Abstract
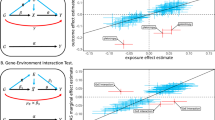
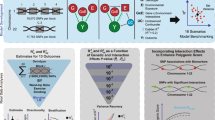
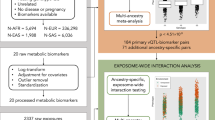
Different exposures, including diet, physical activity, or external conditions can contribute to genotype–environment interactions (G×E). Although high-dimensional environmental data are increasingly available and multiple exposures have been implicated with G×E at the same loci, multi-environment tests for G×E are not established. Here, we propose the structured linear mixed model (StructLMM), a computationally efficient method to identify and characterize loci that interact with one or more environments. After validating our model using simulations, we applied StructLMM to body mass index in the UK Biobank, where our model yields previously known and novel G×E signals. Finally, in an application to a large blood eQTL dataset, we demonstrate that StructLMM can be used to study interactions with hundreds of environmental variables.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The BIOS RNA data can be obtained from the European Genome-phenome Archive (EGA; accession EGAS00001001077). Genotype data are available from the respective biobanks.
References
Hunter, D. J. Gene-environment interactions in human diseases. Nat. Rev. Genet. 6, 287–298 (2005).
Ritz, B. R. et al. Lessons learned from past gene-environment interaction successes. Am. J. Epidemiol. 186, 778–786 (2017).
Brown, A. A. et al. Genetic interactions affecting human gene expression identified by variance association mapping. eLife 3, e01381 (2014).
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 343, 1246949 (2014).
Kraft, P., Yen, Y. C., Stram, D. O., Morrison, J. & Gauderman, W. J. Exploiting gene-environment interaction to detect genetic associations. Hum. Hered. 63, 111–119 (2007).
Rask-Andersen, M., Karlsson, T., Ek, W. E. & Johansson, A. Gene-environment interaction study for BMI reveals interactions between genetic factors and physical activity, alcohol consumption and socioeconomic status. PLoS Genet. 13, e1006977 (2017).
Lin, X., Lee, S., Christiani, D. C. & Lin, X. Test for interactions between a genetic marker set and environment in generalized linear models. Biostatistics 14, 667–681 (2013).
Lin, X. et al. Test for rare variants by environment interactions in sequencing association studies. Biometrics 72, 156–164 (2016).
Casale, F. P., Horta, D., Rakitsch, B. & Stegle, O. Joint genetic analysis using variant sets reveals polygenic gene-context interactions. PLoS Genet. 13, e1006693 (2017).
Kilpelainen, T. O. et al. Physical activity attenuates the influence of FTO variants on obesity risk: a meta-analysis of 218,166 adults and 19,268 children. PLoS Med. 8, e1001116 (2011).
Ahmad, S. et al. Gene x physical activity interactions in obesity: combined analysis of 111,421 individuals of European ancestry. PLoS Genet. 9, e1003607 (2013).
Bjornland, T., Langaas, M., Grill, V. & Mostad, I. L. Assessing gene-environment interaction effects of FTO, MC4R and lifestyle factors on obesity using an extreme phenotype sampling design: Results from the HUNT study. PLoS One 12, e0175071 (2017).
Young, A. I., Wauthier, F. & Donnelly, P. Multiple novel gene-by-environment interactions modify the effect of FTO variants on body mass index. Nat. Commun. 7, 12724 (2016).
Corella, D. et al. Statistical and biological gene-lifestyle interactions of MC4R and FTO with diet and physical activity on obesity: new effects on alcohol consumption. PLoS One 7, e52344 (2012).
Qi, Q. et al. Fried food consumption, genetic risk, and body mass index: gene-diet interaction analysis in three US cohort studies. BMJ 348, g1610 (2014).
Lee, S. et al. Optimal unified approach for rare-variant association testing with application to small-sample case-control whole-exome sequencing studies. Am. J. Hum. Genet. 91, 224–237 (2012).
Crawford, L., Zeng, P., Mukherjee, S. & Zhou, X. Detecting epistasis with the marginal epistasis test in genetic mapping studies of quantitative traits. PLoS Genet. 13, e1006869 (2017).
Gauderman, W. J. et al. Update on the state of the science for analytical methods for gene-environment interactions. Am. J. Epidemiol. 186, 762–770 (2017).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Bycroft, C. et al. Genome-wide genetic data on ~500,000 UK Biobank participants. bioRxiv, https://doi.org/10.1101/166298 (2017).
Richardson, A. S. et al. Moderate to vigorous physical activity interactions with genetic variants and body mass index in a large US ethnically diverse cohort. Pediatr. Obes. 9, e35–e46 (2014).
Ahmad, S. et al. Established BMI-associated genetic variants and their prospective associations with BMI and other cardiometabolic traits: the GLACIER Study. Int. J. Obes. (Lond). 40, 1346–1352 (2016).
Hall, N. G., Klenotic, P., Anand-Apte, B. & Apte, S. S. ADAMTSL-3/punctin-2, a novel glycoprotein in extracellular matrix related to the ADAMTS family of metalloproteases. Matrix Biol. 22, 501–510 (2003).
Zillikens, M. C. et al. Large meta-analysis of genome-wide association studies identifies five loci for lean body mass. Nat. Commun. 8, 80 (2017).
Wen, W. et al. Genome-wide association studies in East Asians identify new loci for waist-hip ratio and waist circumference. Sci. Rep. 6, 17958 (2016).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Zhernakova, D. V. et al. Identification of context-dependent expression quantitative trait loci in whole blood. Nat. Genet. 49, 139–145 (2017).
Westra, H. J. et al. Cell specific eQTL analysis without sorting cells. PLoS Genet. 11, e1005223 (2015).
Cookson, W., Liang, L., Abecasis, G., Moffatt, M. & Lathrop, M. Mapping complex disease traits with global gene expression. Nat. Rev. Genet. 10, 184–194 (2009).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 45, D896–D901 (2017).
Galvez, J. Role of Th17 cells in the pathogenesis of human IBD. ISRN Inflamm. 2014, 928461 (2014).
Day, F. R., Loh, P.-R., Scott, R. A., Ong, K. K. & Perry, J. R. A robust example of collider bias in a genetic association study. Am. J. Hum. Genet. 98, 392–393 (2016).
Listgarten, J., Lippert, C. & Heckerman, D. FaST-LMM-Select for addressing confounding from spatial structure and rare variants. Nat. Genet. 45, 470 (2013).
Wu, M. C. et al. Rare-variant association testing for sequencing data with the sequence kernel association test. Am. J. Hum. Genet. 89, 82–93 (2011).
Lee, S., Wu, M. C. & Lin, X. Optimal tests for rare variant effects in sequencing association studies. Biostatistics 13, 762–775 (2012).
Schaeffer, L. Application of random regression models in animal breeding. Livest. Prod. Sci. 86, 35–45 (2004).
Casale, F. P., Rakitsch, B., Lippert, C. & Stegle, O. Efficient set tests for the genetic analysis of correlated traits. Nat. Methods 12, 755–758 (2015).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Fesinmeyer, M. D. et al. Genetic risk factors for BMI and obesity in an ethnically diverse population: results from the population architecture using genomics and epidemiology (PAGE) study. Obesity 21, 835–846 (2013).
Abraham, G., Qiu, Y. & Inouye, M. FlashPCA2: principal component analysis of Biobank-scale genotype datasets. Bioinformatics 33, 2776–2778 (2017).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Methodol. 57, 289–300 (1995).
Van Greevenbroek, M. M. et al. The cross‐sectional association between insulin resistance and circulating complement C3 is partly explained by plasma alanine aminotransferase, independent of central obesity and general inflammation (the CODAM study). Eur. J. Clin. Invest. 41, 372–379 (2011).
Tigchelaar, E. F. et al. Cohort profile: LifeLines DEEP, a prospective, general population cohort study in the northern Netherlands: study design and baseline characteristics. BMJ Open 5, e006772 (2015).
Hofman, A. et al. The Rotterdam Study: 2014 objectives and design update. Eur. J. Epidemiol. 28, 889–926 (2013).
Skyler, J. S. Pulmonary insulin update. Diabetes Technol. Ther. 7, 834–839 (2005).
Storey, J. D. A direct approach to false discovery rates. J. R. Stat. Soc. Series B Methodol. 64, 479–498 (2002).
Alexa, A. & Rahnenfuhrer, J. topGO: enrichment analysis for gene ontology. R package version 2 (2010).
Lippert, C., Casale, F. P., Rakitsch, B. & Stegle, O. LIMIX: genetic analysis of multiple traits. bioRxiv https://doi.org/10.1101/003905 (2014).
Acknowledgements
The authors thank C. Lippert and L. Parts for helpful discussions. This research was conducted using the UK Biobank Resource (Application Number 14069). R.M. was supported by a PhD fellowship from the Mathematical Genomics and Medicine program, funded by the Wellcome Trust. F.P.C., D.H. and O.S. received support from core funding of the European Molecular Biology Laboratory and the European Union’s Horizon2020 research and innovation program under grant agreement N635290. I.B. acknowledges funding from Wellcome (WT098051 and WT206194). M.J.B. was supported by a fellowship from the EMBL Interdisciplinary Postdoc (EI3POD) program under Marie Skłodowska-Curie Actions COFUND (grant number 664726). The Biobank-Based Integrative Omics Studies (BIOS) Consortium is funded by BBMRI-NL, a research infrastructure financed by the Dutch government (NWO 184.021.007).
Author information
Authors and Affiliations
Consortia
Contributions
R.M., F.P.C., I.B., and O.S. conceived the method. R.M., F.P.C., and D.H. implemented the methods. R.M., F.P.C., and M.J.B. analyzed the data. L.F. and BIOS Consortium provided data resources. R.M., F.P.C., I.B., and O.S. interpreted results and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
F.P.C. was employed at Microsoft while performing the research.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–23, Supplementary Tables 1 and 2, and Supplementary Note
Supplementary Table 3
Interactions identified by StructLMM for BMI in UK Biobank
Supplementary Table 4
Associations identified by StructLMM and LMM in the association analysis of BMI using data from UK Biobank
Supplementary Table 5
Summary table of interaction eQTL analysis in blood cohort
Supplementary Table 6
Pathway enrichment analysis for interaction eQTLs that are in linkage with GWAS loci
Supplementary Data 1
eQTL Manhattan plots for interaction eQTLs that colocalize with disease variants
Supplementary Data 2
Interaction eQTL colocalizing with disease variants
Rights and permissions
About this article
Cite this article
Moore, R., Casale, F.P., Jan Bonder, M. et al. A linear mixed-model approach to study multivariate gene–environment interactions. Nat Genet 51, 180–186 (2019). https://doi.org/10.1038/s41588-018-0271-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0271-0
This article is cited by
-
Effects of sex and gender on the etiologies and presentation of select internalizing psychopathologies
Translational Psychiatry (2024)
-
A fast non-parametric test of association for multiple traits
Genome Biology (2023)
-
Sex differences in the polygenic architecture of hearing problems in adults
Genome Medicine (2023)
-
A versatile, fast and unbiased method for estimation of gene-by-environment interaction effects on biobank-scale datasets
Nature Communications (2023)
-
Environmental neuroscience linking exposome to brain structure and function underlying cognition and behavior
Molecular Psychiatry (2023)