Abstract
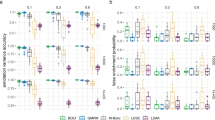
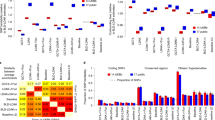
We present SumHer, software for estimating confounding bias, SNP heritability, enrichments of heritability and genetic correlations using summary statistics from genome-wide association studies. The key difference between SumHer and the existing software LD Score Regression (LDSC) is that SumHer allows the user to specify the heritability model. We apply SumHer to results from 24 large-scale association studies (average sample size 121,000) using our recommended heritability model. We show that these studies tended to substantially over-correct for confounding, and as a result the number of genome-wide significant loci was under-reported by about a quarter. We also estimate enrichments for 24 categories of SNPs defined by functional annotations. A previous study using LDSC reported that conserved regions were 13-fold enriched, and found a further six categories with above threefold enrichment. By contrast, our analysis using SumHer finds that none of the categories have enrichment above twofold. SumHer provides an improved understanding of the genetic architecture of complex traits, which enables more efficient analysis of future genetic data.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The simulations and 25 raw GWAS used data from the Wellcome Trust and the eMERGE Network. These were applied for and downloaded from, respectively, the European Genome-phenome Archive (accession codes EGAD00000000001, EGAD00000000002, EGAD00000000003, EGAD00000000004, EGAD00000000005, EGAD00000000006, EGAD00000000007, EGAD00000000008, EGAD00000000009, EGAD00000000021, EGAD00000000022, EGAD00000000023, EGAD00000000024, EGAD00000000025, EGAD00000000057, EGAD00010000124, EGAD00010000264, EGAD00010000506, EGAD00010000634, EGAS00001000672) and from dbGaP (accession code phs000888.v1.p1). To investigate the impact of the reference panel, we used data from the Health and Retirement Study, also available from dbGaP (accession code: phs000428.v2.p2). Results for each of the 24 summary GWAS are available to download from the websites of the corresponding studies (see Table 1 for references).
References
Bulik-Sullivan, B. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Finucane, H. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2016).
Speed, D., Cai, N., Johnson, M. R., Nejentsev, S. & Balding, D. J. Reevaluation of SNP heritability in complex human traits. Nat. Genet. 49, 986–992 (2017).
Speed, D., Hemani, G., Johnson, M. R. & Balding, D. J. Improved heritability estimation from genome-wide SNPs. Am. J. Hum. Genet. 91, 1011–1021 (2012).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Yang, J. et al. Genomic partitioning of genetic variation for complex traits using common SNPs. Nat. Genet. 43, 519–525 (2011).
Lee, S. H., Yang, J., Goddard, M. E., Visscher, P. M. & Wray, N. R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism-derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542 (2012).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
The Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Gottesman, O. et al. The electronic medical records and genomics (eMERGE) network: past, present, and future. Genet. Med. 15, 761 (2013).
Pasaniuc, B. & Price, A. L. Dissecting the genetics of complex traits using summary association statistics. Nat. Rev. Genet. 18, 117–127 (2017).
Speed, D. et al. Describing the genetic architecture of epilepsy through heritability analysis. Brain 137, 2680–2689 (2014).
Verma, S. et al. Imputation and quality control steps for combining multiple genome-wide datasets. Front. Genet. 5, 370 (2015).
Leslie, S. et al. The fine-scale genetic structure of the British population. Nature 519, 309–314 (2015).
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011).
Lippert, C. et al. FaST linear mixed models for genome-wide association studies. Nat. Methods 8, 833–835 (2011).
Yang, J., Zaitlen, N., Goddard, M., Visscher, P. & Price, A. Advantages and pitfalls in the application of mixed-model association methods. Nat. Genet. 46, 100–106 (2014).
Loh, P. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Yu, J. et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38, 203–208 (2006).
Sawcer, S. et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011).
Locke, A. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Global Lipids Genetics Consortium. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013).
The International HapMap 3 Consortium. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Gazal, S. et al. Linkage disequilibrium-dependent architecture of human complex traits shows action of negative selection. Nat. Genet. 49, 1421–1427 (2017).
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011).
Ward, L. & Kellis, M. Evidence of abundant purifying selection in humans for recently acquired regulatory functions. Science 337, 1675–1678 (2012).
Hoffman, M. et al. Integrative annotation of chromatin elements from ENCODE data. Nucleic Acids Res. 41, 827–841 (2013).
Euesden, J., Lewis, C. & O’Reilly, P. PRSice: polygenic risk score software. Bioinformatics 31, 1466–1468 (2015).
Vilhjálmsson, B. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Lambert, J. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Schunkert, H. et al. Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat. Genet. 43, 333–338 (2011).
Liu, J. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
The Tobacco and Genetics Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat. Genet. 42, 441–447 (2010).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Scott, R. et al. An expanded genome-wide association study of type 2 diabetes in Europeans. Diabetes 66, 2888–2902 (2017).
Zheng, H. et al. Whole-genome sequencing identifies EN1 as a determinant of bone density and fracture. Nature 526, 112–117 (2015).
Okbay, A. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 48, 626–633 (2016).
Manning, A. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Soranzo, N. et al. Common variants at 10 genomic loci influence hemoglobin A1(C) levels via glycemic and nonglycemic pathway. Diabetes 59, 3229–3239 (2010).
Wood, A. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Perry, J. et al. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature 514, 92–97 (2014).
Day, F. et al. Large-scale genomic analyses link reproductive aging to hypothalamic signaling, breast cancer susceptibility and BRCA1-mediated DNA repair. Nat. Genet. 47, 1294–1303 (2015).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nat. Genet. 518, 187–196 (2015).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Dempster, E. & Lerner, I. Heritability of threshold characters. Genetics 35, 212–236 (1950).
Lee, S., Wray, N., Goddard, M. & Visscher, P. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Wakefield, J. Bayes factors for genome-wide association studies: comparison with P-values. Genet. Epidemiol. 33, 79–86 (2009).
Pickrell, J. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Delaneau, O., Zagury, J. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 (Bethesda) 1, 457–470 (2011).
Astle, W. & Balding, D. J. Population structure and cryptic relatedness in genetic association studies. Stat. Sci. 24, 451–471 (2009).
Acknowledgements
We thank A. Price, H. Finucane, P. O’Reilly and M. Speed for helpful discussions. Access to the Wellcome Trust Case Control Consortium data was authorized as work related to the project ‘Genome-wide association study of susceptibility and clinical phenotypes in epilepsy’, access to eMERGE Network data was granted under dbGaP Project 14422, ‘Comprehensive testing of SNP-based prediction models’, while access to the Health and Retirement Study was granted under dbGaP Project 15139, ‘Developing summary-statistic tools for analysing genetic association study data’. D.S. is funded by the UK Medical Research Council under grant no. MR/L012561/1, by the European Union’s Horizon 2020 Research and Innovation Programme under the Marie Skłodowska-Curie grant agreement no. 754513, by Aarhus University Research Foundation (AUFF) and the Independent Research Fund Denmark under Project no. 7025-00094B. The eMERGE Network was initiated and funded by NHGRI through the following grants: U01HG006828 (Cincinnati Children’s Hospital Medical Center/Boston Children’s Hospital); U01HG006830 (Children’s Hospital of Philadelphia); U01HG006389 (Essentia Institute of Rural Health, Marshfield Clinic Research Foundation and Pennsylvania State University); U01HG006382 (Geisinger Clinic); U01HG006375 (Group Health Cooperative); U01HG006379 (Mayo Clinic); U01HG006380 (Icahn School of Medicine at Mount Sinai); U01HG006388 (Northwestern University); U01HG006378 (Vanderbilt University Medical Center); and U01HG006385 (Vanderbilt University Medical Center serving as the Coordinating Center). The Health and Retirement Study genetic data is sponsored by the National Institute on Aging (grant nos. U01AG009740, RC2AG036495, and RC4AG039029) and was conducted by the University of Michigan. Analyses were performed with the use of the UCL Computer Science Cluster and the help of the CS Technical Support Group, as well as the use of the UCL Legion High-Performance Computing Facility (Legion@UCL) and associated support services.
Author information
Authors and Affiliations
Contributions
D.S. performed the analysis, D.S. and D.J.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–20, Supplementary Tables 1–20 and Supplementary Note
Rights and permissions
About this article
Cite this article
Speed, D., Balding, D.J. SumHer better estimates the SNP heritability of complex traits from summary statistics. Nat Genet 51, 277–284 (2019). https://doi.org/10.1038/s41588-018-0279-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0279-5
This article is cited by
-
A method to estimate the contribution of rare coding variants to complex trait heritability
Nature Communications (2024)
-
Genetic influences on circulating retinol and its relationship to human health
Nature Communications (2024)
-
Quantile generalized measures of correlation
Statistics and Computing (2024)
-
Accurate and efficient estimation of local heritability using summary statistics and the linkage disequilibrium matrix
Nature Communications (2023)
-
A versatile, fast and unbiased method for estimation of gene-by-environment interaction effects on biobank-scale datasets
Nature Communications (2023)