Abstract
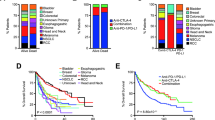
Immune checkpoint inhibitor (ICI) treatments benefit some patients with metastatic cancers, but predictive biomarkers are needed. Findings in selected cancer types suggest that tumor mutational burden (TMB) may predict clinical response to ICI. To examine this association more broadly, we analyzed the clinical and genomic data of 1,662 advanced cancer patients treated with ICI, and 5,371 non-ICI-treated patients, whose tumors underwent targeted next-generation sequencing (MSK-IMPACT). Among all patients, higher somatic TMB (highest 20% in each histology) was associated with better overall survival. For most cancer histologies, an association between higher TMB and improved survival was observed. The TMB cutpoints associated with improved survival varied markedly between cancer types. These data indicate that TMB is associated with improved survival in patients receiving ICI across a wide variety of cancer types, but that there may not be one universal definition of high TMB.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Data necessary to reproduce the figures are provided in Supplementary Data. All data are publicly available at http://www.cbioportal.org/study?id=tmb_mskcc_2018.
References
Callahan, M. K., Postow, M. A. & Wolchok, J. D. Targeting T cell co-receptors for cancer therapy. Immunity 44, 1069–1078 (2016).
Powles, T. et al. Atezolizumab versus chemotherapy in patients with platinum-treated locally advanced or metastatic urothelial carcinoma (IMvigor211): a multicentre, open-label, phase 3 randomised controlled trial. Lancet 391, 748–757 (2018).
Carbone, D. P. et al. First-line nivolumab in stage IV or recurrent non-small-cell lung cancer. N. Engl. J. Med. 376, 2415–2426 (2017).
Cohen, E. E. et al. LBA45_PR Pembrolizumab (pembro) vs standard of care (SOC) for recurrent or metastatic head and neck squamous cell carcinoma (R/M HNSCC): Phase 3 KEYNOTE-040 trial. Ann. Oncol. 28, mdx440.040 (2017).
Bendell, J. et al. LBA-004 Efficacy and safety results from IMblaze370, a randomised Phase III study comparing atezolizumab + cobimetinib and atezolizumab monotherapy vs regorafenib in chemotherapy-refractory metastatic colorectal cancer. Ann. Oncol. 29, mdy208.003 (2018).
Gibney, G. T., Weiner, L. M. & Atkins, M. B. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 17, e542–e551 (2016).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Cheng, D. T. et al. Memorial Sloan Kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Jordan, E. J. et al. Prospective comprehensive molecular characterization of lung adenocarcinomas for efficient patient matching to approved and emerging therapies. Cancer Discov. 7, 596–609 (2017).
Janjigian, Y. Y. et al. Genetic predictors of response to systemic therapy in esophagogastric cancer. Cancer Discov. 8, 49–58 (2018).
Rizvi, H. et al. Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J. Clin. Oncol. 36, 633–641 (2018).
Bouffet, E. et al. Immune checkpoint inhibition for hypermutant glioblastoma multiforme resulting from germline biallelic mismatch repair deficiency. J. Clin. Oncol. 34, 2206–2211 (2016).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Frampton, G. M. et al. Assessment of tumor mutation burden from >60,000 clinical cancer patients using comprehensive genomic profiling. J. Clin. Oncol. 34, 11558 (2016).
Johnson, D. B. et al. Targeted next generation sequencing identifies markers of response to PD-1 blockade. Cancer Immunol. Res. 4, 959–967 (2016).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Chalmers, Z. R. et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 9, 34 (2017).
Roszik, J. et al. Novel algorithmic approach predicts tumor mutation load and correlates with immunotherapy clinical outcomes using a defined gene mutation set. BMC Med. 14, 168 (2016).
Chowell, D. et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 62, eaao4572 (2017).
Balachandran, V. P. et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 551, 512–516 (2017).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012).
Segal, N. H. et al. Epitope landscape in breast and colorectal cancer. Cancer Res. 68, 889–892 (2008).
Verdegaal, E. M. E. et al. Neoantigen landscape dynamics during human melanoma-T cell interactions. Nature 536, 91–95 (2016).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Acknowledgements
We gratefully acknowledge the patients in this study and their families. We thank M. Gonen for statistical advice. We thank members of the Molecular Diagnostics Service in the Department of Pathology and the Marie-Josée and Henry R. Kravis Center for Molecular Oncology. We acknowledge funding sources, including the AACR-AstraZeneca Immunotherapy Fellowship (R.M.S.), Pershing Square Sohn Cancer Research grant (T.A.C.), the PaineWebber Chair (T.A.C.), Stand Up To Cancer (T.A.C.), NIH R01 CA205426, the STARR Cancer Consortium (T.A.C.), NCI R35 CA232097 (T.A.C.), Precision Immunotherapy Kidney Cancer Fund (T.A.C., R.J.M.), The Frederick Adler Fund, Cycle for Survival, NIH K08 DE024774 and R01 DE027738 (L.G.T.M.), and MSKCC through NIH/NCI Cancer Center Support Grant (P30 CA008748).
Author information
Authors and Affiliations
Contributions
R.M.S., R.S., T.A.C., N.R. and L.G.T.M. performed the analyses. C.-H.L., A.N.S., M.D.H., Y.Y.J., D.A.B., S.M.K. and E.J.J. provided clinical annotations. A.Z., M.F.B., D.B.S., M.L. and J.B. established the assays and coordinated data collection. C.-H.L., A.N.S., M.D.H., Y.Y.J., R.J.M., G.J.R., C.A.B., S.B., P.R., A.O., H.A.A., J.R., D.F.B., A.A.H., B.H.B., M.H.V., C.M.R., H.I.S., A.L.H., D.G.P., N.L., R.J.W., V.T., P.H.G., C.W.B., L.M.D., I.K.M., J.E.C., R.Y., N.H.S., R.P.D., L.S., J.D.W., C.E.A., Z.K.S., W.D.T., M.M.G., N.A.R., S.P.D., J.B., P.R., G.Y.K., T.J.K., C.A.K., G.I. and D.B.S. contributed to sample acquisition and patient recruitment. L.G.T.M., D.B.S., N.R. and T.A.C. supervised the study. L.G.T.M., T.A.C., C.-H.L., A.N.S., M.D.H. and R.M.S. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
R.M.S., T.A.C. and L.G.T.M. are inventors on a provisional patent application (62/569,053) filed by Memorial Sloan Kettering (MSK) relating to the use of TMB in cancer immunotherapy. M.D.H., N.A.R. and T.A.C. are inventors on a PCT patent application (PCT/US2015/062208) filed by MSK relating to the use of TMB in lung cancer immunotherapy. MSK and the inventors may receive a share of commercialization revenue from license agreements relating to these patent applications. C.-H.L. received research funding from Eisai, BMS, Exelixis, Pfizer and Calithera, and consulting fees from Exelixis and Eisai. A.N.S. has received research support from Bristol Myers Squibb, Immunocore, Astra-Zeneca and Xcovery and serves on the advisory board for Bristol Myers Squibb, Immunocore and Castle Biosciences; he also receives royalties from UpToDate. M.D.H. receives research funding from Bristol-Myers Squibb and is a paid consultant to Merck, Bristol-Myers Squibb, AztraZeneca, Genentech/Roche, Janssen, Nektar, Syndax, Mirati and Shattuck Labs. Y.Y.J. received research funding from Boehringer Ingelheim, Bayer, Genentech/Roche, Bristol-Myers Squibb, Eli Lilly and Merck, and served on advisory boards for Merck Serono, Bristol-Myers Squibb, Eli Lilly, Pfizer, Bayer, Imugene and Merck. S.B. currently works for Flatiron Health, which is a for-profit company. R.J.M. received research support from Pfizer, Genentech/Roche, Bristol Myers Squibb and Eisai, and consulting fees from Pfizer, Genentech/Roche, Merck, Incyte, Novartis, Eisai and Exelixis. M.H.V. received commercial research grants from Bristol-Myers Squibb and Genentech/Roche; honoraria from Novartis; travel/accommodation from Eisai, Novartis and Takeda; consultant/advisory board member for Alexion Pharmaceuticals, Bayer, Calithera Biosciences, Corvus Pharmaceuticals, Exelixis, Eisai, GlaxoSmithKline, Natera, Novartis and Pfizer. J.R. receives consulting fees from Merck, AstraZeneca, Bristol-Myers Squibb, EMD-Serono, Roche/Genentech, Sanofi, Seattle Genetics, Agensys, Bayer, Inovio, Lilly, Adicet Bio, Sensei, Chugai and Inovio. B.H.B. receives consulting fees from Genentech. G.J.R. received research funding from Novartis, Roche/Genentech, Millennium, GlaxoSmithKline, Pfizer, Infinity Pharmaceuticals and Takeda, and received travel expense compensation from Merck. A.L.H. receives research funding from Eisai, Bristol-Myers Squibb, Kura Oncology, AstraZeneca, Genentech Roche, Celldex, Pfizer, Lilly and Bayer; consulting fees from Bristol-Myers Squibb, Merck, Novartis, AstraZeneca, Regeneron, Sanofi Aventis, Sun Pharmaceuticals, Eisai, Genentech/Roche, Genzyme and Ayala Pharmaceuticals; and travel fees from Ignyta and Kura Oncology. C.A.B. receives research funding from Merck, Amgen and Bristol-Myers Squibb. J.D.W. was a consultant for Adaptive Biotech, Amgen, Apricity, Array BioPharma, Ascentage Pharma, Beigene, Bristol-Myers Squibb, Celgene, Chugai, Elucida, Eli Lilly, F Star, Genentech, Imvaq, Kleo Pharma, MedImmune, Merck, Neon Therapuetics, Ono, Polaris Pharma, Polynoma, Psioxus, Puretech, Recepta, Trienza, Sellas Life Sciences, Serametri, Surface Oncology and Syndax; received research support from Bristol-Myers Squibb, Medimmune, Merck Pharmaceutical, and Genentech; and holds equity in Potenza Therapeutics, Tizona Pharmaceuticals, Adaptive Biotechnologies, Elucida, Imvaq, Beigene and Trienza. J.B. is on the Board of Directors for Varian Medical Systems, Bristol-Myers Squibb and Foghorn, and is a past board member of Grail, Aura Biosciences and Infinity Pharmaceuticals. He has performed consulting and/or advisory work for Grail, PMV Pharma, ApoGen, Juno, Roche, Lilly, Novartis and Northern Biologics. He has stock or other ownership interests in PMV Pharma, Grail, Juno, Varian, Foghorn, Aura, Infinity and ApoGen, as well Tango and Venthera, for which he is a cofounder. He has previously received honoraria and/or travel expenses from Roche, Novartis and Lilly. G.Y.K. received research funding and consulting fees from AstraZeneca, Bristol-Myers Squibb and Merck. I.K.M. reports research funding from GE and consulting/speaker fees from Agios Pharmaceuticals, Debiopharm Group, Roche, Merck, Puma Biotechnology and Deciphera Pharmaceuticals. W.D.T. reports personal fees from Eli Lilly, personal fees from EMD Serono, personal fees from Novartis, personal fees from Eisai, personal fees from Janssen, personal fees from Immune Design, personal fees from Adaptimmune, personal fees from Daiichi Sankyo, personal fees from Blueprint, personal fees from Loxo, personal fees from GlaxoSmithKline and personal fees from Agios Pharmaceuticals and from Plexxikon Pharmaceuticals, outside the submitted work. In addition, W.D.T. has a patent (Companion Diagnostic for CDK4 inhibitors–14/854,329) pending to MSKCC/SKI, and a patent (Methods of Treating Metastatic Sarcoma using Talimogene Laherparepvec (T-Vec) and Pembrolizumab Combination Therapy—62/671,625) pending to MSKCC/SKI, and Scientific Advisory Board—Certis Oncology Solutions, stock ownership; Scientific Advisory Board—Atropos Therapeutics, stock ownership. C.M.R. has consulted on oncology drug development with AbbVie, Amgen, Ascentage, Bristol-Myers Squibb, Celgene, Daiichi Sankyo, Genentech/Roche, Harpoon, Loxo, Pharmamar and Seattle Genetics. V.T. is a cofounder and consultant for BluRock Therapeutics. N.A.R. received consulting fees from Merck, AstraZeneca, Roche, Bristol-Myers Squibb, Novartis, Pfizer, Lilly, Abbvie, Merck KGaA, Regeneron and Janssen; is a cofounder and shareholder in Gritstone Oncology; and serves on the advisory board or Neogenomics, OncoMed and Bellcum. M.L. has received ad hoc advisory board compensation from AstraZeneca, Bristol-Myers Squibb, Takeda and Bayer, and research support from LOXO Oncology and Helsinn Healthcare. M.F.B. received research support from Illumina and consulting fees from Roche. D.B.S. received honoraria/consulted for Pfizer, Loxo Oncology, Illumina, Intezyne and Vivideon Therapuetics. N.R. receives research support from Bristol-Myers Squibb and Pfizer and speakers fees from Illumina. T.A.C. is a cofounder of Gritstone Oncology and holds equity. T.A.C. holds equity in An2H. T.A.C. acknowledges grant funding from Bristol-Myers Squibb, AstraZeneca, Illumina, Pfizer, An2H and Eisai. T.A.C. has served as an advisor for Bristol-Myers Squibb, Illumina, Eisai and An2H. MSK has licensed the use of TMB for the identification of patients that benefit from immune checkpoint therapy to PGDx. MSK and T.A.C. receive royalties as part of this licensing agreement. L.G.T.M. received consulting fees from Rakuten Aspyrian and speaker fees from Physician Educational Resources.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 1 and 2
Supplementary Data
Data file containing patient-level data for ICI- and non-ICI-treated patients
Rights and permissions
About this article
Cite this article
Samstein, R.M., Lee, CH., Shoushtari, A.N. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51, 202–206 (2019). https://doi.org/10.1038/s41588-018-0312-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0312-8
This article is cited by
-
Mutation status of the KMT2 family associated with immune checkpoint inhibitors (ICIs) therapy and implicating diverse tumor microenvironments
Molecular Cancer (2024)
-
Tumor mutational burden predictability in head and neck squamous cell carcinoma patients treated with immunotherapy: systematic review and meta-analysis
Journal of Translational Medicine (2024)
-
Discontinuation risk from adverse events: immunotherapy alone vs. combined with chemotherapy: a systematic review and network meta-analysis
BMC Cancer (2024)
-
Biomarker discovery with quantum neural networks: a case-study in CTLA4-activation pathways
BMC Bioinformatics (2024)
-
Refining mutanome-based individualised immunotherapy of melanoma using artificial intelligence
European Journal of Medical Research (2024)