mind: functions in metabolism, memory, and mood. Mol. Metab. 5, 589–601 (2016).
McIntyre, R. S. et al. A randomized, double-blind, controlled trial evaluating the effect of intranasal insulin on neurocognitive function in euthymic patients with bipolar disorder. Bipolar Disord. 14, 697–706 (2012).
Desfossés, J., Stip, E., Bentaleb, L. A. & Potvin, S. Endocannabinoids and schizophrenia. Pharmaceuticals 3, 3101–3126 (2010).
Zamberletti, E., Rubino, T. & Parolaro, D. The endocannabinoid system and schizophrenia: integration of evidence. Curr. Pharm. Des. 18, 4980–4990 (2012).
Serra, G. & Fratta, W. A possible role for the endocannabinoid system in the neurobiology of depression. Clin. Pract. Epidemiol. Ment. Health 3, 25 (2007).
MacCabe, J. H. et al. Excellent school performance at age 16 and risk of adult bipolar disorder: national cohort study. Br. J. Psychiatry 196, 109–115 (2010).
Vreeker, A. et al. High educational performance is a distinctive feature of bipolar disorder: a study on cognition in bipolar disorder, schizophrenia patients, relatives and controls. Psychol. Med. 46, 807–818 (2016).
Ripke, S. Ricopili: a Tool for Visualizing Regions of Interest in Select GWAS Data Sets (Broad Institute, 2014).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: Polygenic Risk Score software. Bioinformatics 31, 1466–1468 (2015).
Finucane, H. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
O’Dushlaine, C. et al. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat. Neurosci. 18, 199–209 (2015).
1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Olier, I., Vellido, A. & Giraldo, J. Kernel generative topographic mapping. In Proc. 18th European Symposium on Artificial Neural Networks (ed. Verleysen, M.) 481–486 (ESANN, 2010).
Acknowledgements
This paper is dedicated to the memory of Psychiatric Genomics Consortium (PGC) founding member and Bipolar Disorder Working Group co-chair Pamela Sklar. We thank the participants who donated their time, experiences and DNA to this research, and the clinical and scientific teams that worked with them. We are deeply indebted to the investigators who comprise the PGC. Analyses were carried out on the NL Genetic Cluster Computer (http://www.geneticcluster.org), hosted by SURFsara, and the Mount Sinai high performance computing cluster (http://hpc.mssm.edu). PGC members have received major funding from the US National Institute of Mental Health. This work was funded in part by the Brain and Behavior Research Foundation, Stanley Medical Research Institute, University of Michigan, Pritzker Neuropsychiatric Disorders Research Fund L.L.C., Marriot Foundation and the Mayo Clinic Center for Individualized Medicine, the NIMH Intramural Research Program; Canadian Institutes of Health Research; the UK Maudsley NHS Foundation Trust, NIHR, NRS, MRC, Wellcome Trust; European Research Council; German Ministry for Education and Research, German Research Foundation IZKF of Münster, Deutsche Forschungsgemeinschaft, ImmunoSensation, the Dr Lisa-Oehler Foundation, University of Bonn; the Swiss National Science Foundation; French Foundation FondaMental and ANR; Spanish Ministerio de Economía, CIBERSAM, Industria y Competitividad, European Regional Development Fund (ERDF), Generalitat de Catalunya, EU Horizon 2020 Research and Innovation Programme; BBMRI-NL; South-East Norway Regional Health Authority and Mrs Throne-Holst; Swedish Research Council, Stockholm County Council, Söderström Foundation; Lundbeck Foundation, Aarhus University; Australia NHMRC, NSW Ministry of Health, Janette M. O’Neil and Betty C. Lynch. The views expressed are those of the authors and not necessarily those of their institutions or any funding or regulatory bodies. Additional acknowledgements, including funding sources, are presented in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Writing group: E.A.S., G.B., A.J.F., A. McQuillin, S.R., J.R.I.C., N.M., N.R.W., A.D.F., H.J.E., S.C., R.A.O., L.J.S., O.A.A. and J.K. PGC BD principal investigator group: E.A.S., G.B., A.J.F., A. McQuillin, D. Curtis, R.H.P., R.A., I.A., M.A., L.B., B.T.B., F.B., W.H.B., J.M.B., D.H.R.B., M. Boehnke, A.D.B., A.C., N.C., U.D., T. Esko, B.E., M. Frye, J.M.F., E.S.G., M.G., F.G., M.G.-S., J.H., D.M.H., C.M.H., I.J., L.A.J., R.S.K., M. Landén, M. Leboyer, C.M.L., Q.S.L., J. Lissowska, N.G.M., S.L.M., A.M.M., F.J.M., I.M., A. Metspalu, P.B. Mitchell, G.M., O.M., P.B. Mortensen, B.M.-M., R.M.M., B.M.N., V.N., M.N., M.M.N., M.C.O’D., K.J.O., M.J.O., S.A.P., C.P., M.T.P., J.A.R.-Q., M. Ribasés, M. Rietschel, G.A.R., M. Schalling, P.R.S., T.G.S., A.S., J.W.S., H.S., K.S., E. Stordal, G.T., A.E.V., E.V., J.B.V., T.W., J.I.N., A.D.F., H.J.E., S.C., R.A.O., L.J.S., O.A.A., J.K. and P.S. Bioinformatics: E.A.S., G.B., A.J.F., J.R.I.C., H.A.G., P.A.H., S.E.B., D.W.C., V.E.-P., C.G., M.L.H., S.H., R. Karlsson, M. Leber, C. Liu, F. Meng, L.M.O.L., A.P.S.O., C.S.R., P.R., P.D.S., M. Steffens, S. Szelinger, T.E.T., S.X., P. Zandi, eQTLGen Consortium, BIOS Consortium, T. Esko, E.S.G., Q.S.L., G.A.R. and H.S. Clinical: A. McQuillin, M.M., E.A., N.A.-R., A.A., N.B., M. Bauer, C.B.P., E.B., M.P.B., M. Budde, M. Burmeister, W. Byerley, M.C., P.C., W.C., D. Curtis, P.M.C., J.R.D., T. Elvsåshagen, L. Forty, C.F., K.G., J. Garnham, M.G.P., K.G.-S., M.J.G., J. Grove, J.G.-P., M. Hautzinger, U.H., M. Hipolito, A.J., J.L.K., S.K.-S., M.K., R. Kupka, C. Lavebratt, J. Lawrence, W.B.L., S.L., D.J.M., P.B. Mahon, W.M., L. Martinsson, P.M., M.G.M., H.M., A.N.A., E.A.N., C.O’D., L.O., U.Ö., R.H.P., A. Perry, A. Pfennig, J.B.P., E.J.R., A.R., J.P.R., F.R., M. Rivera, W.A.S., C.S.W., E. Sigurdsson, C.S., O.B.S., J.L.S., A.T.S., D.S.C., J.S.S., F.S., J.S., R.C.T., H.V., T.W.W., A.H.Y., R.A., I.A., M.A., B.T.B., F.B., D.H.R.B., A.D.B., A.C., N.C., U.D., B.E., M. Frye, E.S.G., M.G., F.G., M.G.-S., J.H., I.J., L.A.J., R.S.K., G.K., M. Landén, J. Lissowska, N.G.M., F. Mayoral, S.L.M., A.M.M., F.J.M., I.M., P.B. Mitchell, G.M., O.M., P.B. Mortensen, V.N., M.C.O’D., K.J.O., M.J.O., C.P., M.T.P., J.A.R.-Q., M. Rietschel, T.G.S., A.S., J.W.S., E. Stordal, A.E.V., E.V., J.I.N. and A.D.F. Genomic assays/data generation: G.B., A.J.F., E.A., D.A., M.B.-H., C.B.P., J.B.-G., T.-K.C., D.W.C., C. Cruceanu, F.D., J.D.-F., S.D., S.B.F., J.F., M.G.P., E.K.G., P.H., S.J., R. Kandaswamy, A.C.K., S.E.L., A. Maaser, J.D.M., L. Milani, G.W.M., D.W.M., T.W.M., E.R., C.S.-M., T.S., C.S.H., S. Szelinger, J.T., S.H.W., P. Zhang, A.C., T. Esko, J.M.F., E.S.G., M.G., D.M.H., R.M.M., M.M.N., M. Ribasés, G.A.R., G.T. and S.C. Obtained funding for BD samples: G.B., H.A., N.A.-R., J.D.B., W. Bunney, J.R.D., N.B.F., L. Frisén, M.J.G., S.J., J.A.K., C. Lavebratt, S.L., P.M., G.W.M., U.Ö., J.B.P., M. Rivera, A.F.S., C.S.W., S.J.W., T.W.W., A.H.Y., M.A., B.T.B., W.H.B., J.M.B., M.Boehnke, A.D.B., A.C., N.C., M. Frye, J.M.F., E.S.G., M.G., M.G.-S., I.J., L.A.J., M. Landén, M. Leboyer, C.M.L., N.G.M., A. Metspalu, P.B. Mitchell, O.M., P.B. Mortensen, B.M.N., M.N., M.M.N., M.C.O’D., M.J.O., C.P., M.T.P., M. Rietschel, G.A.R., P.R.S., T.G.S., J.W.S., G.T., J.B.V., T.W., J.I.N., H.J.E., R.A.O. and P.S. Statistical analysis: E.A.S., G.B., S.R., V.T., M.M., Y.W., J.R.I.C., H.A.G., C.A.d.L., S. Steinberg, J.M.W.P., M.T., E.M.B., T.H.P., P.A.H., A.L.R., L.A., N.A.-R., T.D.A., V.A., S.A., J.A.B., R.B., S.E.B., J.B., F.C., K.C., A.W.C., D. Chen, C. Churchhouse, A.M.D., S.d.J., A.L.D., A.D., V.E.-P., C.C.F., M. Flickinger, T.M.F., D.G., C.G., J. Goldstein, S.D.G., T.A.G., J. Grove, W.G., M.L.H., D.H., L.H., J.S.J., R. Karlsson, M. Leber, P.H.L., J.Z.L., S.M., S.E.M., D.W.M., N.M., H.N., C.M.N., L.M.O.L., A.P.S.O., S.M.P., C.S.R., P.R., D.M.R., N.J.S., O.B.S., T.E.T., W.W., W.X., P. Zandi, P. Zhang, S.Z., eQTLGen Consortium, BIOS Consortium, J.M.B., A.D.B., M.J.D., E.S.G., F.G., Q.S.L., B.M.-M., D.P., H.S., P.F.S., N.R.W. and P.S.
Corresponding authors
Ethics declarations
Competing interests
T.E.T., S. Steinberg, H.S. and K.S. are employed by deCODE Genetics/Amgen. Multiple additional authors work for pharmaceutical or biotechnology companies in a manner directly analogous to academic co-authors and collaborators. A.H.Y. has given paid lectures and is on advisory boards for the following companies with drugs used in affective and related disorders: Astra Zeneca, Eli Lilly, Janssen, Lundbeck, Sunovion, Servier, Livanova. A.H.Y. is Lead Investigator for Embolden Study (Astra Zeneca), BCI Neuroplasticity Study and Aripiprazole Mania Study, which are investigator-initiated studies from Astra Zeneca, Eli Lilly, Lundbeck and Wyeth. J.N. is an investigator for Janssen. P.F.S. reports the following potentially competing interests: Lundbeck (advisory committee), Pfizer (Scientific Advisory Board member) and Roche (grant recipient, speaker reimbursement). G.B. reports consultancy and speaker fees from Eli Lilly and Illumina and grant funding from Eli Lilly. O.A.A. has received speaker fees from Lundbeck. J.A.R.-Q. was on the speakers’ bureau and/or acted as consultant for Eli Lilly, Janssen-Cilag, Novartis, Shire, Lundbeck, Almirall, Braingaze, Sincrolab and Rubió in the last 5 years. He also received travel awards (air tickets and hotel) for taking part in psychiatric meetings from Janssen-Cilag, Rubió, Shire and Eli Lilly. The Department of Psychiatry chaired by him received unrestricted educational and research support from the following companies in the last 5 years: Eli Lilly, Lundbeck, Janssen-Cilag, Actelion, Shire, Ferrer, Oryzon, Roche, Psious and Rubió. E.V. has received grants and served as consultant, advisor or CME speaker for the following entities: AB-Biotics, Abbott, Allergan, Angelini, Astra Zeneca, Bristol-Myers Squibb, Dainippon Sumitomo Pharma, Farmindustria, Ferrer, Forest Research Institute, Gedeon Richter, Glaxo-Smith-Kline, Janssen, Lundbeck, Otsuka, Pfizer, Roche, SAGE, Sanofi-Aventis, Servier, Shire, Sunovion, Takeda, the Brain and Behaviour Foundation, the Catalan Government (AGAUR and PERIS), the Spanish Ministry of Science, Innovation, and Universities (AES and CIBERSAM), the Seventh European Framework Programme and Horizon 2020 and the Stanley Medical Research Institute. T. Elvsåshagen has received speaker fees from Lundbeck. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and Supplementary Notes
Supplementary Tables
Supplementary Tables 1–18
Supplementary Data 1
Regional association plots for all 31 lead SNPs in the 30 genome-wide-significant loci.
Supplementary Data 2
Forest plots of the 32 primary GWAS datasets for all 31 lead SNPs in the 30 genome-wide-significant loci.
Supplementary Data 3
Forest plots of the 7 follow-up datasets for all 31 lead SNPs in the 30 genome-wide-significant loci.
Rights and permissions
About this article
Cite this article
Stahl, E.A., Breen, G., Forstner, A.J. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat Genet 51, 793–803 (2019). https://doi.org/10.1038/s41588-019-0397-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0397-8
This article is cited by
-
Causal relationships between atopic dermatitis and psychiatric disorders: a bidirectional two-sample Mendelian randomization study
BMC Psychiatry (2024)
-
Association of major depression, schizophrenia and bipolar disorder with thyroid cancer: a bidirectional two-sample mendelian randomized study
BMC Psychiatry (2024)
-
The causal relationship of female infertility and psychiatric disorders in the European population: a bidirectional two-sample Mendelian randomization study
BMC Women's Health (2024)
-
Whole-exome sequencing in UK Biobank reveals rare genetic architecture for depression
Nature Communications (2024)
-
Cortico-striatal differences in the epigenome in attention-deficit/ hyperactivity disorder
Translational Psychiatry (2024)