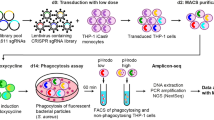
Abstract
Phagocytosis is required for a broad range of physiological functions, from pathogen defense to tissue homeostasis, but the mechanisms required for phagocytosis of diverse substrates remain incompletely understood. Here, we developed a rapid magnet-based phenotypic screening strategy, and performed eight genome-wide CRISPR screens in human cells to identify genes regulating phagocytosis of distinct substrates. After validating select hits in focused miniscreens, orthogonal assays and primary human macrophages, we show that (1) the previously uncharacterized gene NHLRC2 is a central player in phagocytosis, regulating RhoA-Rac1 signaling cascades that control actin polymerization and filopodia formation, (2) very-long-chain fatty acids are essential for efficient phagocytosis of certain substrates and (3) the previously uncharacterized Alzheimer’s disease–associated gene TM2D3 can preferentially influence uptake of amyloid-β aggregates. These findings illuminate new regulators and core principles of phagocytosis, and more generally establish an efficient method for unbiased identification of cellular uptake mechanisms across diverse physiological and pathological contexts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The RNA-seq FASTQ files are available at GEO under accession GSE107566.
References
Lim, J. J., Grinstein, S. & Roth, Z. Diversity and versatility of phagocytosis: roles in innate immunity, tissue remodeling, and homeostasis. Front. Cell. Infect. Microbiol. 7, 191 (2017).
Gordon, S. Phagocytosis: an immunobiologic process. Immunity 44, 463–475 (2016).
Arandjelovic, S. & Ravichandran, K. S. Phagocytosis of apoptotic cells in homeostasis. Nat. Immunol. 16, 907–917 (2015).
Hong, S., Dissing-Olesen, L. & Stevens, B. New insights on the role of microglia in synaptic pruning in health and disease. Curr. Opin. Neurobiol. 36, 128–134 (2016).
Vargas, M. E. & Barres, B. A. Why is Wallerian degeneration in the CNS so slow? Annu. Rev. Neurosci. 30, 153–179 (2007).
Chao, M. P. et al. Anti-CD47 antibody synergizes with rituximab to promote phagocytosis and eradicate non-Hodgkin lymphoma. Cell 142, 699–713 (2010).
Chen, J. et al. SLAMF7 is critical for phagocytosis of haematopoietic tumour cells via Mac-1 integrin. Nature 544, 493–497 (2017).
Ziegenfuss, J. S., Doherty, J. & Freeman, M. R. Distinct molecular pathways mediate glial activation and engulfment of axonal debris after axotomy. Nat. Neurosci. 15, 979–987 (2012).
Juncadella, I. J. et al. Apoptotic cell clearance by bronchial epithelial cells critically influences airway inflammation. Nature 493, 547–551 (2013).
Chung, W. S. et al. Astrocytes mediate synapse elimination through MEGF10 and MERTK pathways. Nature 504, 394–400 (2013).
Freeman, S. A. & Grinstein, S. Phagocytosis: receptors, signal integration, and the cytoskeleton. Immunol. Rev. 262, 193–215 (2014).
Diakonova, M., Bokoch, G. & Swanson, J. A. Dynamics of cytoskeletal proteins during Fcgamma receptor-mediated phagocytosis in macrophages. Mol. Biol. Cell 13, 402–411 (2002).
Fairn, G. D. & Grinstein, S. How nascent phagosomes mature to become phagolysosomes. Trends Immunol. 33, 397–405 (2012).
Swanson, J. A. Shaping cups into phagosomes and macropinosomes. Nat. Rev. Mol. Cell Biol. 9, 639–649 (2008).
Hedgecock, E. M., Sulston, J. E. & Thomson, J. N. Mutations affecting programmed cell deaths in the nematode Caenorhabditis elegans. Science 220, 1277–1279 (1983).
Ellis, R. E., Jacobson, D. M. & Horvitz, H. R. Genes required for the engulfment of cell corpses during programmed cell death in Caenorhabditis elegans. Genetics 129, 79–94 (1991).
Gumienny, T. L. et al. CED-12/ELMO, a novel member of the CrkII/Dock180/Rac pathway, is required for phagocytosis and cell migration. Cell 107, 27–41 (2001).
Tosello-Trampont, A. C., Brugnera, E. & Ravichandran, K. S. Evidence for a conserved role for CRKII and Rac in engulfment of apoptotic cells. J. Biol. Chem. 276, 13797–13802 (2001).
Logan, M. A. et al. Negative regulation of glial engulfment activity by Draper terminates glial responses to axon injury. Nat. Neurosci. 15, 722–730 (2012).
Silva, E., Au-Yeung, H. W., Van Goethem, E., Burden, J. & Franc, N. C. Requirement for a Drosophila E3-ubiquitin ligase in phagocytosis of apoptotic cells. Immunity 27, 585–596 (2007).
Garver, L. S., Wu, J. & Wu, L. P. The peptidoglycan recognition protein PGRP-SC1a is essential for Toll signaling and phagocytosis of Staphylococcus aureus in Drosophila. Proc. Natl Acad. Sci. USA 103, 660–665 (2006).
Shen, K., Sidik, H. & Talbot, W. S. The Rag-Ragulator complex regulates lysosome function and phagocytic flux in microglia. Cell Rep. 14, 547–559 (2016).
Kocks, C. et al. Eater, a transmembrane protein mediating phagocytosis of bacterial pathogens in Drosophila. Cell 123, 335–346 (2005).
Ramet, M., Manfruelli, P., Pearson, A., Mathey-Prevot, B. & Ezekowitz, R. A. Functional genomic analysis of phagocytosis and identification of a Drosophila receptor for E. coli. Nature 416, 644–648 (2002).
Philips, J. A., Rubin, E. J. & Perrimon, N. Drosophila RNAi screen reveals CD36 family member required for mycobacterial infection. Science 309, 1251–1253 (2005).
Stroschein-Stevenson, S. L., Foley, E., O’Farrell, P. H. & Johnson, A. D. Identification of Drosophila gene products required for phagocytosis of Candida albicans. PLoS Biol. 4, e4 (2006).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882 e1821 (2016).
Koike-Yusa, H., Li, Y., Tan, E. P., Velasco-Herrera Mdel, C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
Morgens, D. W. et al. Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat. Commun. 8, 15178 (2017).
Parnas, O. et al. A genome-wide CRISPR screen in primary immune cells to dissect regulatory networks. Cell 162, 675–686 (2015).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Zhou, Y. et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Wang, L. et al. High-throughput functional genetic and compound Screens identify targets for senescence induction in cancer. Cell Rep 21, 773–783 (2017).
Larrick, J. W., Fischer, D. G., Anderson, S. J. & Koren, H. S. Characterization of a human macrophage-like cell line stimulated in vitro: a model of macrophage functions. J. Immunol. 125, 6–12 (1980).
Morgens, D. W., Deans, R. M., Li, A. & Bassik, M. C. Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes. Nat. Biotechnol. 34, 634–636 (2016).
Rotty, J. D. et al. Arp2/3 complex is required for macrophage integrin functions but is dispensable for FcR phagocytosis and in vivo motility. Dev. Cell. 42, 498–513 e496 (2017).
Bassik, M. C. et al. A systematic mammalian genetic interaction map reveals pathways underlying ricin susceptibility. Cell 152, 909–922 (2013).
Brumell, J. H. et al. Expression of the protein kinase C substrate pleckstrin in macrophages: association with phagosomal membranes. J. Immunol. 163, 3388–3395 (1999).
Ma, A. D. & Abrams, C. S. Pleckstrin induces cytoskeletal reorganization via a Rac-dependent pathway. J. Biol. Chem. 274, 28730–28735 (1999).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Bradley, W. D., Hernandez, S. E., Settleman, J. & Koleske, A. J. Integrin signaling through Arg activates p190RhoGAP by promoting its binding to p120RasGAP and recruitment to the membrane. Mol. Biol. Cell 17, 4827–4836 (2006).
Jongstra-Bilen, J. & Jongstra, J. Leukocyte-specific protein 1 (LSP1): a regulator of leukocyte emigration in inflammation. Immunol. Res. 35, 65–74 (2006).
Schlam, D. et al. Phosphoinositide 3-kinase enables phagocytosis of large particles by terminating actin assembly through Rac/Cdc42 GTPase-activating proteins. Nat. Commun. 6, 8623 (2015).
Kihara, A. Very long-chain fatty acids: elongation, physiology and related disorders. J. Biochem. 152, 387–395 (2012).
Collins, S. R. et al. Using light to shape chemical gradients for parallel and automated analysis of chemotaxis. Mol. Syst. Biol. 11, 804 (2015).
Kajkowski, E. M. et al. beta-Amyloid peptide-induced apoptosis regulated by a novel protein containing a g protein activation module. J. Biol. Chem. 276, 18748–18756 (2001).
Jakobsdottir, J. et al. Rare functional variant in TM2D3 is associated with late-onset Alzheimer’s disease. PLoS Genet. 12, e1006327 (2016).
Stine, W. B., Jungbauer, L., Yu, C. & LaDu, M. J. Preparing synthetic Abeta in different aggregation states. Methods Mol. Biol. 670, 13–32 (2011).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Uusimaa, J. et al. NHLRC2 variants identified in patients with fibrosis, neurodegeneration, and cerebral angiomatosis (FINCA): characterisation of a novel cerebropulmonary disease. Acta Neuropathol. 135, 727–742 (2018).
Nishi, K. et al. ROS-induced cleavage of NHLRC2 by caspase-8 leads to apoptotic cell death in the HCT116 human colon cancer cell line. Cell Death Dis. 8, 3218 (2017).
Horsthemke, M. et al. Multiple roles of filopodial dynamics in particle capture and phagocytosis and phenotypes of Cdc42 and Myo10 deletion. J. Biol. Chem. 292, 7258–7273 (2017).
Larocca, J. N. & Norton, W. T. Isolation of myelin. Curr. Protoc. Cell. Biol. Chapter 3, Unit 325 (2007).
Mosser, D. M. & Zhang, X. Measuring opsonic phagocytosis via Fcgamma receptors and complement receptors on macrophages. Curr. Protoc. Immunol. Chapter 14, Unit 14 27 (2011).
Dunkley, P. R., Jarvie, P. E. & Robinson, P. J. A rapid Percoll gradient procedure for preparation of synaptosomes. Nat. Protoc. 3, 1718–1728 (2008).
Kampmann, M., Bassik, M. C. & Weissman, J. S. Integrated platform for genome-wide screening and construction of high-density genetic interaction maps in mammalian cells. Proc. Natl Acad. Sci. USA 110, E2317–E2326 (2013).
Kramer, N. J. et al. CRISPR-Cas9 screens in human cells and primary neurons identify modifiers of C9ORF72 dipeptide-repeat-protein toxicity. Nat. Genet. 50, 603–612 (2018).
Liu, N. et al. Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators. Nature 553, 228–232 (2018).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome. Biol. 15, 554 (2014).
Acknowledgements
We thank the members of the Bassik and Barres laboratories for feedback and support throughout, and S. Grinstein, J. Pritchard, J. Pluvinage, and T. Wyss-Coray for helpful suggestions. We thank T. Flores, J. Mulholland and J. Perrino of the Stanford Electron Microscopy facility for excellent assistance with SEM images, and D. Lysko and W. Talbot for experimental advice. This work was supported by the Christopher and Dana Reeve Foundation International Research Consortium on Spinal Cord Injury, the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation, the JPB Foundation, the Novartis Institute of Basic Research, a National Institutes of Health (NIH) grant to B.A.B. (R01 DA015043), a Jane Coffin Childs Fellowship to R.A.K., the Howard Hughes Medical Institute (J.A.T. and D.T.) and the Stanford Medical Scientist Training Program NIH T32-GM007365 (for L.L.), a Kimmel Scholar Award, an NIH Director’s New Innovator Award (DP2 HD094656) to S.R.C., an NIH Director’s New Innovator Award (1DP2HD084069-01) and Stanford Neuroscience Institute Brain Rejuvenation Project Award to M.C.B., and generous contributions from Vincent and Stella Coates. C.J.B. was supported by the Damon Runyon Cancer Research Foundation (DRG-2125-12).
Author information
Authors and Affiliations
Contributions
C.J.B., M.S.H., B.A.B., S.R.C. and M.C.B. conceptualized the study. C.J.B., M.S.H. and D.W.M. generated and analyzed screen data. A.L. assisted in the cloning of sgRNA libraries. K.T. performed FACS-based phagocytosis screening. J.A.O. and B.E. assisted in generating sgRNA-expressing U937 and RAW 264.7 cell lines and clonally derived knockout lines. H.C. and A.T. assisted in performing phagocytosis microscopy assays. J.A.O. made BirA-NHLRC2 construct and prepared samples for mass spectrometry with advice from E.C.; J.C. and L.J. ran mass spectrometry samples and assisted with analysis; E.R. performed neutrophil migration assays with S.R.C.; J.T., D.V., L.L. and R.L. advised on microscopy experiments. R.L. advised on and performed phalloidin microscopy for NHLRC2 knockouts and frustrated phagocytosis assay.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Differentiated U937 cells are highly phagocytic.
a,b, Staining of undifferentiated (a) or PMA-differentiated (b) U937 cells with a CD11b antibody (green) and DAPI (blue). Representative of two independent experiments. c, Flow cytometry of CD11b levels in undifferentiated or PMA-differentiated U937 cells. Representative of two independent experiments. d, Phagocytic index of differentiated (red), undifferentiated (green), or differentiated U937 cells treated with the actin polymerization inhibitor cytochalasin D (blue). Phagocytic index is measured by live-cell microscopy using 1.3-μm beads labeled with the pH-sensitive dye pHrodo, which becomes fluorescent upon reaching lysosomal pH. Phagocytic index was calculated at each time point by measuring the total area of pHrodo signal normalized by the total area of live cells (indicated by calcein AM), and the index is presented relative to the average value of the control condition at the time point at 5 h. Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. e, Phagocytic index for pHrodo-labeled zymosan in differentiated U937 cells (red) and differentiated U937 cells treated with the actin polymerization inhibitor cytochalasin D (blue). Values represent the means ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. f, Proportion of WT differentiated U937 cells in the bound fraction of a magnetic column after the addition 1.3-μm magnetic beads (red) and proportion of WT differentiated U937 cells treated with the actin polymerization inhibitor cytochalasin D in the bound fraction of a magnetic column after the addition of 1.3-μm magnetic beads. g, Schematic of manually categorized genes passing 10% FDR (by casTLE) according to cellular process or compartment.
Supplementary Figure 2 Generalization of the magnetic based phagocytosis screening method to diverse substrates.
a, Myelin phagocytosis by differentiated U937 cells (red) is inhibited by addition of the actin polymerization inhibitor cytochalasin D (blue). Phagocytic index was calculated at each time point by measuring the total area of pHrodo signal normalized by the total area of live cells (indicated by calcein AM), and the index is presented relative to the average value of the control condition at the time point at 5 h. Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. b, Titration of IONP concentration applied to myelin shows an increase in the proportion of differentiated U937 cells bound to the magnetic column after phagocytosis (red), while titration of IONPs alone shows minimal increase in differentiated U937 cells bound to a magnetic column (blue). Magnetic labeling density chosen for genome-wide screens is indicated with the gray dotted line. c, Schematic of the pilot experiment to demonstrate the method of separating differentiated U937 cells based on the amount of IONP-labeled material phagocytosed. Myelin labeled with pHrodo-red and IONPs was applied to unlabeled U937 cells, allowing for 6 h of phagocytosis. Alternatively, myelin labeled with pHrodo-red and IONPs was applied to a second population of U937 cells labeled with calcein AM (green), allowing for 2 h of phagocytosis. These two populations were then mixed 1:1 and separated by a magnetic column. d, Microscopy shows an even distribution of calcein AM–labeled U937 cells (2 h of phagocytosis) and U937 cells with pHrodo-red myelin (6 h of phagocytosis) before magnetic separation, but an enrichment of pHrodo-red-labeled myelin in the bound column (without calcein AM) and a corresponding enrichment of green-labeled cells in the unbound column. Representative of three independent experiments. e, Proportion of differentiated U937 cells bound to the magnetic column after IONP-labeled myelin was applied for 2 or 6 h. Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. f, Proportion of a 1:1 mixture of green and unlabeled differentiated U937 cells in each fraction before and after separation on a magnetic column. g, Microscopy demonstrating enrichment of pHrodo- (red) and IONP-labeled red blood cells opsonized with IgG in the bound fraction of a magnetic column compared to the unbound fraction. Live cells were labeled with calcein AM dye (green). Representative of two independent experiments. h, Microscopy demonstrating enrichment of pHrodo- (red) and IONP-labeled zymosan in the bound fraction of a magnetic column compared to the unbound fraction. Live cells were labeled with calcein AM dye (green). Representative of two independent experiments. i, Demonstration that red blood cells labeled with IONPs but not opsonized with IgG (gray) are not phagocytosed at the same rate as red blood cells labeled with IONPs and opsonized with IgG (red). This phagocytosis was eliminated with the actin polymerization inhibitor cytochalasin D (blue). Values represent the means ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. j, Demonstration that red blood cells labeled with IONPs but not opsonized with C3b (gray) are not phagocytosed at the same rate as red blood cells labeled with IONPs and opsonized with C3b (red). Phagocytosis was eliminated with the actin polymerization inhibitor cytochalasin D (blue). Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments.
Supplementary Figure 3 mRNA expression in myeloid cells of genes discovered to be phagocytosis screen hits and correlation of replicate screens.
a, mRNA expression levels of genes passing 10% FDR for each substrate in differentiated U937 cells and primary human microglia. Plots illustrate mean (bar), minimum, and maximum values, with each gene’s mean FPKM value derived from n = 4 sequenced replicates. b, Correlation of replicate genome-wide screens. Signed confidence score (casTLE score) of each genome-wide screen plotted against a biological replicate screen. The signed confidence score is negative if gene knockout inhibits phagocytosis and positive if it promotes phagocytosis. Each casTLE score is derived from n = 2 replicate screens. c, Correlation of replicate-focused batch retest screens. The signed confidence score (casTLE score) of each focused batch retest screen is plotted against a biological replicate screen. The signed confidence score is negative if gene knockout inhibits phagocytosis and positive if it promotes phagocytosis. Each casTLE score is derived from n = 2 replicate screens.
Supplementary Figure 4 Validation of genome-wide screen results using focused sublibraries and individual gene KO cell lines.
a, Comparison of genome-wide and focused sublibrary screens. Scatterplots of confidence scores for genes in the focused sublibrary batch retest screens and original genome-wide results across various substrates. Confidence scores are calculated by casTLE and are negative if the gene KO inhibits phagocytosis and positive if the gene KO promotes phagocytosis. Hits discovered in the genome-wide screen for a given substrate are highlighted in red. Genes that were present in the batch library but were not hits for the given substrate in the genome-wide screen are in gray. Note that the focused sublibrary screen reveals phenotypes for many genes that were missed in the genome-wide screens (false negatives). b, Representation of the number of hits for each focused sublibrary screen. Hits are defined as the gene KO having a nonzero effect on phagocytosis within a 95% credible interval. c, Summary of all microscopy-based zymosan validations. U937 cells containing the indicated guide were treated with pHrodo-labeled zymosan and phagocytosis was monitored over time using automated microscopy. The phagocytic index of the time point at 5 h is presented. Data are the average of four independent replicate wells, and error bars represent the s.e.m. d, Summary of all microscopy-based positive bead validations. U937 cells containing the indicated guide were treated with pHrodo-labeled positively charged beads and phagocytosis was monitored over time using automated microscopy. The phagocytic index of the time point at 5 h is presented. Data are the average of four independent replicate wells, and error bars represent s.e.m. e, Validation by magnetic separation of magnetized zymosan and positively charged beads for predicted phagocytosis-promoting hits ICAM1 and CCNC. U937 cells containing the indicated guide were treated with magnetized zymosan or magnetic positively charge beads. Cells were separated by magnetic column as conducted in genome-wide screens. Cells in the unbound and bound fractions were counted and the fold enrichment of the bound:unbound cell ratio was calculated. f, Validation by magnetic separation of magnetized zymosan and positively charged beads for the predicted substrate-specific hits PLEK and TLN1. U937 cells containing the indicated guide were treated with magnetized zymosan or magnetic positively charge beads. Cells were separated by magnetic column as conducted in genome-wide screens. Cells in the unbound and bound fractions were counted and the fold enrichment of the bound:unbound cell ratio was calculated. g, Validation of sgRNAs editing target genes in U937 cells. Genomic DNA from U937 cells containing the indicated guide was submitted to Sanger sequencing at the location of sgRNA homology. The percentage of edited cells at the sgRNA target location was calculated using the ICE software package. Sanger sequencing data are available upon request. h, Validation of sgRNAs editing target genes in clonal U937 and RAW 264.7 cells. Genomic DNA from U937 and RAW 264.7 pooled cell populations containing the indicated guide was single-cell sorted, clonally expanded and submitted to Sanger sequencing at the location of sgRNA homology. The percentage of edited cells at the sgRNA target location was calculated using the ICE software package. Sanger sequencing data are available upon request. i, Validation of sgRNAs editing target genes in primary human macrophage cells. Genomic DNA from primary cells containing the indicated guide was submitted to Sanger sequencing at the location of sgRNA homology. The percentage of edited cells at the sgRNA target location was calculated using the ICE software package. Sanger sequencing data are available upon request.
Supplementary Figure 5 NHLRC2 knockout inhibits phagocytosis.
a, Multiple independent sgRNAs reduce phagocytosis in pools of RAW 264.7 cells expressing sgRNAs targeting Nhlrc2 (blue lines) or Nckap1l (red, orange lines) when compared to a control sgRNA (gray). Values represent the means ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. b, Validation of phagocytic impairment in the RAW 264.7 Nhlrc2 homozygous clonal knockout line (blue) by measuring the phagocytic index of pHrodo-labeled zymosan, compared to RAW 264.7 cells expressing a control sgRNA (gray) or the same line with addition of the actin polymerization inhibitor cytochalasin D (red). Values represent the means ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. c, Amount of active (GTP-bound) RAC1 in RAW 264.7 cells expressing a control sgRNA (red) or confirmed Nhlrc2 clonal knockout RAW 264.7 cells (blue) as determined by ELISA with or without addition of a phagocytic substrate (beads). ELISA signal was normalized against recombinant GTP-bound RAC1. Values represent mean ± s.e.m. of n = 3 replicate wells. Representative of two independent experiments. d, Additional images of F-actin staining of RAW 264.7 cells expressing a control sgRNA (left) and a confirmed Nhlrc2 KO cell line (right) imaged on a spinning-disk confocal microscope. Images were generated from extended-focus projections of confocal sections acquired at 0.4-μm intervals of cells stained with AlexaFluor-488 phalloidin. Representative of four independent experiments. Scale bar, 10 μm. e, Additional SEM images of RAW 264.7 cells expressing a control sgRNA (top), a confirmed Nckap1l KO cell line (middle) and a confirmed Nhlrc2 KO cell line (bottom). Cells were either exposed to IgG-opsonized 7-μm beads for 10 min prior to fixation or fixed without exposure to beads. Representative of two independent experiments.
Supplementary Figure 6 Phagocytic and motility defects in ELOVL1 knockout and knockdown cells.
a, Additional confocal microscopy of RAW 264.7 cells expressing a control sgRNA (left) or ELOVL1 sgRNA (right). Cells were incubated with IgG-coated 7-µm beads for 10 min and fixed. Donkey anti-rabbit IgG signal (red) marks exposed bead surfaces that are not obstructed by contact with a cell. Cells were stained with phalloidin (green) and Hoechst nuclear stain (blue). A white asterisk marks a fully internalized bead. Representative of four independent experiments. b, Additional SEM of RAW 264.7 cells expressing a control sgRNA (right) or ELOVL1 sgRNA (left) that were incubated with IgG-coated 7-μm beads for 10 min and fixed. A white asterisk marks a fully internalized bead. Representative of two independent experiments. c, Differentiated PLB-985 cells were imaged in an under-agarose chemotaxis assay both before and after generation of a chemoattractant gradient by photo-uncaging of a caged derivative of the chemoattractant fMLF. Each experimental well contained both mCherry-labeled experimental cells expressing dCas9 and an sgRNA targeting the indicated gene, and mTurquoise-labeled control cells expressing dCas9 without an sgRNA. RAC2 and ACTR2 were included as positive controls for migration defects, and FPR1 (formyl peptide receptor) was included as a positive control for response to attractant stimulation. Mean cell speed was measured by tracking cell movement without chemoattractant gradient generation (basal migration speed). Shown are the mean cell speeds for the experimental cells, normalized according to the speed of the control cells in the same well and then normalized by the mean of the no-sgRNA wells on each day. A total of at least n = 39 replicate wells (technical replicates) were measured for each sgRNA over a total of five independent days. Error bars indicate the s.e.m. for normalized well-average speed measurements. d, Mean cell speed was measured by tracking cell movement without chemoattractant gradient generation (post-stimulation). Shown are the mean cell speeds for the experimental cells, normalized according to the speed of the control cells in the same well and then normalized by the mean of the no-sgRNA wells on each day. A total of at least n = 39 replicate wells (technical replicates) were measured for each sgRNA over a total of five independent days. Error bars indicate the s.e.m. for normalized well-average speed measurements. e, qPCR was used to measure the transcript abundance of ELOVL1 for cells expressing dCas9 and the indicated sgRNA.
Supplementary Figure 7 Substrate-specific modifiers of phagocytosis.
Assay of U937 cell lines expressing sgRNAs that are simultaneously challenged with two phagocytic substrates within the same well, one substrate labeled with pHrodo green and the other with pHrodo red. Phagocytosis of both substrates is measured over time using live-cell microscopy. a, Ratio of the total red area (indicating zymosan phagocytosis) to the total green area (indicating 1.3-μm bead phagocytosis) at the time point at 5 h of the phagocytosis assay in cells expressing control sgRNAs (gray) or sgRNAs targeting ITGB2, TLN1, FERMT3 and PLEK (blue). Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments. b, Ratio of the total red area (indicating C3b-opsonized RBC phagocytosis) to the total green area (1.3-μm bead phagocytosis) at the time point at 5 h of the phagocytosis assay in cells expressing control sgRNAs (gray) or sgRNAs targeting ITGB2, TLN1, FERMT3, and PLEK (blue). Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of two independent experiments. c, Phagocytosis of pHrodo-labeled amyloid-β (Aβ) aggregates over time in differentiated U937 cells expressing control sgRNAs (black and gray), a clonal NCKAP1L KO U937 line (light blue), a clonal NHLRC2 KO U937s line (dark blue), and a U937 line expressing a control sgRNA with the actin polymerization inhibitor cytochalasin D (red). Values represent mean ± s.e.m. of n = 4 replicate wells. Representative of three independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note
Supplementary Table 1
Results from genome-wide KO phagocytosis screens for eight phagocytic substrates
Supplementary Table 2
Sequencing counts from genome-wide screens
Supplementary Table 3
Labeling densities of IONPs and pHrodo for each substrate tested
Supplementary Table 4
RNA-seq results of differentiated and undifferentiated U937 cells
Supplementary Table 5
Results from targeted (batch library) KO phagocytosis FACS and magnetic-based validation screens
Supplementary Table 6
Sequencing counts from batch library screens
Supplementary Table 7
Single-gene validation data of phagocytosis phenotypes using time-lapse fluorescence microscopy
Supplementary Table 8
Results from BioID followed by mass spectrometry
Supplementary Video 1
Frustrated phagocytosis assay of control-sgRNA-expressing RAW 264.7 cells. Representative of four independent experiments
Supplementary Video 2
Frustrated phagocytosis assay of control of NHLRC2 KO RAW 264.7 cells. Representative of four independent experiments
Supplementary Video 3
z stack of control-sgRNA-expressing RAW 264.7 cells. Representative of two independent experiments
Supplementary Video 4
z stack of ELOVL1 KO RAW 264.7 cells. Representative of two independent experiments
Rights and permissions
About this article
Cite this article
Haney, M.S., Bohlen, C.J., Morgens, D.W. et al. Identification of phagocytosis regulators using magnetic genome-wide CRISPR screens. Nat Genet 50, 1716–1727 (2018). https://doi.org/10.1038/s41588-018-0254-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0254-1
This article is cited by
-
Genome-wide screens identify SEL1L as an intracellular rheostat controlling collagen turnover
Nature Communications (2024)
-
APOE4/4 is linked to damaging lipid droplets in Alzheimer’s disease microglia
Nature (2024)
-
TM2D3, a mammalian homologue of Drosophila neurogenic gene product Almondex, regulates surface presentation of Notch receptors
Scientific Reports (2023)
-
Roles and regulation of microglia activity in multiple sclerosis: insights from animal models
Nature Reviews Neuroscience (2023)
-
Role of NCKAP1 in the Defective Phagocytic Function of Microglia-Like Cells Derived from Rapidly Progressing Sporadic ALS
Molecular Neurobiology (2023)