Abstract
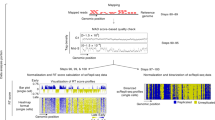
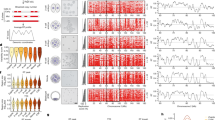
Here, we report a single-cell DNA replication sequencing method, scRepli-seq, a genome-wide methodology that measures copy number differences between replicated and unreplicated DNA. Using scRepli-seq, we demonstrate that replication-domain organization is conserved among individual mouse embryonic stem cells (mESCs). Differentiated mESCs exhibited distinct profiles, which were also conserved among cells. Haplotype-resolved scRepli-seq revealed similar replication profiles of homologous autosomes, while the inactive X chromosome was clearly replicated later than its active counterpart. However, a small degree of cell-to-cell replication-timing heterogeneity was present, which was smallest at the beginning and the end of S phase. In addition, developmentally regulated domains were found to deviate from others and showed a higher degree of heterogeneity, thus suggesting a link to developmental plasticity. Moreover, allelic expression imbalance was found to strongly associate with replication-timing asynchrony. Our results form a foundation for single-cell-level understanding of DNA replication regulation and provide insights into three-dimensional genome organization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All RT profiles (BrdU-IP, 100 cells and single-cell Repli-seq) and RNA-seq data generated in this study have been deposited in the NCBI Gene Expression Omnibus (GEO) database under accession code GSE108556.
References
Berezney, R., Dubey, D. D. & Huberman, J. A. Heterogeneity of eukaryotic replicons, replicon clusters, and replication foci. Chromosoma. 108, 471–484 (2000).
Rhind, N. & Gilbert, D. M. DNA Replication timing. Cold Spring Harb. Perspect. Biol. 5, 1–26 (2013).
Prioleau, M. N. & MacAlpine, D. M. DNA replication origins: where do we begin? Gene. Dev. 30, 1683–1697 (2016).
Rivera-Mulia, J. C. & Gilbert, D. M. Replication timing and transcriptional control: beyond cause and effect: part III. Curr. Opin. Cell Biol. 40, 168–178 (2016).
Hiratani, I. et al. Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res. 20, 155–169 (2010).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Ryba, T. et al. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 20, 761–770 (2010).
Ryba, T., Battaglia, D., Pope, B. D., Hiratani, I. & Gilbert, D. M. Genome-scale analysis of replication timing: from bench to bioinformatics. Nat. Protoc. 6, 870–895 (2011).
Hansen, R. S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl Acad. Sci. USA 107, 139–144 (2010).
Takebayashi, S.-I., Manders, E. M. M., Kimura, H., Taguchi, H. & Okumura, K. Mapping sites where replication initiates in mammalian cells using dna fibers. Exp. Cell Res. 271, 263–268 (2001).
Norio, P. et al. Progressive activation of DNA replication initiation in large domains of the immunoglobulin heavy chain locus during B cell development. Mol. Cell 20, 575–587 (2005).
Lebofsky, R., Heilig, R., Sonnleitner, M., Weissenbach, J. & Bensimon, A. DNA replication origin interference increases the spacing between initiation events in human cells. Mol. Biol. Cell. 17, 5337–5345 (2006).
Dileep, V. & Gilbert, D. M. Single-cell replication profiling to measure stochastic variation in mammalian replication timing. Nat. Commun. 9, 427 (2018).
Hiratani, I. et al. Global reorganization of replication domains during embryonic stem cell differentiation. PLoS Biol. 6, 2220–2236 (2008).
Woodfine, K. et al. Replication timing of the human genome. Hum. Mol. Genet. 13, 191–202 (2004).
Koren, A. et al. Genetic variation in human DNA replication timing. Cell 159, 1015–1026 (2014).
Mukhopadhyay, R. et al. Allele-specific genome-wide profiling in human primary erythroblasts reveal replication program organization. PLoS Genet. 10, e1004319 (2014).
Jiang, X. R. et al. Telomerase expression in human somatic cells does not induce changes associated with a transformed phenotype. Nat. Genet. 21, 111–114 (1999).
Baslan, T. et al. Optimizing sparse sequencing of single cells for highly multiplex copy number profiling. Genome Res. 125, 714–724 (2015).
Morgani, S., Nichols, J. & Hadjantonakis, A.-K. The many faces of pluripotency: in vitro adaptations of a continuum of in vivo states. BMC Dev. Biol. 17, 7 (2017).
Takahashi, S., Kobayashi, S. & Hiratani, I. Epigenetic differences between naïve and primed pluripotent stem cells. Cell. Mol. Life Sci. 75, 1191–1203 (2018).
Rivera-Mulia, J. C. et al. Allele-specific control of replication timing and genome organization during development. Genome Res. 28, 800–811 (2018).
Dileep, V. et al. Topologically associating domains and their long-range contacts are established during early G1 coincident with the establishment of the replication timing program. Genome Res. 25, 1104–1113 (2015).
Besnard, E. et al. Unraveling cell type-specific and reprogrammable human replication origin signatures associated with G-quadruplex consensus motifs. Nat. Struct. Mol. Biol. 19, 837–844 (2012).
Murakami, K., Araki, K., Ohtsuka, S., Wakayama, T. & Niwa, H. Choice of random rather than imprinted X inactivation in female embryonic stem cell-derived extra-embryonic cells. Development 138, 197–202 (2011).
Takada, T. et al. The ancestor of extant Japanese fancy mice contributed to the mosaic genomes of classical inbred strains. Genome Res. 23, 1329–1338 (2013).
Taylor, J. H. Asynchronous duplication of chromosomes in cultured cells of chinese hamster. J. Cell. Biol. 7, 455–463 (1960).
Morishima, A., Grumbach, M. M. & Taylor, J. H. Asynchronous duplication of human chromosomes and the origin of sex chromatin. Proc. Natl Acad. Sci. USA 48, 756–763 (1962).
Hiratani, I. & Gilbert, D. M. Autosomal lyonization of replication domains during early mammalian development. Adv. Exp. Med. Biol. 695, 41–58 (2010).
Hiratani, I., Takebayashi, S., Lu, J. & Gilbert, D. M. Replication timing and transcriptional control: beyond cause and effect-part II. Curr. Opin. Genet. Dev. 19, 142–149 (2009).
Gimelbrant, A. A. & Chess, A. An epigenetic state associated with areas of gene duplication. Genome Res. 16, 723–729 (2006).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Flyamer, I. M. et al. Single-cell Hi-C reveals unique chromatin reorganization at oocyte-tozygote transition. Nature 544, 110–114 (2017).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Takebayashi, S. I., Ogata, M. & Okumura, K. Anatomy of mammalian replication domains. Genes 8, 110 (2017).
Yamazaki, S., Hayano, M. & Masai, H. Replication timing regulation of eukaryotic replicons: Rif1 as a global regulator of replication timing. Trends Genet. 29, 449–460 (2013).
Niwa, H. How is pluripotency determined and maintained? Development 134, 635–646 (2007).
Meshorer, E. et al. Hyperdynamic plasticity of chromatin proteins in pluripotent embryonic stem cells. Dev. Cell. 10, 105–116 (2006).
Fussner, E. et al. Constitutive heterochromatin reorganization during somatic cell reprogramming. EMBO J. 30, 1778–1789 (2011).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Kind, J. et al. Genome-wide maps of nuclear lamina interactions in single human cells. Cell 163, 134–147 (2015).
Das, S. P. et al. Replication timing is regulated by the number of MCMs loaded at origins. Genome Res. 25, 1886–1892 (2015).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Dimitrova, D. S. & Gilbert, D. M. The spatial position and replication timing of chromosomal domains are both established in early G1 phase. Mol. Cell 4, 983–993 (1999).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Nora, E. P. et al. Targeted degradation of ctcf decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 3, e157 (2005).
Pope, B. D. et al. Replication-timing boundaries facilitate cell-type and species-specific regulation of a rearranged human chromosome in mouse. Hum. Mol. Genet. 21, 4162–4170 (2012).
Macaulay, I. C., Ponting, C. P. & Voet, T. Single-cell multiomics: multiple measurements from single cells. Trends Genet. 33, 155–168 (2017).
Calaway, J. D. et al. Genetic architecture of skewed X inactivation in the laboratory mouse. PLoS Genet. 9, e1003853 (2013).
Hayashi, K. & Saitou, M. Generation of eggs from mouse embryonic stem cells and induced pluripotent stem cells. Nat. Protoc. 8, 1513–1524 (2013).
Eiraku, M. & Sasai, Y. Mouse embryonic stem cell culture for generation of three-dimensional retinal and cortical tissues. Nat. Protoc. 7, 69–79 (2012).
Rathjen, J. & Rathjen, P. D. Lineage specific differentiation of mouse ES cells: formation and differentiation of early primitive ectoderm-like (EPL) cells. Methods Enzymol. 365, 3–25 (2003).
Kadota, M. et al. CTCF binding landscape in jawless fish with reference to Hox cluster evolution. Sci. Rep. 7, 4957 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17, 10–12 (2011).
Sakata, Y. et al. Defects in dosage compensation impact global gene regulation in the mouse trophoblast. Development 144, 2784–2797 (2017).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Bakker, B. et al. Single-cell sequencing reveals karyotype heterogeneity in murine and human malignancies. Genome. Biol. 17, 1–15 (2016).
Cayrou, C. et al. The chromatin environment shapes DNA replication origin organization and defines origin classes. Genome Res. 25, 1873–1885 (2015).
Acknowledgements
We thank S. Kuraku and members of his laboratory for assistance with NGS, F. Matsuzaki for the use of FACS and A. Tanigawa and Y. Kondo for technical assistance. We also thank D. M. Gilbert for exchanging unpublished observations; H. Niwa and K. Araki for CBMS1 mESCs; and B. D. Pope, M. Sugimoto and H. Masai for helpful discussions. This work was supported by a RIKEN CDB/BDR intramural grant to I.H., the Special Postdoctoral Researcher (SPDR) Program of RIKEN to S.T. and a Grant-in-Aid for Scientific Research on Innovative Areas (16H01405) to S-i.T., 18H05530 to I.H., 18K14681 to S.T., 15K06942, 15H01462 and 17H06426 to K.N., from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT).
Author information
Authors and Affiliations
Contributions
S.T., H.M., T.S., S-i.T. and I.H. conceived the project. S.T., T.S. and S-i.T. developed and conducted scRepli-seq and BrdU-IP experiments. S.T. and I.H. performed mESC culture, differentiation and sample collection. T.S. and S-i.T. performed hTERT-RPE1 cell culture and sample collection. K.N. and C.O. constructed diploid reference genome and helped with the haplotype-resolved analysis pipeline setup. H.M. and S-i.T. performed bioinformatics analyses. K.O. and M.O. supported for the design and execution of the project. S.T., H.M., S-i.T. and I.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9
Rights and permissions
About this article
Cite this article
Takahashi, S., Miura, H., Shibata, T. et al. Genome-wide stability of the DNA replication program in single mammalian cells. Nat Genet 51, 529–540 (2019). https://doi.org/10.1038/s41588-019-0347-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0347-5
This article is cited by
-
scAbsolute: measuring single-cell ploidy and replication status
Genome Biology (2024)
-
Emergence of replication timing during early mammalian development
Nature (2024)
-
Polycomb repressive complexes 1 and 2 are each essential for maintenance of X inactivation in extra-embryonic lineages
Nature Cell Biology (2023)
-
Replication dynamics identifies the folding principles of the inactive X chromosome
Nature Structural & Molecular Biology (2023)
-
Replication stress generates distinctive landscapes of DNA copy number alterations and chromosome scale losses
Genome Biology (2022)