Abstract
More than 15 years have passed since the identification, through linkage, of ‘first-wave’ susceptibility genes for common cancers (BRCA1, BRCA2, MLH1 and MSH2). These genes have strong frequency-penetrance profiles, such that the associated clinical utility probably remains relevant regardless of the context of ascertainment. ‘Second-wave’ genes, not tractable by linkage, were subsequently identified by mutation screening of candidate genes (PALB2, ATM, CHEK2, BRIP1, RAD51C and RAD51D). Their innately weaker frequency-penetrance profiles have rendered delineation of cancer associations, risks and variant pathogenicity challenging, thereby compromising their clinical application. Early germline exome-sequencing endeavors for common cancers did not yield the long-anticipated slew of ‘next-wave’ genes but instead implied a highly polygenic genomic architecture requiring much larger experiments to make any substantive inroads into gene discovery. As such, the ‘genetic economics’ of frequency penetrance clearly indicates that focused identification of carriers of first-wave-gene mutations is most impactful for cancer control. With screening, prevention and early detection at the forefront of the cancer management agenda, we propose that the time is nigh for the initiation of national population-testing programs to identify carriers of first-wave gene mutation. To fully deliver a precision prevention program, long-term, large-scale mutation studies that capture longitudinal clinical data and serial biosamples are required.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
12 December 2018
In the version of this article originally published, there was an error in the second-to-last sentence of the abstract. In this sentence, the final phrase “to identify carriers of first-wave gene mutation carriers” should have instead read “to identify carriers of first-wave gene mutation.” The error has been corrected in the HTML and PDF versions of the paper.
References
Cancer Research UK. Cancer Incidence Statistics (Cancer Research UK, London, 2017).
Burrell, R. A. & Swanton, C. Tumour heterogeneity and the evolution of polyclonal drug resistance. Mol. Oncol. 8, 1095–1111 (2014).
Prasad, V. Perspective: the precision-oncology illusion. Nature 537, S63 (2016).
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer: analyses of cohorts of twins from Sweden, Denmark, and Finland. N. Engl. J. Med. 343, 78–85 (2000).
Chanock, S. Charting a course toward precision cancer prevention. Cancer Currents Blog https://www.cancer.gov/news-events/cancer-currents-blog/2016/precision-prevention-chanock/ (2016).
Houlston, R. & Peto, J. Genetics and the common cancers. in Genetic Predisposition to Cancer (eds. Eeles, R., Ponder, B.A., Easton, D. & Horwich, A.) 208–226 (Chapman & Hall Medical, London, 1996).
Fletcher, O. & Houlston, R. S. Architecture of inherited susceptibility to common cancer. Nat. Rev. Cancer 10, 353–361 (2010).
Wooster, R. et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 378, 789–792 (1995).
Miki, Y. et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266, 66–71 (1994).
Papadopoulos, N. et al. Mutation of a mutL homolog in hereditary colon cancer. Science 263, 1625–1629 (1994).
Leach, F. S. et al. Mutations of a mutS homolog in hereditary nonpolyposis colorectal cancer. Cell 75, 1215–1225 (1993).
Hussussian, C. J. et al. Germline p16 mutations in familial melanoma. Nat. Genet. 8, 15–21 (1994).
Elston, R. C. & Cordell, H. J. Overview of model-free methods for linkage analysis. Adv. Genet. 42, 135–150 (2001).
Risch, N. The genetic epidemiology of cancer: interpreting family and twin studies and their implications for molecular genetic approaches. Cancer Epidemiol. Biomarkers Prev. 10, 733–741 (2001).
Al-Tassan, N. et al. Inherited variants of MYH associated with somatic G:C→T:A mutations in colorectal tumors. Nat. Genet. 30, 227–232 (2002).
Loveday, C. et al. Germline RAD51C mutations confer susceptibility to ovarian cancer. Nat. Genet. 44, 475–476 (2012).
Loveday, C. et al. Germline mutations in RAD51D confer susceptibility to ovarian cancer. Nat. Genet. 43, 879–882 (2011).
Rahman, N. et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat. Genet. 39, 165–167 (2007).
Renwick, A. et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nat. Genet. 38, 873–875 (2006).
Seal, S. et al. Truncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility alleles. Nat. Genet. 38, 1239–1241 (2006).
Meijers-Heijboer, H. et al. Low-penetrance susceptibility to breast cancer due to CHEK2*1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat. Genet. 31, 55–59 (2002).
Erkko, H. et al. A recurrent mutation in PALB2 in Finnish cancer families. Nature 446, 316–319 (2007).
Turnbull, C. & Rahman, N. Genetic predisposition to breast cancer: past, present, and future. Annu. Rev. Genomics Hum. Genet. 9, 321–345 (2008).
Cybulski, C. et al. Germline RECQL mutations are associated with breast cancer susceptibility. Nat. Genet. 47, 643–646 (2015).
Heikkinen, K. et al. RAD50 and NBS1 are breast cancer susceptibility genes associated with genomic instability. Carcinogenesis 27, 1593–1599 (2006).
Tommiska, J. et al. Evaluation of RAD50 in familial breast cancer predisposition. Int. J. Cancer 118, 2911–2916 (2006).
Slavin, T. P. et al. The contribution of pathogenic variants in breast cancer susceptibility genes to familial breast cancer risk. NPJ Breast Cancer 3, 22 (2017).
Buys, S. S. et al. A study of over 35,000 women with breast cancer tested with a 25-gene panel of hereditary cancer genes. Cancer 123, 1721–1730 (2017).
Easton, D. F. et al. No evidence that protein truncating variants in BRIP1 are associated with breast cancer risk: implications for gene panel testing. J. Med. Genet. 53, 298–309 (2016).
Sopik, V. & Foulkes, W. D. Risky business: getting a grip on BRIP. J. Med. Genet. 53, 296–297 (2016).
Rafnar, T. et al. Mutations in BRIP1 confer high risk of ovarian cancer. Nat. Genet. 43, 1104–1107 (2011).
Pelttari, L. M. et al. RAD51C is a susceptibility gene for ovarian cancer. Hum. Mol. Genet. 20, 3278–3288 (2011).
Couch, F. J. et al. Associations between cancer predisposition testing panel genes and breast cancer. JAMA Oncol. 3, 1190–1196 (2017).
Osorio, A. et al. Predominance of pathogenic missense variants in the RAD51C gene occurring in breast and ovarian cancer families. Hum. Mol. Genet. 21, 2889–2898 (2012).
Meindl, A. et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat. Genet. 42, 410–414 (2010).
Pharoah, P. D. P. et al. PPM1D mosaic truncating variants in ovarian cancer cases may be treatment-related somatic mutations. J. Natl. Cancer Inst. 108, djv347 (2016).
Swisher, E. M. et al. Somatic mosaic mutations in PPM1D and TP53 in the blood of women with ovarian carcinoma. JAMA Oncol. 2, 370–372 (2016).
Zajkowicz, A. et al. Truncating mutations of PPM1D are found in blood DNA samples of lung cancer patients. Br. J. Cancer 112, 1114–1120 (2015).
Ruark, E. et al. Mosaic PPM1D mutations are associated with predisposition to breast and ovarian cancer. Nature 493, 406–410 (2013).
Southey, M. C. et al. PALB2, CHEK2 and ATM rare variants and cancer risk: data from COGS. J. Med. Genet. 53, 800–811 (2016).
Antoniou, A. C. et al. Breast-cancer risk in families with mutations in PALB2. N. Engl. J. Med. 371, 497–506 (2014).
Concannon, P. ATM heterozygosity and cancer risk. Nat. Genet. 32, 89–90 (2002).
Gatti, R. A., Tward, A. & Concannon, P. Cancer risk in ATM heterozygotes: a model of phenotypic and mechanistic differences between missense and truncating mutations. Mol. Genet. Metab. 68, 419–423 (1999).
Swift, M., Reitnauer, P. J., Morrell, D. & Chase, C. L. Breast and other cancers in families with ataxia-telangiectasia. N. Engl. J. Med. 316, 1289–1294 (1987).
Schmidt, M. K. et al. Age- and tumor subtype-specific breast cancer risk estimates for CHEK2*1100delC carriers. J. Clin. Oncol. 34, 2750–2760 (2016).
Hale, V., Weischer, M. & Park, J. Y. CHEK2*1100delC mutation and risk of prostate cancer. Prostate Cancer 2014, 294575 (2014).
Han, F. F., Guo, C. L. & Liu, L. H. The effect of CHEK2 variant I157T on cancer susceptibility: evidence from a meta-analysis. DNA Cell Biol. 32, 329–335 (2013).
Liu, C., Wang, Q. S. & Wang, Y. J. The CHEK2 I157T variant and colorectal cancer susceptibility: a systematic review and meta-analysis. Asian Pac. J. Cancer Prev. 13, 2051–2055 (2012).
Weischer, M., Bojesen, S. E., Ellervik, C., Tybjaerg-Hansen, A. & Nordestgaard, B. G. CHEK2*1100delC genotyping for clinical assessment of breast cancer risk: meta-analyses of 26,000 patient cases and 27,000 controls. J. Clin. Oncol. 26, 542–548 (2008).
Schutte, M. et al. Variants in CHEK2 other than 1100delC do not make a major contribution to breast cancer susceptibility. Am. J. Hum. Genet. 72, 1023–1028 (2003).
Dong, C. et al. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum. Mol. Genet. 24, 2125–2137 (2015).
Findlay, G.M. et al. Accurate functional classification of thousands of BRCA1 variants with saturation genome editing. Preprint at https://www.biorxiv.org/content/early/2018/04/05/294520 (2018).
Starita, L. M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Easton, D. F. et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 372, 2243–2257 (2015).
Osher, D. J. et al. Mutation analysis of RAD51D in non-BRCA1/2 ovarian and breast cancer families. Br. J. Cancer 106, 1460–1463 (2012).
Tung, N. et al. Counselling framework for moderate-penetrance cancer-susceptibility mutations. Nat. Rev. Clin. Oncol. 13, 581–588 (2016).
DeRycke, M. S. et al. Targeted sequencing of 36 known or putative colorectal cancer susceptibility genes. Mol. Genet. Genomic Med. 5, 553–569 (2017).
Hampel, H. et al. Screening for Lynch syndrome (hereditary nonpolyposis colorectal cancer) among endometrial cancer patients. Cancer Res. 66, 7810–7817 (2006).
Hampel, H. et al. Cancer risk in hereditary nonpolyposis colorectal cancer syndrome: later age of onset. Gastroenterology 129, 415–421 (2005).
Antoniou, A. C., Pharoah, P. P., Smith, P. & Easton, D. F. The BOADICEA model of genetic susceptibility to breast and ovarian cancer. Br. J. Cancer 91, 1580–1590 (2004).
Antoniou, A. et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: a combined analysis of 22 studies. Am. J. Hum. Genet. 72, 1117–1130 (2003).
Antoniou, A. C. et al. A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br. J. Cancer 86, 76–83 (2002).
Rich, T., Lotito, M., Kidd, J., Saam, J. & Lancaster, J. Abstract PD7–03: characterization of Li-Fraumeni syndrome diagnosed using a 25-gene hereditary cancer panel. Cancer Res. 76, PD7–03 (2016).
Susswein, L. R. et al. Pathogenic and likely pathogenic variant prevalence among the first 10,000 patients referred for next-generation cancer panel testing. Genet. Med. 18, 823–832 (2016).
de Andrade, K. C. et al. Higher-than-expected population prevalence of potentially pathogenic germline TP53 variants in individuals unselected for cancer history. Hum. Mutat. 38, 1723–1730 (2017).
Kim, J., Field, A., Schultz, K. A. P., Hill, D. A. & Stewart, D. R. The prevalence of DICER1 pathogenic variation in population databases. Int. J. Cancer 141, 2030–2036 (2017).
Loveday, C. et al. p.Val804Met, the most frequent pathogenic mutation in RET, confers a very low lifetime risk of medullary thyroid cancer. J. Clin. Endocrinol. Metab. https://doi.org/10.1210/jc.2017-02529 (2018).
Narod, S. A. The tip of the iceberg: a countercurrents series. Curr. Oncol. 19, 129–130 (2012).
Easton, D. F. & Eeles, R. A. Genome-wide association studies in cancer. Hum. Mol. Genet. 17, R109–R115 (2008).
Easton, D. F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Hunter, D. J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat. Genet. 39, 870–874 (2007).
Broderick, P. et al. A genome-wide association study shows that common alleles of SMAD7 influence colorectal cancer risk. Nat. Genet. 39, 1315–1317 (2007).
Tomlinson, I. et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat. Genet. 39, 984–988 (2007).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Michailidou, K. et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat. Genet. 45, 353–361 (2013).
Turnbull, C. et al. Genome-wide association study identifies five new breast cancer susceptibility loci. Nat. Genet. 42, 504–507 (2010).
Michailidou, K. et al. Association analysis identifies 65 new breast cancer risk loci. Nature 551, 92–94 (2017).
Amos, C. I. et al. The OncoArray Consortium: a network for understanding the genetic architecture of common cancers. Cancer Epidemiol. Biomarkers Prev. 26, 126–135 (2017).
Al Olama, A. A. et al. A meta-analysis of 87,040 individuals identifies 23 new susceptibility loci for prostate cancer. Nat. Genet. 46, 1103–1109 (2014).
Sakoda, L. C., Jorgenson, E. & Witte, J. S. Turning of COGS moves forward findings for hormonally mediated cancers. Nat. Genet. 45, 345–348 (2013).
Sud, A., Kinnersley, B. & Houlston, R. S. Genome-wide association studies of cancer: current insights and future perspectives. Nat. Rev. Cancer 17, 692–704 (2017).
Guo, M. H. et al. Determinants of power in gene-based burden testing for monogenic disorders. Am. J. Hum. Genet. 99, 527–539 (2016).
Chubb, D. et al. Rare disruptive mutations and their contribution to the heritable risk of colorectal cancer. Nat. Commun. 7, 11883 (2016).
Manchanda, R. et al. Current detection rates and time-to-detection of all identifiable BRCA carriers in the Greater London population. J. Med. Genet. 55, 538–545 (2018).
Manchanda, R. et al. Cost-effectiveness of population-based BRCA1, BRCA2, RAD51C, RAD51D, BRIP1, PALB2 mutation testing in unselected general population women. J. Natl. Cancer Inst. 110, 714–725 (2018).
Manchanda, R. et al. Cost-effectiveness of population based BRCA testing with varying Ashkenazi Jewish ancestry. Am. J. Obstet. Gynecol. 217, 578.e1–578.e12 (2017).
Manchanda, R. et al. Cost-effectiveness of population screening for BRCA mutations in Ashkenazi jewish women compared with family history-based testing. J. Natl. Cancer Inst. 107, 380 (2014).
Manchanda, R. et al. Population testing for cancer predisposing BRCA1/BRCA2 mutations in the Ashkenazi-Jewish community: a randomized controlled trial. J. Natl. Cancer Inst. 107, 379 (2014).
Lieberman, S. et al. Population screening for BRCA1/BRCA2 founder mutations in Ashkenazi Jews: proactive recruitment compared with self-referral. Genet. Med. 19, 754–762 (2017).
Levy-Lahad, E., Lahad, A. & King, M.-C. Precision medicine meets public health: population screening for BRCA1 and BRCA2. J. Natl. Cancer Inst. 107, 420 (2014).
Pashayan, N. et al. Polygenic susceptibility to prostate and breast cancer: implications for personalised screening. Br. J. Cancer 104, 1656–1663 (2011).
Garcia-Closas, M., Gunsoy, N. B. & Chatterjee, N. Combined associations of genetic and environmental risk factors: implications for prevention of breast cancer. J. Natl. Cancer Inst. 106, dju305 (2014).
Mavaddat, N. et al. Prediction of breast cancer risk based on profiling with common genetic variants. J. Natl. Cancer Inst. 107, djv036 (2015).
Frampton, M. J. et al. Implications of polygenic risk for personalised colorectal cancer screening. Ann. Oncol. 27, 429–434 (2016).
Litchfield, K. et al. Polygenic susceptibility to testicular cancer: implications for personalised health care. Br. J. Cancer 114, e22 (2016).
Kong, S. W. et al. Summarizing polygenic risks for complex diseases in a clinical whole-genome report. Genet. Med. 17, 536–544 (2015).
Li, H. et al. Breast cancer risk prediction using a polygenic risk score in the familial setting: a prospective study from the Breast Cancer Family Registry and kConFab. Genet. Med. 19, 30–35 (2017).
Gray, E. et al. Evaluation of a stratified national breast screening program in the United Kingdom: an early model-based cost-effectiveness analysis. Value Health 20, 1100–1109 (2017).
Evans, D. G. et al. The impact of a panel of 18 SNPs on breast cancer risk in women attending a UK familial screening clinic: a case-control study. J. Med. Genet. 54, 111–113 (2017).
Jervis, S. et al. A risk prediction algorithm for ovarian cancer incorporating BRCA1, BRCA2, common alleles and other familial effects. J. Med. Genet. 52, 465–475 (2015).
Lee, A. J. et al. Incorporating truncating variants in PALB2, CHEK2, and ATM into the BOADICEA breast cancer risk model. Genet. Med. 18, 1190–1198 (2016).
Jones, M. R., Kamara, D., Karlan, B. Y., Pharoah, P. D. P. & Gayther, S. A. Genetic epidemiology of ovarian cancer and prospects for polygenic risk prediction. Gynecol. Oncol. 147, 705–713 (2017).
Møller, P. et al. Cancer incidence and survival in Lynch syndrome patients receiving colonoscopic and gynaecological surveillance: first report from the prospective Lynch syndrome database. Gut 66, 464–472 (2017).
Milne, R. L. & Antoniou, A. C. Modifiers of breast and ovarian cancer risks for BRCA1 and BRCA2 mutation carriers. Endocr. Relat. Cancer 23, T69–T84 (2016).
Cancer Research UK. UK Cancer Incidence Statistics (Cancer Research UK, London, 2018).
Wilson, J. M. G. & Jungner, G. Principles and Practices of Screening for Disease. Report no. Public Health Papers 34 (World Health Organization, Geneva, 1968).
Yurgelun, M. B., Chenevix-Trench, G. & Lippman, S. M. Translating germline cancer risk into precision prevention. Cell 168, 566–570 (2017).
Spira, A. et al. Leveraging premalignant biology for immune-based cancer prevention. Proc. Natl. Acad. Sci. USA 113, 10750–10758 (2016).
Nolan, E. et al. RANK ligand as a potential target for breast cancer prevention in BRCA1-mutation carriers. Nat. Med. 22, 933–939 (2016).
Kloor, M. et al. Vaccination of MSI-H colorectal cancer patients with frameshift peptide antigens: A phase I/IIa clinical trial. J. Clin. Oncol. 33, 3020 (2015).
Milne, R. L. & Antoniou, A. C. Genetic modifiers of cancer risk for BRCA1 and BRCA2 mutation carriers. Ann. Oncol. 22 (Suppl. 1), i11–i17 (2011).
Spurdle, A. B. et al. BRCA1 R1699Q variant displaying ambiguous functional abrogation confers intermediate breast and ovarian cancer risk. J. Med. Genet. 49, 525–532 (2012).
Moghadasi, S. et al. The BRCA1 c. 5096G>A p.Arg1699Gln (R1699Q) intermediate risk variant: breast and ovarian cancer risk estimation and recommendations for clinical management from the ENIGMA consortium. J. Med. Genet. 55, 15–20 (2018).
Rebbeck, T. R. et al. Association of type and location of BRCA1 and BRCA2 mutations with risk of breast and ovarian cancer. JAMA 313, 1347–1361 (2015).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Cheung, R. et al. Large-scale screening of rare genetic variants in humans reveals frequent splicing disruptions. Preprint at https://www.biorxiv.org/content/early/2017/10/08/199927 (2017).
Acknowledgements
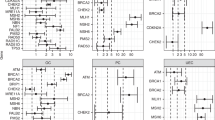
We thank our many colleagues for decades of interesting discussions around these themes, in particular W. Foulkes, M. Tischkowitz, H. Hanson, A. Taylor, K. Snape and A. Kulkani for their invaluable and wise thoughts. We also thank D. Easton (University of Cambridge, UK) and P. Devilee (Leiden University Medical Center, the Netherlands) for providing the data used to generate Fig. 1a. R.S.H. is supported by Cancer Research UK (C1298/A8362 Bobby Moore Fund for Cancer Research UK). C.T. is supported by the Movember Foundation.
Author information
Authors and Affiliations
Contributions
C.T., A.S. and R.S.H. researched, reviewed, drafted and edited the manuscript. A.S. and C.T. generated the images.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Turnbull, C., Sud, A. & Houlston, R.S. Cancer genetics, precision prevention and a call to action. Nat Genet 50, 1212–1218 (2018). https://doi.org/10.1038/s41588-018-0202-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0202-0
This article is cited by
-
The complementary roles of genome-wide approaches in identifying genes linked to an inherited risk of colorectal cancer
Hereditary Cancer in Clinical Practice (2023)
-
Repurposing of neprilysin inhibitor ‘sacubitrilat’ as an anti-cancer drug by modulating epigenetic and apoptotic regulators
Scientific Reports (2023)
-
Mendelian inheritance revisited: dominance and recessiveness in medical genetics
Nature Reviews Genetics (2023)
-
Identification of novel leads as potent inhibitors of HDAC3 using ligand-based pharmacophore modeling and MD simulation
Scientific Reports (2022)
-
Single-cell profiling of tumour evolution in multiple myeloma — opportunities for precision medicine
Nature Reviews Clinical Oncology (2022)