Abstract
Modern sugarcanes are polyploid interspecific hybrids, combining high sugar content from Saccharum officinarum with hardiness, disease resistance and ratooning of Saccharum spontaneum. Sequencing of a haploid S. spontaneum, AP85-441, facilitated the assembly of 32 pseudo-chromosomes comprising 8 homologous groups of 4 members each, bearing 35,525 genes with alleles defined. The reduction of basic chromosome number from 10 to 8 in S. spontaneum was caused by fissions of 2 ancestral chromosomes followed by translocations to 4 chromosomes. Surprisingly, 80% of nucleotide binding site-encoding genes associated with disease resistance are located in 4 rearranged chromosomes and 51% of those in rearranged regions. Resequencing of 64 S. spontaneum genomes identified balancing selection in rearranged regions, maintaining their diversity. Introgressed S. spontaneum chromosomes in modern sugarcanes are randomly distributed in AP85-441 genome, indicating random recombination among homologs in different S. spontaneum accessions. The allele-defined Saccharum genome offers new knowledge and resources to accelerate sugarcane improvement.
Similar content being viewed by others
Main
Cultivated sugarcanes (Saccharum spp., Poaceae) are unusual among leading crops in that they are polyploid interspecific hybrids, with singularly complex genomes. Domesticated in New Guinea ~10,000 years ago, ‘reeds that produce honey without bees’ were considered a luxury and an expensive spice from the sixth to fourth centuries bc. After the introduction of sugarcane to the Old World around the eighth century1, its spread to Caribbean, South American, Indian Ocean and Pacific island nations drove large human migrations, including slave labor2. Now the world’s number one crop by harvested tonnage and its fifth most valuable crop (FAO, 2012), sugarcane is cultivated on ~26 million hectares of land in >90 countries, and 1.83 billion metric tonnes are harvested annually with a gross production value approaching $57 billion, providing 80% of the world’s sugar and 40% of its ethanol as the primary sugar and biofuel feedstock crop.
While the high sugar content of modern sugarcane cultivars derives from cultivated ‘noble’ forms of Saccharum officinarum, their hardiness, disease resistance and ratooning capacity were obtained during ‘nobilization’, specifically backcrossing into S. officinarum selected traits from a sugar-poor relative, Saccharum spontaneum3. ‘Noble’ S. officinarum cultivars, typically 2n = 8x = 80 (n is the haploid chromosome number in eukaryotic organisms and x is the monoploid chromosome number in polyploid organisms), accumulate sucrose in the stem reaching up to 50% of the dry weight but are vulnerable to biotic and abiotic stresses. Dutch breeders in Java made interspecific crosses between S. officinarum and a wild relative, S. spontaneum, to obtain disease resistance and stress tolerance traits of S. spontaneum while backcrossing to S. officinarum to recover high biomass and high sugar content4. Consequently, modern sugarcane cultivars are interspecific hybrids with approximately 80% chromosomes from S. officinarum, 10–15% chromosomes from S. spontaneum, and 5–10% recombinant chromosomes5.
The lowest chromosome number recorded for natural Saccharum accession is a 2n = 5x = 40 S. spontaneum that no longer exists; however, exactly one haploid (1n = 4x = 32) S. spontaneum, AP85-441, generated from a culture of octoploid SES2086, provides a foundation for assembly of a prototypical version of the sugarcane chromosome set. This study illuminates the hereditary blueprint and evolutionary history of one of our most important, and most complex, crop genomes.
Results
Genome sequencing and assembly
The genome size of AP85-441 was estimated at 3.36 Gbp by flow cytometry7. From a BAC library of AP85-441, 35,156 BAC clones were pooled into 712 libraries (mostly of 48 BACs; Supplementary Table 1), and individual BAC pools were sequenced independently by a Hiseq 2500 with PE250 (paired-end model and 250-bp read length), yielding 267.5 Gbp of data that were assembled using three different assemblers: ALLPATHS-LG8, SPAdes9 and SOAPdenovo210, yielding a 2.56-Gbp assembly with contig N50 of 7.4 kb (Supplementary Tables 2 and 3). To reduce fragmentation, 295 Gbp of data from the Pacific Biosciences (PacBio) RS II system (Supplementary Table 4) were used for self-correction and assembly by Canu11, incorporating assembled BAC sequences and correcting and polishing with 90× Illumina paired-end sequences, yielding 3.13 Gbp with contig N50 of 45 kb (Supplementary Table 5). The hybrid assembled contigs and BAC contigs correspond with ~99.72% accuracy (Supplementary Table 6).
High-throughput chromatin conformation capture (Hi-C) is an extension of chromosome conformation capture (3C) technology, in which cross-linked chromatin is digested with an appropriate restriction enzyme and then ligated to obtain an interacting fragment12. This approach, which was pioneered by Lieberman-Aiden et al.13 and Burton et al.14, was used previously in grasses in the assemblies of barley15 and wild emmer wheat16. To provide a scaffold for contig assembly, four Hi-C libraries were constructed from young leaves of AP85-441. Chimeric fragments representing the original cross-linked long-distance physical interactions were processed into paired-end sequencing libraries, then 1 billion 150-bp paired-end Illumina reads were produced and uniquely mapped onto the draft assembly contigs. Due to polyploidy, existing Hi-C scaffolding programs such as LACHESIS14 and SALSA17 link S. spontaneum allelic haplotypes together and are no longer suitable for this autopolyploid genome. We developed a Hi-C-based scaffolding algorithm (ALLHIC) that integrates four functions—pruning, partition, optimization and building—to select contigs specific for polyploid genome assembly (see Online Methods and Supplementary Figs. 1–3). The quality of Hi-C sequencing was evaluated using HiC-Pro18 (Supplementary Table 7 and Supplementary Fig. 4).
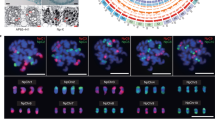
A Hi-C-based physical map was used to assemble 32 pseudo-chromosomes that anchor 2.9 Gbp of the genome, including 97% of the gene content. A high-density genetic map of 998,370 SNPs was used to verify the Hi-C assembly, supporting that the two methods are consistent in both chromosomal assignment and order for 89% of contigs (Supplementary Table 8). The 32 pseudo-chromosomes comprise 8 homologous groups with 4 sets of monoploid chromosomes: A, B, C and D (Fig. 1). A total of 219 (88.3 %) complete gene models among 248 ultraconserved core eukaryotic genes (CEGs) in CEGMA19 and 1,397 (97.01 %) among 1,440 conserved genes in BUSCO20 were recalled in our assembly (Supplementary Tables 9 and 10). Further, 1,624 million (97.01%) of 1,674 million Illumina short reads were alignable and covered 97.3% of the assembly (Supplementary Table 11). The assembly allowed us to predict 28 potential centromeric regions along the 32 chromosomes, with length ranging from 0.25 to 11.85 Mbp (Supplementary Table 12).
A set of 4 homologous chromosomes aligned to a single sorghum chromosome. The reduction of basic chromosomes from 10 to 8 in S. spontaneum is caused by chromosome fissions followed by translocations of two ancestral chromosomes homologous to sorghum chromosomes 5 and 8. Inset: sorghum stratum SSA, which is mirrored in the alignment of SsChr5 to SbChr08 and SsChr7 to SbChr05 at the tip of the short arm. Inset reproduced from ref. 25, http://www.plantcell.org, Copyright American Society of Plant Biologists. Mya, million years ago.
Allele-specific annotation
A high level of homologous gene retention was detected from sequencing multiple haplotypes in sugarcane, despite extreme autopolyploid redundancy21. In autopolyploid genomes, homologous genes at the same locus on homologous chromosomes are defined as alleles22. Using two rounds of MAKER followed by manual annotation to separate genes and alleles, we annotated 35,525 genes with alleles defined, including 4,289 (12.7%) genes with four alleles, 9,792 (27.6%) with three, 14,797 (41.7%) with two, and 6,647 (18.7%) with one. The total number of alleles was 82,773, with an average 2.3 alleles per gene. In unanchored sequences, 3,130 gene/alleles were annotated. We annotated 1,256 tandemly duplicated genes and 3,375 dispersedly duplicated paralogs (Table 1). The cytochrome P450 gene families illustrated the importance of annotating alleles in polyploid genomes, with a total of 1,465 manually annotated alleles in 387 genes (Supplementary Fig. 5). Without allele-specific annotation, the number of P450 genes in this genome would be 1,465, not 387.
Among the predicted gene models, 90.0% could be found in the sorghum genome23 and 80% in collinear positions. Comparison with rice, sorghum, maize and Arabidopsis (Supplementary Fig. 6) showed that among 21,661 gene families, 1,278 (6%) were unique to S. spontaneum. Gene Ontology (GO) enrichment analysis showed that these S. spontaneum-specific genes were enriched in a list of GO categories, including response to wounding/external stimulus, serine-type endopeptidase/peptidase inhibitor activity and ribosomal subunit (both false discovery rate (FDR) and P < 0.01, Fisher’s exact test; Supplementary Table 13).
AP85-441 contains 1,842 Mbp of repetitive sequences, accounting for 58.65% of the assembled genome (Supplementary Table 14). Long terminal repeat (LTR) retrotransposons account for 45.62% of the genome, including 14.19% Ty1/copia and 26.04% Ty3/gypsy. Kimura distances analysis indicated a more recent LTR burst (Supplementary Fig. 7), including Ty1/copia and Ty3/gypsy superfamilies that occurred between 0.72 and 2.9 million years ago.
Basic chromosome number reduction
The AP85-441 genome assembly showed chromosome reduction from 10 to 8 in S. spontaneum to involve a paleo-duplicated pair of chromosomes that have experienced frequent recombinations. Alignment to sorghum showed chromosome fissions in ancestral homologs of sorghum chromosomes 5 and 8, paleo-duplicated chromosome pairs A5 and A11 in grasses (Fig. 2). The ancestor of SbChr05 (A12) split into two major segments, C5S (A12S) and C5L (A12L)24, that translocated into ancestors of SbChr06 (A2) and SbChr07 (A5), respectively (Fig. 2c, event 1). The ancestor of SbChr08 (A11) split into two major segments, C8S (A11S) and C8L (A11L), and translocated into ancestors of SbChr09 (A6) and SbChr02 (A7 + A9), respectively. The short fragments that appear to be homologous between SbChr08 and SsChr5 and between SbChr05 and SsChr7 are remains of homeologous genes in sorghum stratum SSA formed 13.4 million years ago, well before sorghum and Saccharum diverged25. Strikingly, even the smaller SSA region in S5 and the larger SSA region in S8 were conserved in the rearranged AP85-441 genome, reflected in the sparse alignment of SsChr5 to SbChr08 and the dense alignment of SsChr7 to SbChr05 at the tip of the short arm (Fig. 1), validating the high quality and accuracy of the AP85-441 genome assembly.
a, Evolution of chromosome numbers in Poaceae, from n = 10 in sorghum to n = 8 in S. spontaneum. Chromosomes are represented with color codes to illuminate the evolution of segments from a common ancestor with 5 chromosomes. Ancestral genomes are labeled with AX (X is a number from 1 to 12) according to Salse et al.24. The S. spontaneum rearranged chromosomes are marked with dashed boxes. The inversion events that occurred in chromosome segments are shown with red curved arrows, and the chromosomes recombined with inverted chromosome segments are marked with red plus signs. b, Genomic alignments between Ss2, Ss5, Ss6 and Ss7 and Sb02, Sb05, Sb06, Sb07, Sb08 and Sb09 are shown; the inverted regions are marked with red ribbons. c, The genome duplications and rearrangements in S. spontaneum, Miscanthus (genetic map), and sorghum are shown. 1, The basic chromosome number reduction from 10 to 8 in S. spontaneum as described in the text. 2, Two inversions occurred after the first round of WGD as shown by pairs of inversions in SsChr2AB and SsChr7AB. 3, Three chromosomal fragments in SsChr6ABD are in an inverted position. 4, The ancestral chromosome SbChr07 (A5) fused into SbChr04 (A4) after an allopolyploidization event in Miscanthus27. 5, Four chromosomal fragments in SsChr4ABCD are in an inverted position. This is a sorghum-specific inversion in SbChr04 (A4), because the orientation of this chromosomal fragment is the same in rice, Miscanthus, and S. spontaneum.
Polyploidization in S. spontaneum
We assessed whether the two rounds of whole-genome duplication (WGD) affecting sugarcane were allopolyploidization followed by autopolyploidization, as proposed26, or just two rounds of autopolyploidization. Although the sequenced genome is haploid, each gamete contains four sets of homologous or hom(e)ologous chromosomes, representing two WGDs (that is, from one to two to four). Comparison among hom(e)ologous haplotypes A, B, C and D revealed 7.7 million SNPs, 1.03 million short indels and 3,637 structural variations (SVs), accounting for 11.2 Mbp of sequence and indicating heterozygosity of 0.98% in the S. spontaneum AP85-441 genome (Fig. 3 and Supplementary Table 15). To exploit the fact that paralogs are often located on all four of a set of hom(e)ologous chromosomes, we developed a framework to carry out a more sensitive study of gene pair similarities (Supplementary Note). However, no clear partition reflecting two events could be inferred, with each of three complementary approaches suggesting random association among the four members of most homologous series.
The rings indicate (from outermost to innermost) monoploid genome in Mbp (a), SNP density among haplotypes (b), gene density (c), expression (d) and nucleotide diversity (e). HA, HB, HC and HD indicate four haplotypes in ring B, respectively, and these four haplotypes were ordered from outside to inside in rings C, D and E.
It is clear, however, from comparisons of chromosomal rearrangements that there were two discrete WGDs, rather than a single event. Two fissions in ancestral homologs of sorghum chromosomes SbChr05 and SbChr08 that resulted in translocations to a set of two chromosomes each occurred before the two rounds of WGDs in Saccharum and after the divergence of Saccharum and Miscanthus (Fig. 2c). However, inversions in ancestral SbChr08S and SbChr08L in two pairs, SsChr2AB and SsChr7AB, indicate that these occurred after the first WGD but before the second (Fig. 2c, event 2). Among three regions showing collapsing of homologous sequences (upper region of SsChr1C, middle region of SsChr3D and upper region of SsChr8C), SsChr3B and SsChr8A have about 2× greater depth of Illumina short reads, suggesting that they are the collapsed homologs. The SsChr1C region showed equal distribution among three homologs, indicating a deletion in SsChr1C (Supplementary Table 16 and Supplementary Fig. 8).
Two inversions involving single chromosomes, ancestral SbChr05L (A12L) (homologous chromosome C of SsChr6) (Fig. 2c, event 3) and the bottom of SsChr5C (Fig. 2c, event 3), presumably occurred after the two rounds of WGD. Chromosome reduction in Miscanthus was caused by fusion of one set of chromosomes homologous to SbChr04 and SbChr0727. Inversions involving all four homologous chromosomes between SsChr4ABCD appear to have occurred before the two rounds of WGD, but it is actually an inversion that occurred in SbChr04 after Saccharum and Sorghum diverged from a common ancestor (Fig. 2c, event 5). These analyses indicate that the two rounds of WGD are autopolyploidization and that they occurred with a brief time in between.
Allelic expression dominance
The homologous genome expression levels of the four homologous genomes were similar in examined tissues (Supplementary Fig. 9), indicating no significant global homologous genome dominance in S. spontaneum. To mitigate differential gene content among the homologous genomes, we further extracted 4,289 sets of genes with four alleles in high collinearity from AP85-441 (Supplementary Fig. 10), but overall gene expression level from each haplotype was similar for the four homologous genomes (Supplementary Fig. 9). These results are not surprising—even recent allopolyploids such as Brassica napus28, Gossypium hirsutum29, Triticum aestivum30 and Brassica juncea31 displayed no homologous chromosome dominance. S. spontaneum is a recent autopolyploid, and homologous genomes are fluid and changing after each meiosis.
For breeding polyploid crops such as sugarcane, the segregation of alleles with different expression levels may contribute to the segregation of traits in a breeding population. To simplify the analysis of the allelic expression, we sorted the expression levels of four alleles for 4,289 genes in descending order from I to IV. Three allele pairs (I/II, II/III and III/IV) were compared for analyzing the differentially expressed alleles (Supplementary Figs. 11 and 12). We defined the pairs with less than twofold difference in expression within a pair as neutral and others as non-neutral. Of 4,289 genes, on average, 37.6% displayed neutral expression and 62.4% displayed non-neutral expression, suggesting that the expression of alleles varied. We further analyzed the variations in gene number among tissues; the numbers of both neutral and non-neutral genes were similar among the examined tissues. However, the genes of these two expressional patterns varied among examined tissues. On average, 36.3% of the neutral and 56.4% of the non-neutral genes were conserved across all the examined tissues (Supplementary Fig. 13).
Nicotinamide adenine dinucleotide phosphate-malic enzyme-type C4 pathway
The C4 photosynthesis pathway was discovered in sugarcane32,33. We identified 24 genes for 7 key enzymes related to the nicotinamide adenine dinucleotide phosphate-malic enzyme (NADP-ME) C4 pathway (Supplementary Fig. 14). Increased expression of core C4 enzymes played a major role in the evolution of C4 photosynthesis34. Based on gene expression and phylogenetic analysis, 8 genes—SsCA1, SsCA2, SsPEPC1, SsPEPC-k1, SsNADP-MDH2, SsNADP-ME2, SsPPDK1 and SsPPDK-RP2—were identified as C4-type genes (Supplementary Table 17). A tandem duplication of SsNADP-ME2, SsNADP-ME1, also displayed a C4 expression profile similar to that of SsNADP-ME2. But the ortholog of SsNADP-ME1 in maize, ZmNADP-ME (GRMZM2G122479), displayed non-C4-type expression35, suggesting that neofunctionalization of SsNADP-ME1 for C4 in sugarcane occurred after the divergence of maize and tribe Andropogoneae.
Sugar transporters
Sucrose transporters (SUTs) are hypothesized to load sucrose into the phloem of leaf minor veins and also function to retrieve sucrose from the apoplasm during transport36,37,38. In the step prior to phloem loading, SWEETs (sugars will eventually be exported transporters) are potentially responsible for sucrose efflux into the cell wall space from phloem parenchyma cells and the bundle sheath39,40. SWEETs play various important roles in multiple physiological processes41. In sugarcane and sweet sorghum, the stems are the principal sink tissues that store very high concentrations of sugars within the parenchyma cells42,43,44. Tonoplast sugar transporters (TSTs) have been characterized as sucrose transporters highly associated with vacuolar sucrose accumulation from sugar beet taproot45, sugarcane46 and sweet sorghum stems47 and watermelon fruit48. Whereas there are 3 TST genes in the sorghum genome47, the family has expanded in the S. spontaneum genome, which has 4 genes consisting of 13 homologs. Hence, it is reasonable to hypothesize that TSTs are the most promising players to sequester sucrose into the vacuoles of the sugarcane stem46,47,49.
In the S. spontaneum genome, we identified 123 sugar transporters from 9 subfamilies, including 4 in the TST family, 4 in the vacuolar glucose transporter (VGT) family, 3 in the plastidic glucose transporter (pGlcT) family, 4 in the inositol transporters (INT) family, 31 in the polyol transporter (PLT) family, 14 in the early response to dehydration 6-like (SFP) family, 6 in the SUT family, 22 in the SWEET family, and 35 in the sugar transporters family or hexose transporter family (STP) (Supplementary Table 18). Phylogenetic analysis of those sugar transporters showed gene family expansion in the STP and PLT families compared with sorghum (22 in STP and 17 in PLT), rice (21 in STP and 11 in PLT) and Arabidopsis (14 in STP and 9 in PLT) (Supplementary Fig. 15). Tandem duplication analysis indicated that 19 and 23 genes of STP and PLT, respectively, could be assigned to tandem duplication, compared with 11 and 9 genes in sorghum. The cause of STP and PLT family expansions in S. spontaneum is tandem duplication.
Disease resistance genes
S. spontaneum contributed disease resistance genes to modern sugarcane hybrid cultivars. We identified 361 sequences putatively encoding nucleotide-binding site (NBS) proteins, including 22 N-type, 169 NL-type, 68 CN-type and 102 CNL-type. The number of NBS-encoding genes is larger than that in sorghum50, owing to the species-specific tandem duplication in S. spontaneum. Surprisingly, 80% of the NBS-encoding genes located in the four rearrangement chromosomes (SsChr02, SsChr05, SsChr06 and SsChr07) and 51% of those were in the rearranged regions, including SsChr5 (Sb05S) 57.6–89.1 Mbp, SsChr6 (Sb05L) 54.6–90.6 Mbp, SsChr7 (Sb08S) 62.0–83.3 Mbp, SsChr2 (Sb08L) 98.5–125.9 Mbp (Supplementary Table 19). Resistance genes are seven times more likely to locate in the four rearranged regions than in other chromosomes or regions (P < 2.2 × 1016, Fisher’s exact test; Supplementary Table 20).
S. spontaneum fraction in hybrid sugarcane cultivars
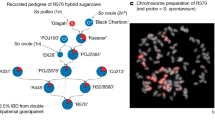
Modern sugarcane cultivars are the product of complex and repeated hybridization between S. officinarum and S. spontaneum, resulting in complex hybrids with chromosome numbers and morphologies that differ from those of their progenitors. Previous studies estimated that the S. spontaneum genome contributed approximately 10–20% of the modern hybrid sugarcane genome. In the modern hybrid sugarcane SP80-3280, approximately 12.25% of sequences are contributed by S. spontaneum. We mapped the sequences back to the AP85-441 genome, and they were randomly and evenly distributed, not constituting a set or sets of chromosomes as expected (Fig. 4). Analysis of the integrated S. spontaneum fraction in 15 resequenced hybrid genomes also yielded random distribution throughout the genome (Supplementary Fig. 16).
Origin and genetic diversity of S. spontaneum
Most of the genetic diversity found within S. spontaneum has not been introgressed into commercial sugarcane, and in principle, this germplasm represents a rich source of desirable agronomic traits related to stress tolerance and biomass accumulation51. S. spontaneum has a broad natural range extending throughout Asia, the Indian subcontinent, the Mediterranean and Africa52, and natural populations display a wide range of phenotypic, genetic and ploidy-level diversity.
In practice, however, nucleotide diversity (π) across S. spontaneum was estimated to be 0.00021 ± 0.000002 (Supplementary Tables 21 and 22 and Supplementary Fig. 17), much lower than that in other clonally propagated crops such as potato53, cassava54, grape55 and citrus56. We resequenced 64 diverse S. spontaneum accessions from the world germplasm collection, identifying 4.48 million high-confidence variants that included 3,961,408 SNPs, 201,854 insertions and 291,346 deletions, averaging 1.52 variants per kb. We identified 671,265 variants (15%) in genic regions, including 41,960 synonymous, 101,826 nonsynonymous and 491,493 intronic variants.
Both principal component analysis (PCA) and admixture-based analyses clustered the 64 S. spontaneum accessions into three distinct groups (Fig. 5a,c; for other K-values, see Supplementary Fig. 18a,b) that were also supported by phylogenetic relationships among the 64 accessions inferred by bootstrapping and geographic origins (Supplementary Table 23), with group 1 originating from China, the Philippines, Indonesia and Papua New Guinea and groups 2 and 3 originating from India, Pakistan and Iran. The regions of Pan-Malaysia might be the ancient hybrid zones among three groups. Ploidy varies widely within the three groups, from 6x to 16x. By mapping the ploidy levels on the bootstrapped tree (Fig. 5b), the topology shows that the accessions of different ploidy levels (from hexaploid to hexadecaploid) diverged independently from ancestors in three groups, suggesting that the fluid ploidy levels may have independently evolved from ancestral progenitors.
a, Principal components (PCs) of accession variation. b, Bootstrapped tree of 64 S. spontaneum accessions based on genetic distance. Color bars indicate accessions with different ploidy levels (P6, hexaploid; P7, heptaploid; P8, octoploid; P9, enneaploid; P10, decaploid; P11–16, from hendecaploid to hexadecaploid). The scale bar shows substitutions per site. c, ADMIXTURE plot for S. spontaneum, showing the distribution of K = 3 genetic clusters with the smallest cross-validation error.
Regions of S. spontaneum with larger-scale chromosomal rearrangements compared with sorghum have higher genetic diversity (higher π value) than non-rearranged regions and may have undergone much stronger balancing selection (Supplementary Table 22 and Supplementary Fig. 19). Although several individual chromosomes do not show significant differences, comparisons averaging values on all chromosomes show nucleotide diversity (π) in rearranged regions (0.00025 ± 0.00003) to be much higher than in non-rearranged regions (0.00021 ± 0.00001, P = 0.000234). The Tajima’s D in rearranged regions (−0.659 ± 0.052) is much higher than in non-rearranged regions (−0.720 ± 0.011, P = 0.005013). SNP density is also higher in rearranged regions (360.27 ± 48.41) than in non-rearranged regions (297.46 ± 12.65, P = 0.001798). In addition, the GO enrichment analyses showed that the non-rearranged regions are enriched in GOs related to basic life cycles, primarily in photosynthesis, respiration and ATP synthesis (both FDR and P < 0.05, Fisher’s exact test; Supplementary Table 23), whereas the rearranged regions were enriched in many GOs related to secondary life processes, whole celluar process and intracellular anabolic and catabolic processes, transmembrane transport and ion binding (both FDR and P < 0.05, Fisher’s exact test; Supplementary Table 24).
An intriguing question is whether genomic rearranged regions might have had a role in adaptation to different habitats. The rearranged regions in S. spontaneum with high levels of genomic diversity (π value) might result from a preponderance of adaptive genes related to habitat or stress adaptation, such as responses to various abiotic stresses (drought, salinity, alkaline, metal ions and so on), which are controlled by genes of whole celluar process and intracellular anabolic and catabolic processes, transmembrane transport and ion binding, as detected in these regions. Conversely, it is also meaningful that the non-rearranged regions mainly controlling basic life cycles maintain a lower level of genomic diversity. The rearranged regions have undergone stronger balancing selection after the polyploidization events. Adding further support to this notion is that 80% of the NBS-encoding genes are located on the four rearrangement chromosomes.
Discussion
The identification of 80% of disease resistance genes on rearranged chromosomes suggests that reduction of basic chromosome number might have contributed to the retention of disease-resistance genes. Following chromosome fissions and translocations in a diploid ancestor, translocated fragments may have undergone little recombination. Following WGD, additional chromosomal rearrangements in these translocated regions may have further suppressed recombination (Fig. 1). Population genomic analyses detected balancing selection in these rearranged regions, a mechanism to maintain genetic diversity. It is likely an unintended consequence that these rearranged chromosome arms are enriched with NBS-encoding genes, resulting in more disease-resistance genes being retained in S. spontaneum, which leads to higher resistance to disease and abiotic stresses in S. spontaneum than in other Saccharum species and makes S. spontaneum the source of disease and stress tolerance in sugarcane breeding program.
Integration of S. spontaneum chromosome segments into modern sugarcane hybrid cultivars by three to four generations of backcrossing at random would result in about one set of monoploid S. spontaneum chromosomes. The S. spontaneum fraction of the sugarcane hybrid cultivar SP80-3280 and of 15 resequenced hybrid genomes each appear randomly distributed in the reference AP85-441 genome, indicating random recombination of homologous chromosomes in different accessions that have undergone many rounds of meiosis after their separation. This is indirect evidence that S. spontaneum is autopolyploid, and it reinforces the importance of allele-specific annotation for mining effective alleles of resistance genes in hybrid cultivars.
Defining alleles in an autopolyploid genome clarifies gene or gene family analysis, as demonstrated in P450 and other gene families. This reference genome offers substantial new knowledge and unprecedented genomic resources for sugarcane breeders and researchers to mine disease resistance and other alleles in rearranged chromosomes from historic hybrid cultivars, and to track them in breeding populations to shorten the 13-year breeding cycle.
URLs
FGENESH online version, http://www.softberry.com/berry.phtml?topic=fgenesh&group=help&subgroup=gfind; FigTree, http://tree.bio.ed.ac.uk/software/figtree/; National Center for Biotechnology Information non-redundant (NCBI NR) database for Oryza sativa, ftp://ftp.ncbi.nih.gov/blast/db; RepeatModeler, http://www.repeatmasker.org/RepeatModeler/.
Methods
Genome sequencing
Illumina short reads sequencing
DNA was extracted from leaf tissue of a single soil-grown plant using the Qiagen DNeasy Plant Mini Kit and applied to 280-bp and 500-bp paired-end library construction using the NEBNext Ultra DNA Library Prep Kit for Illumina sequencing. Sequencing was performed using the Illumina HiSeq 2500 platform.
Construction of BAC libraries and sequencing
Nuclei were isolated from the young leaf tissues of AP85-441 following the method described by Ming et al.57. The high-molecular-weight DNA embedded in agarose was partially digested using HindIII. Fractions at approximately 100 kb were recovered and cloned into the pSMART BAC vector (Lucigen). A total of 38,400 BAC clones were constructed and selected for sequencing; 48 BAC clones were pooled together, and DNA libraries were prepared with the PhasePrep BAC DNA Kit (Sigma) following the manufacturer’s protocols. BAC DNA libraries were sequenced using the Illumina HiSeq 2500 with a 250-bp paired-end sequencing strategy.
PacBio library construction and sequencing
More than 5 µg of sheared and concentrated DNA was applied to size selection by the BluePippin system. Approximately 20-kb SMRTbell libraries were prepared according to the released protocol from PacBio. A total of 176 Single-Molecule, Real-Time (SMRT) cells were run on the PacBio RS II system with P6-C4 chemistry.
Hi-C library construction and sequencing
Four Hi-C libraries were created from tender leaves of AP85-441 at BioMarker Technologies Company as described previously58. Briefly, the leaves were fixed with formaldehyde and lysed, and then the cross-linked DNA was digested with HindIII overnight. Sticky ends were biotinylated and proximity-ligated to form chimeric junctions that were enriched for and then physically sheared to a size of 500–700 bp. Chimeric fragments representing the original cross-linked long-distance physical interactions were then processed into paired-end sequencing libraries, and 1,001 million 150-bp paired-end reads were produced on the Illumina HiSeq X Ten platform.
Genome assembly overview
The sugarcane AP85-441 contig-level assembly incorporated sequencing data from a mixture of sequencing technologies (Supplementary Fig. 1), including BAC pools sequenced with Illumina HiSeq 2500 and whole-genome shotgun sequencing with PacBio RS II as well as Hi-C reads, followed by Illumina short reads polishing. Each BAC pool was independently assembled using ALLPATHS-LG8, SPAdes9 and SOAPdenovo210, and best results were retained. For PacBio assembly, Canu v1.511 was used, as it is capable of avoiding collapsed repetitive regions and haplotypes. Self-correction was performed with parameter corOutCoverage = 100, which allowed us to correct all of the input PacBio reads. The corrected reads, along with BAC-assembled contigs, were imported to the assembly step. Chromosomal assembly was constructed based on proximity-guided assembly using our newly developed program, ALLHIC, which is designed for polyploid genome scaffolding (see Supplementary Note for details).
Genome annotation
Repeat prediction
We first customized a de novo repeat library of the genome using RepeatModeler (see URLs), which can automatically execute two de novo repeat finding programs, including RECON (version 1.08)59 and RepeatScout (version 1.0.5)60. The consensus transposable element (TE) sequences generated above were imported to RepeatMasker (version 4.05)61 to identify and cluster repetitive elements. Unknown TEs were further classified using TEclass (version 2.1.3)62. To identify tandem repeats within the genome, the Tandem Repeat Finder (TRF) package (version 4.07)63 was used with the modified parameters of ‘1 1 2 80 5 200 2,000 –d –h’ to find high-order repeats. Telomeres and centromeres were identified based on the .dat output files above. Repeat sequences with more than ten monomers ‘AAACCT’ were identified as telomeres. For centromere identification, we used a similar method described in the Oropetium thomaeum genome64. The largest repeat arrays were identified and clustered as centromeres. To further investigate LTRs, we applied the LTR_retriever pipeline65, which can integrate results from public programs such as LTR_FINDER66 and LTRharvest67 and efficiently remove false positives from the initial predictions. The predicted LTRs were further classified into intact and non-intact LTRs, and the insertion time was estimated as T = K/2μ (K is the divergence rate, and μ is the neutral mutation rate; the default is 1.38 × 10−8 in LTR_retriever) using the scripts implemented in the LTR_retriever package65.
Gene annotation
To get high-quality annotation of protein-coding genes, we carried out two rounds of MAKER running, following extensive and careful manual inspections in JBrowse68.
In the first round of MAKER running, ten selected RNA sequencing (RNA-seq) samples were imported into Trinity de novo assembly and genome-guided assembly pipelines with default parameters69. RSEM was used to calculate transcript abundance70. Transcripts with FPKM (fragments per kilobase of exon per million fragments mapped) < 1 and iso-percentage < 3% were removed from further analysis. The filtered transcripts were imported to the PASA program for construction of comprehensive transcripts, as PASA is able to take advantage of the high sensitivity of reference-based assembly while leveraging the ability of de novo assembly to detect novel transcripts71. The PASA-assembled transcripts described above were used for training. The nearly ‘full-length’ transcripts were evaluated by comparing with the UniProt plant protein database (last accessed on 8 December 2016), and proteins that were covered at least 95% were retained as candidates. Then ab initio gene predictors, including SNAP72, GENEMARK73 and AUGUSTUS74, were each trained with those selected proteins. After that, the MAKER pipeline was used to integrate multiple tiers of coding evidence, including ab initio gene prediction, transcript evidence and protein evidence and generate a comprehensive set of protein-coding genes.
In the second round of MAKER running, the predicted gene models with AED score equal to 0 were extracted for retraining using SNAP72, GENEMARK73 and AUGUSTUS74. In addition, the RNA-seq reads were mapped to the AP85-441 genome using HiSAT275 version 2.10 and reassembled using StringTie76 version 1.3.4, which is a reference-based RNA assembler. Meanwhile, published full-length transcripts based on IsoSeq in sugarcane were also recruited for annotation77. The four haplotypes (A, B, C and D) were split into four sub-genomes, each containing eight pseudo-molecules.
Gene structures were visualized in JBrowse68 along with RNA-seq-assembled transcripts and homologs from the sorghum, maize and rice genomes. We compared the two rounds of MAKER annotation and selected the better ones if their structures were better supported by homologous proteins or RNA-seq-assembled transcripts. Genes in the first round of annotation were kept if their structures did not improve significantly in the second round.
Extensive manual inspection of the annotation identified that 28,306 gene models had a significant difference in protein length or sequence similarity compared with other reported protein sequences in the NCBI NR database. The corresponding genomic DNA sequences of these genes were extracted for further careful annotation using the online version of FGENESH (see URLs) with pretrained parameters for sorghum genes. Genes with significant improvement were replaced with the FGENESH annotation.
BUSCO20 version 3 was used for evaluation of annotation completeness. Out of 1,440 conserved genes, 1,397 (97.1%) were re-annotated in the AP85-441 genome, among which 1,101 (76.5%) were complete and duplicated BUSCO genes.
Allelic variation analysis
Construction of a monoploid genome
To compare the allelic variations among the four haplotypes, we first generated a monoploid genome. The concept of the monoploid genome is aimed at retaining consensus sequences among four haplotypes and covering as many genetic materials as possible. The longest pseudo-molecule was used as reference for each set of haplotypes, and the other three haplotypes were mapped against the reference for SNP/indel and SV calling using the nucmer78 program. Mapping results were filtered, and only the best hits were retained. The program show-snps, implemented in the MUMmer package78, was used to identify SNPs and indels with parameters –Clr, which means only SNPs/indels from ambiguous mapping were reported. Consensus sequences were extracted using a homemade PERL script. Insertions larger than 50 bp were identified on Assemblytics79, a Web-based SV analytics tool, and further inserted into the reference genome. Finally, a monoploid genome, containing eight representative pseudomolecules and 785 Mbp of sequence, was generated for further analysis (Fig. 3).
Identification of alleles
Identification of alleles in the AP85-441 genome was based on two strategies: (1) synteny-based and (2) coordinate-based approaches. Interhaplotype syntenic blocks were identified by MCScanX80 and organized into a four-column table containing allele A, B, C or D. In addition, genes that were not shown in that table were mapped to the monoploid genome using GMAP81, and those with at least 50% overlaps on coordinates were considered as potential alleles. Sequence similarities were checked among alleles on the basis of reciprocal blast, and genes without significant similarities to any other allele were removed from the table.
Analysis of allelic variations
We use a reference-based strategy to identify SNPs/indels and SVs. Similar to the approach described in the previous section, the nucmer78 program was used to map haplotypes A, B, C and D to the monoploid genome and SNPs were extracted from ambiguous best mapping. Short indels (1–10 bp) and large structural variations were recalled by Assemblytics79 on the basis of the alignments above.
Identification of the sequences in hybrid sugarcane that originate from S. spontaneum
The SP80-3280 genome was first masked using the customized TE library and then split into 1-kb fragments. Each of the fragments was blast against the AP85-441 and LA-purple (unpublished) masked genomes, respectively, and the mapping score was calculated for each blast hit using the following formula:
where S indicates mapping score, N indicates the number of matched bases and I indicates identity in each blast hit.
Fragments were further classified as sequences from AP85-441 and sequences from LA-purple if they had a best mapping score in the corresponding category. Sequences were classified as fragments from both if they had similar mapping scores (<5% difference) in the LA-purple and AP85-441 genomes.
Allelic expression dominance
Tissues including leaves, stems and roots were collected from mature plants, and RNA-seq analysis in this part was based on these three samples. RNA-seq reads were trimmed by Trimmomatic and then mapped to the AP85-441 genome by HiSAT275. FPKM was calculated on the basis of unique mapping reads using the StringTie package76. To analyze allelic expression dominance, we extracted 4,289 genes with full of four alleles from the AP85-441 annotation files. To simplify the analysis of the allelic expression, the expression levels of the genes were sorted in descending order from I to IV. Three allele pairs (I/II, II/III and III/IV) were compared to analyze the differentially expressed alleles. Allele pairs with less than twofold difference in expression were defined as a neutral pair and all others as non-neutral.
Resequencing and population analysis
Reads mapping and variants calling
The raw pair-end reads of 64 S. spontaneum accessions were trimmed to remove the adaptors and low-quality bases using Trimmomatic82 after quality control by FastQC83. The reads were filtered with a sliding window of size 7, with average Phred score scale of 20 within the window. The trimmed reads were mapped to the S. spontaneum genome using Bowtie284 with default parameters. The mapped reads were sorted, and duplicated reads were removed using SAMtools85. We estimated the rate of uniquely mapped reads outputted from both BWA86 and Bowtie284. Bowtie2 generated ten times as many uniquely mapped reads as BWA did (Supplementary Fig. 21).
The Realigner Target Creator and Indel Realigner programs from the Genome Analysis Toolkit (GATK) package87 were used for global realignment of reads around indels from the sorted BAM files. The HaplotypeCaller was used to estimate the SNPs and indels for putative diploids using the default parameters. The HaplotypeCaller outputted 42,585,337 unfiltered variants (SNPs and indels). The distribution of calling depths (DP) of each raw variant was estimated as a criterion for variants filtering. Low depths and repetitive variants were removed from the raw VCF file if they had DP < 2 or DP > 45, minQ < 30. Variants with more than 15% missing data were removed. These filtering strategies reduced the raw unfiltered set of 42.59 million variants (SNPs and Indels) to the working set of 4.48 million (4,476,608) variants. SnpEff (v3.6c)88 was used to assign variants effects on the basis of gene models from S. spontaneum genome annotation. The variants sites were annotated as the SNPs and Indels, as well as intergenic and genic regions (including the synonymous, nonsynonymous, intronic, upstream and downstream variants).
Genome-wide genetic diversity estimation
Population genetic statistics of SNP density, π and Tajima’s D were calculated directly from the filtered VCF file in 1,000-kb window and 500-kb step for π, and non-overlapping intervals for SNPs density and Tajima’s D in VCFtools89. The high confidence 4,476,608 variant set was used for statistical estimations.
PCA
PCA was performed using the GCTA software on the filtered 4,476,608 variants. The input Plink binary files are transformed from the filtered VCFs file using VCFtools89 and PLINK90. The top three principal components were used for assigning the 64 accessions and downstream population structure analysis.
Phylogeny
Bi-allelic and polymorphic SNPs (3,969,408) were used for reconstructing the phylogenetic relationships among 64 accessions. Before tree construction, we filtered and pruned the SNPs with MAF (minor allele frequency) < 0.2, missing rate > 0.15, and LD (linkage disequilibrium) threshold = 0.2. Finally, a total of 37,617 SNPs were selected for the constructed tree using SNPhylo software. The multiple consensus sequences were aligned using MUSCLE91. Maximum likelihood trees were constructed using the maximum likelihood method by running DNAML programs in the PHYLIP package92. In addition, a bootstrapped tree was constructed by bootstrapping (bootstrap = 10,000) analysis using the PHANGORN package93. Figtree v.1.4 (see URLs) was used to visualize the trees.
Population genetic structure
Ancestral population stratification among 64 accessions was inferred using Admixture software. The optimal ancestral population structure was estimated from the same variants set with STRUCTURE94 using ancestral population sizes K = 1–20 and choosing the population with the smallest cross-validation error. The parameter standard errors were estimated using bootstrapping (bootstrap = 200) when doing the admixture analyses. DISTRUCT95 was used to plot the population stratification results for K = 1 through K = 20 (Supplementary Fig. 18).
Differentiation of genomic diversity among four homologous haploid sets
The reads mapped to each of four homologous haploid sets (A, B, C and D) of the AP85-441 genome were retrieved for each of 64 accessions using SAMtools85 and Bedtools96. The four sets of retrieved reads for each of 64 accessions were mapped separately to each of eight chromosomes in a consensus monoploid genome using Bowtie2 with default parameters. The variants were called from a cohort of 256 BAM files generated from the previous step for each of the eight chromosomes. The HaplotypeCaller of GATK was used to estimate the SNPs and indels for putative diploids using the default parameters. The HaplotypeCaller outputted 17,531,765 unfiltered variants (SNPs and indels). The distribution of calling depths (DP) of each raw variant was estimated as a criterion for variant filtering. Low depths and repetitive variants were removed from the raw VCF file if they had DP < 1 or DP > 5, minQ < 20. We allowed the variant sites with max-missing rate as 50%. These filtering strategies reduced the raw unfiltered set of variants (SNPs and indels) to the working set of 68,911 variants. These filtering strategies reduced the raw unfiltered set of variants (SNPs and indels) to the working set of 68,911 variants. The working variant set was then used for estimating the population genetics statistics π among four homologous haploid sets.
Genomic diversity of genomic rearranged regions
To test whether genomic rearranged regions (RAR) have a genetic difference from non-rearranged regions (non-RAR), we compared the population genetic statistics π, SNP density, and Tajima’s D between rearranged and non-rearranged regions in each of four sets (A, B, C and D) of chromosomes 2, 5, 6 and 7. The genomic rearranged regions inferred by collinear dot plot and alleles phasing are shown Supplementary Table 21. We used the T-test and Mann-Whitney U test with the one-tailed hypothesis to compare the differences of statistics (π, SNP density and Tajima’s D) between RAR and non-RAR. To find the difference in gene functions between RAR and non-RAR, we conducted GO enrichment analysis for the gene models in RAR and non-RAR. We first blastX the S. spontaneum gene models in the NCBI NR database of Oryza sativa (see URLs). Then, the functional annotation and GO enrichment analyses of gene models were conducted in Blast2Go v4.1 software97. We used gene models of RAR or non-RAR as tested gene sets and the whole gene models as reference. The significance of enrichment was valued using the Fisher’s exact test.
Genomic diversity among different polyploidy accessions
To test the effects of polyploidization on genetic diversity, we compared the population nucleotide diversity (π) among accessions with different ploidy levels. We used a 1,000-kb sliding window and a 500-kb step to calculate the values of each statistic. In addition, we divided the 64 accessions into four groups (ploidy 6, 8, 10 and 13–16) depending on their ploidy level. The four groups were used to calculate the pairwise Weir and Cockerham’s FST between the two of them using VCFtools version 0.1.12b46 with a 1,000-kb sliding window and a 500-kb step.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The genome assembly and gene annotation have been deposited in the NCBI database under accession number QVOL00000000, BioProject number PRJNA483885 and BioSample number SAMN09753102. The data can also be downloaded from the following link: http://www.life.illinois.edu/ming/downloads/Spontaneum_genome/.
Change history
13 November 2018
In the version of this article originally published, the accession codes listed in the data availability section were incorrect and the section was incomplete. The text for this section should have read “The genome assembly and gene annotation have been deposited in the NCBI database under accession number QVOL00000000, BioProject number PRJNA483885 and BioSample number SAMN09753102. The data can also be downloaded from the following link: http://www.life.illinois.edu/ming/downloads/Spontaneum_genome/.” The errors have been corrected in the HTML and PDF versions of the article.
References
Watson, A. M. Agricultural Innovation in the Early Islamic World: The Diffusion of Crops and Farming Techniques, 700–1100 (Cambridge Univ. Press, 2008).
Mintz, S. W. Sweetness and Power: The Place of Sugar in Modern History (Penguin, 1986).
Roach, B. T. Nobilization of sugarcane. Proc. Int. Soc. Sugar Cane Technol. 14, 206–216 (1972).
Brandes, E. W. & Sartoris, G. B. Sugarcane: its origin and improvement. Yearb. U.S. Dep. Agric. 1936, 561–624 (1936).
D’Hont, A. et al. Characterisation of the double genome structure of modern sugarcane cultivars (Saccharum spp.) by molecular cytogenetics. Mol. Gen. Genet. 250, 405–413 (1996).
Moore, P. H., Nagai, C. & Fitch, M. M. M. Production and evaluation of sugarcane haploids. Proc. Int. Soc. Sugar Cane Technol. 20, 599–607 (1989).
Zhang, J. et al. Genome size variation in three Saccharum species. Euphytica 185, 511–519 (2012).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl. Acad. Sci. USA 108, 1513–1518 (2011).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18 (2012).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Dekker, J. The three ‘C’ s of chromosome conformation capture: controls, controls, controls. Nat. Methods 3, 17–21 (2006).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Mascher, M. et al. A chromosome conformation capture ordered sequence of the barley genome. Nature 544, 427–433 (2017).
Avni, R. et al. Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 357, 93–97 (2017).
Ghurye, J., Pop, M., Koren, S. & Chin, C.-S. Scaffolding of long read assemblies using long range contact information. BMC Genomics 18, 527 (2017).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Garsmeur, O. et al. High homologous gene conservation despite extreme autopolyploid redundancy in sugarcane. New Phytol. 189, 629–642 (2011).
Osborn, T. C. et al. Understanding mechanisms of novel gene expression in polyploids. Trends Genet. 19, 141–147 (2003).
Paterson, A. H. et al. The Sorghum bicolor genome and the diversification of grasses. Nature 457, 551 (2009).
Salse, J. et al. Identification and characterization of shared duplications between rice and wheat provide new insight into grass genome evolution. Plant Cell 20, 11–24 (2008).
Wang, X., Tang, H. & Paterson, A. H. Seventy million years of concerted evolution of a homoeologous chromosome pair, in parallel, in major Poaceae lineages. Plant Cell 23, 27–37 (2011).
Kim, C. et al. Comparative analysis of Miscanthus and Saccharum reveals a shared whole-genome duplication but different evolutionary fates. Plant Cell 26, 2420–2429 (2014).
Swaminathan, K. et al. A framework genetic map for Miscanthus sinensis from RNAseq-based markers shows recent tetraploidy. BMC Genomics 13, 142 (2012).
Chalhoub, B. et al. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, 950–953 (2014).
Said, J. I. et al. A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G. hirsutum × G. barbadense populations. Mol. Genet. Genom. 290, 1003–1025 (2015).
International Wheat Genome Sequencing Consortium (IWGSC). A chromosome-based draft sequence of thehexaploid bread wheat (Triticum aestivum) genome. Science 345, 1251788 (2014).
Yang, J. et al. The genome sequence of allopolyploid Brassica juncea and analysis of differential homoeolog gene expression influencing selection. Nat. Genet. 48, 1225–1232 (2016).
Kortschak, H. P., Hartt, C. E. & Burr, G. O. Carbon dioxide fixation in sugarcane leaves. Plant Physiol. 40, 209–213 (1965).
Hatch, M. D. & Slack, C. R. Photosynthesis by sugar-cane leaves. A new carboxylation reaction and the pathway of sugar formation. Biochem. J. 101, 103–111 (1966).
Sage, R. F. The evolution of C4 photosynthesis. New Phytol. 161, 341–370 (2004).
Wang, L. et al. Comparative analyses of C4 and C3 photosynthesis in developing leaves of maize and rice. Nat. Biotechnol. 32, 1158–1165 (2014).
Julius, B. T., Leach, K. A., Tran, T. M., Mertz, R. A. & Braun, D. M. Sugar transporters in plants: new insights and discoveries. Plant Cell Physiol. 58, 1442–1460 (2017).
Baker, R. F. et al. Sucrose transporter ZmSut1 expression and localization uncover new insights into sucrose phloem loading. Plant Physiol. 172, 1876–1898 (2016).
Slewinski, T. L. & Braun, D. M. Current perspectives on the regulation of whole-plant carbohydrate partitioning. Plant Sci. 178, 341–349 (2010).
Williams, L. E., Lemoine, R. & Sauer, N. Sugar transporters in higher plants—a diversity of roles and complex regulation. Trends Plant. Sci. 5, 283–290 (2000).
Chen, L. Q. et al. Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335, 207–211 (2012).
Eom, J. S. et al. SWEETs, transporters for intracellular and intercellular sugar translocation. Curr. Opin. Plant Biol. 25, 53–62 (2015).
Bihmidine, S., Hunter, C. T., Johns, C. E., Koch, K. E. & Braun, D. M. Regulation of assimilate import into sink organs: update on molecular drivers of sink strength. Front. Plant Sci. 4, 177 (2013).
Patrick, J. W., Botha, F. C. & Birch, R. G. Metabolic engineering of sugars and simple sugar derivatives in plants. Plant. Biotechnol. J. 11, 142–156 (2013).
Bihmidine, S., Baker, R. F., Hoffner, C. & Braun, D. M. Sucrose accumulation in sweet sorghum stems occurs by apoplasmic phloem unloading and does not involve differential sucrose transporter expression. BMC Plant Biol. 15, 186 (2015).
Jung, B. et al. Identification of the transporter responsible for sucrose accumulation in sugar beet taproots. Nat. Plants 1, 14001 (2015).
Casu, R. E. et al. Tissue-specific transcriptome analysis within the maturing sugarcane stalk reveals spatial regulation in the expression of cellulose synthase and sucrose transporter gene families. Plant Mol. Biol. 89, 607–628 (2015).
Bihmidine, S., Julius, B. T., Dweikat, I. & Braun, D. M. Tonoplast sugar transporters (SbTSTs) putatively control sucrose accumulation in sweet sorghum stems. Plant Signal. Behav. 11, e1117721 (2016).
Ren, Y. et al. A tonoplast sugar transporter underlies a sugar accumulation QTL in watermelon. Plant Physiol. 176, 836–850 (2018).
Julius, B. T., Leach, K. A., Tran, T. M., Mertz, R. A. & Braun, D. M. Sugar transporters in plants: new insights and discoveries. Plant Cell Physiol. 58, 1442–1460 (2017).
Cheng, X. et al. A genomic analysis of disease-resistance genes encoding nucleotide binding sites in Sorghum bicolor. Genet. Mol. Biol. 33, 292–297 (2010).
Wang, L.-P. et al. Evaluation of sugarcane × Saccharum spontaneum progeny for biomass composition and yield components. Crop Sci. 48, 951–961 (2008).
Panje, R. & Babu, C. Studies in Saccharum spontaneum distribution and geographical association of chromosome numbers. Cytologia 25, 152–172 (1960).
Hardigan, M. A. et al. Genome diversity of tuber-bearing Solanum uncovers complex evolutionary history and targets of domestication in the cultivated potato. Proc. Natl. Acad. Sci. USA 114, E9999–E10008 (2017).
Bredeson, J. V. et al. Sequencing wild and cultivated cassava and related species reveals extensive interspecific hybridization and genetic diversity. Nat. Biotechnol. 34, 562 (2016).
Myles, S. et al. Genetic structure and domestication history of the grape. Proc. Natl. Acad. Sci. USA 108, 3530–3535 (2011).
Wu, G. A. et al. Genomics of the origin and evolution of Citrus. Nature 554, 311–316 (2018).
Ming, R. et al. Construction and characterization of a papaya BAC library as a foundation for molecular dissection of a tree-fruit genome. Theor. Appl. Genet. 102, 892–899 (2001).
Xie, T. et al. De novo plant genome assembly based on chromatin interactions: a case study of Arabidopsis thaliana. Mol. Plant 8, 489–492 (2015).
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Res. 12, 1269–1276 (2002).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21(Suppl. 1), i351–i358 (2005).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0, http://www.repeatmasker.org.
Abrusan, G., Grundmann, N., DeMester, L. & Makalowski, W. TEclass—a tool for automated classification of unknown eukaryotic transposable elements. Bioinformatics 25, 1329–1330 (2009).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
VanBuren, R. et al. Single-molecule sequencing of the desiccation-tolerant grass Oropetium thomaeum. Nature 527, 508–511 (2015).
Ou, S. & Jiang, N. LTR_retriever: a highly accurate and sensitive program for identification of LTR retrotransposons. Plant Physiol. 176, 1410–1422 (2018).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinformatics 9, 18 (2008).
Buels, R. et al. JBrowse: a dynamic web platform for genome visualization and analysis. Genome Biol. 17, 66 (2016).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 21, 5654–5666 (2003).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506 (2005).
Stanke, M., Schoffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinformatics 7, 62 (2006).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Hoang, N. V. et al. A survey of the complex transcriptome from the highly polyploid sugarcane genome using full-length isoform sequencing and de novo assembly from short read sequencing. BMC Genomics 18, 395 (2017).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Nattestad, M. & Schatz, M. C. Assemblytics: a web analytics tool for the detection of variants from an assembly. Bioinformatics 32, 3021–3023 (2016).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Andrews, S. FastQC, https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (Babraham Bioinformatics, 2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w 1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Plotree, D. & Plotgram, D. PHYLIP-phylogeny inference package (version 3.2). Cladistics 5, 6 (1989).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592 (2011).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Rosenberg, N. A. DISTRUCT: a program for the graphical display of population structure. Mol. Ecol. Res. 4, 137–138 (2004).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Acknowledgements
We thank L. McHale for reviewing and commenting on the section on disease resistance genes. This project was supported by a startup fund from Fujian Agriculture and Forestry University to R.M., the International Consortium for Sugarcane Biotechnology (project #35, R.M.), the US Department of Energy (DOE; DE-SC0010686 to R.M.), the European Bioinformatics Institute (BP2012OO2J17 to R.M.), the US National Science Foundation Plant Genome Research Program (grant IOS-1025976 to D.M.B.), the 863 program (2013AA102604 to J.Z.), the Natural Science Foundation of China (31201260 to J.Z.), the Program for New Century Excellent Talents in Fujian Province (J.Z.), and the São Paulo Research Foundation (FAPESP; grants 2008/52146-0, 2012/51062-3 and 2014/50921-8 to G.M.S. and 2008/52074-8 to M.-A.V.S.). This work was funded in part by the DOE Center for Advanced Bioenergy and Bioproducts Innovation (US DOE, Office of Science, Office of Biological and Environmental Research under Award Number DE-SC 18420 to M.H. and L.-Q.C.).
Author information
Authors and Affiliations
Contributions
R.M. conceived this genome project and coordinated research activities; R.M., C.N., J.Z. and Q.Y. designed the experiments; H.T. and Xingtan Z. developed novel algorithms to assemble autopolyploid genomes; T.J., P.H.M. and C.N. generated and maintained haploid plant materials and mapping populations; J.Z., Q.Z., X.H., Z.Li, Y.W., L.W. and J.X. collected and maintained plant materials; R.M. and Q.Y. generated BAC libraries; C.M.W., J.A., J.O.H., S.Chakrabarty, M.P. and A.S. isolated BAC DNA; S.Chen, L.H., W.Zhang, Yanhong M., Z.Y., F.D. and X.C. sequenced and processed the raw data; Xingtan Z., H.T., J.Z. and Q.Z. conducted comparative genomic analysis; H.T., Xingtan Z., J.Z., Q.Z., X.X., H.X., P.C., Z.K. and L.L. assembled and annotated the genome; J.B., R.M., Q.Z., Xingtan Z. J.Z. and Xingtan Z. constructed the linkage map; Q.Z., H.T., Y.S., S.Chen. and X.X. studied genome evolution; J.Z., X.H., X.Zhu, M.L., F.C. and G.Z. conducted the C4 photosynthesis analysis; J.Z., Xingtan Z., Q.Z., X.H., Y.S., L.H., Z.Li, Y.W., W.H. and Jishan L. analyzed gene families; R.M., Xingtan Z., F.Z., J.Z., Q.Z., X.H., X.M., Y.S., S.Chen, X.X., J.Y., L.H., Z.Li, H.X., D.Zhou, Y.W., W.H., L.W., Jishan L., Y.D., P.M., Z.Y., F.D., Z.Lin, Hai L., Hong L., H.Y., W.Zhong, P.L., Gang W., Guofeng W.,Y.Y., J.Song, J.Shi, J.H., Jingxian L., Q.S., Q.J., P.Z., Yaying M., Xunxiao Z., R.X., J.Liu, Y.Z., H.J., Q.M., R.Q., Z.Z., Y.D., N.P., M.M., D.Zerpa, J.K.N., L.W., L.Y., Y.X., G.D., H.F., J.J., J.F., Z.C., Z.L., M.H., M.W., T.M., X.Y. and L.G. manually checked the gene allele annotation; S.R.D., D.M.B., S.E.H. and L.-Q.C. analyzed sugar transporters; D.N. and M.S. analyzed P450 genes; Z.Z. and K.W. analyzed chromosome centromeres; C.Z. and D.S. tested whole-genome duplications; X.M., R.V.B., C.M.W., X.Y.J., J.W., J.H.T., M.C.M. and J.J.R. analyzed resequenced populations; M.C.S., D.H., M.-A.V.S. and G.M.S. contributed the SP80-3280 genome; and R.M., A.H.P., J.Z., H.T. and Xingtan Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Text and Figures
Supplementary Figures 1–21, Supplementary Tables 1, 2, 4–16 and 19–26, and Supplementary Note
Supplementary Table 3
Statistics of individual BAC pool assemblies
Supplementary Table 17
Comparison of C4-type genes between S. spontaneum and Sorghum
Supplementary Table 18
Gene IDs for 123 sugar transporters from nine subfamilies
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Zhang, X., Tang, H. et al. Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L.. Nat Genet 50, 1565–1573 (2018). https://doi.org/10.1038/s41588-018-0237-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0237-2
This article is cited by
-
Haplotype-resolved genome assembly provides insights into evolutionary history of the Actinidia arguta tetraploid
Molecular Horticulture (2024)
-
Two putative calcium-dependent protein kinases are involved in the regulation of sugarcane defense genes
Phytopathology Research (2024)
-
Genomic investigation of duplication, functional conservation, and divergence in the LRR-RLK Family of Saccharum
BMC Genomics (2024)
-
Chromosome-level genome provides insights into environmental adaptability and innate immunity in the common dolphin (delphinus delphis)
BMC Genomics (2024)
-
Phylogenomic profiles of whole-genome duplications in Poaceae and landscape of differential duplicate retention and losses among major Poaceae lineages
Nature Communications (2024)