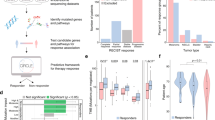
Abstract
Tumor mutational burden correlates with response to immune checkpoint blockade in multiple solid tumors, although in microsatellite-stable tumors this association is of uncertain clinical utility. Here we uniformly analyzed whole-exome sequencing (WES) of 249 tumors and matched normal tissue from patients with clinically annotated outcomes to immune checkpoint therapy, including radiographic response, across multiple cancer types to examine additional tumor genomic features that contribute to selective response. Our analyses identified genomic correlates of response beyond mutational burden, including somatic events in individual driver genes, certain global mutational signatures, and specific HLA-restricted neoantigens. However, these features were often interrelated, highlighting the complexity of identifying genetic driver events that generate an immunoresponsive tumor environment. This study lays a path forward in analyzing large clinical cohorts in an integrated and multifaceted manner to enhance the ability to discover clinically meaningful predictive features of response to immune checkpoint blockade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Topalian, S. L., Taube, J. M., Anders, R. A. & Pardoll, D. M. Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat. Rev. Cancer 16, 275–287 (2016).
Brahmer, J. et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N. Engl. J. Med. 373, 123–135 (2015).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Sharma, P. Immune checkpoint therapy and the search for predictive biomarkers. Cancer J. 22, 68–72 (2016).
Carbognin, L. et al. Differential activity of nivolumab, pembrolizumab and MPDL3280A according to the tumor expression of programmed death-ligand-1 (PD-L1): sensitivity analysis of trials in melanoma, lung and genitourinary cancers. PLoS One 10, e0130142 (2015).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Roh, W. et al. Integrated molecular analysis of tumor biopsies on sequential CTLA-4 and PD-1 blockade reveals markers of response and resistance. Sci. Trans. Med. 9, eaah3560 (2017).
Colli, L. M. et al. Burden of nonsynonymous mutations among TCGA cancers and candidate immune checkpoint inhibitor responses. Cancer Res. 76, 3767–3772 (2016).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Riaz, N. et al. Recurrent SERPINB3 and SERPINB4 mutations in patients who respond to anti-CTLA4 immunotherapy. Nat. Genet. 48, 1327–1329 (2016).
Johnson, D. B. et al. Impact of NRAS mutations for patients with advanced melanoma treated with immune therapies. Cancer Immunol. Res. 3, 288–295 (2015).
Gao, J. et al. Loss of IFN-γ pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167, 397–404 (2016).
Kato, S. et al. Hyper-progressors after immunotherapy: analysis of genomic alterations associated with accelerated growth rate. Clin. Cancer Res. 23, 4242–4250 (2017).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Davoli, T., Uno, H., Wooten, E. C. & Elledge, S. J. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355, eaaf8399 (2017).
Sucker, A. et al. Acquired IFNγ resistance impairs anti-tumor immunity and gives rise to T-cell-resistant melanoma lesions. Nat. Commun. 8, 15440 (2017).
Van Allen, E. M. et al. Long-term benefit of PD-L1 blockade in lung cancer associated with JAK3 activation. Cancer Immunol. Res. 3, 855–863 (2015).
George, S. et al. Loss of PTEN is associated with resistance to anti-PD-1 checkpoint blockade therapy in metastatic uterine leiomyosarcoma. Immunity 46, 197–204 (2017).
Mouw, K. W. et al. Genomic evolution after chemoradiotherapy in anal squamous cell carcinoma. Clin. Cancer Res. 23, 3214–3222 (2017).
Garofalo, A. et al. The impact of tumor profiling approaches and genomic data strategies for cancer precision medicine. Genome Med. 8, 79 (2016).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Wolchok, J. D. et al. Guidelines for the evaluation of immune therapy activity in solid tumors: immune-related response criteria. Clin. Cancer Res. 15, 7412–7420 (2009).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Govindan, R. et al. Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150, 1121–1134 (2012).
Rizvi, H. et al. Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J. Clin. Oncol. 36, 633–641 (2018).
de Bruin, E. C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Henderson, S., Chakravarthy, A., Su, X., Boshoff, C. & Fenton, T. R. APOBEC-mediated cytosine deamination links PIK3CA helical domain mutations to human papillomavirus-driven tumor development. Cell Rep. 7, 1833–1841 (2014).
Mullane, S. A. et al. Correlation of APOBEC mRNA expression with overall survival and PD-L1 expression in urothelial carcinoma. Sci. Rep. 6, 27702 (2016).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Peng, W. et al. Loss of PTEN promotes resistance to T cell–mediated immunotherapy. Cancer Discov. 6, 202–216 (2016).
Pan, D. et al. A major chromatin regulator determines resistance of tumor cells to T cell–mediated killing. Science 359, 770–775 (2018).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Hodges, C., Kirkland, J. G. & Crabtree, G. R. The many roles of BAF (mSWI/SNF) and PBAF complexes in cancer. Cold Spring Harb. Perspect. Med. 6, a026930 (2016).
Gettinger, S. et al. Nivolumab monotherapy for first-line treatment of advanced non-small-cell lung cancer. J. Clin. Oncol. 34, 2980–2987 (2016).
Van Allen, E. M. et al. Whole-exome sequencing and clinical interpretation of formalin-fixed, paraffin-embedded tumor samples to guide precision cancer medicine. Nat. Med. 20, 682–688 (2014).
Cibulskis, K. et al. ContEst: estimating cross-contamination of human samples in next-generation sequencing data. Bioinformatics 27, 2601–2602 (2011).
Taylor-Weiner, A. et al. DeTiN: overcoming tumor-in-normal contamination. Nat. Methods. 15, 531–534 (2018).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res. 41, e67 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor–normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Gao, J. et al. 3D clusters of somatic mutations in cancer reveal numerous rare mutations as functional targets. Genome Med. 9, 4 (2017).
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Brastianos, P. K. et al. Genomic characterization of brain metastases reveals branched evolution and potential therapeutic targets. Cancer Discov. 5, 1164–1177 (2015).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Hoof, I. et al. NetMHCpan, a method for MHC class I binding prediction beyond humans. Immunogenetics 61, 1–13 (2009).
Nielsen, M. & Andreatta, M. NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Med. 8, 33 (2016).
Nielsen, M. et al. NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and -B locus protein of known sequence. PLoS One 2, e796 (2007).
Acknowledgements
This work was supported by BroadIgnite (E.M.V.A.), BroadNext10 (E.M.V.A., D.M.), NIH R01CA227388 (E.M.V.A.) and NIH K08CA188615 (E.M.V.A.). D.M. was supported by the Howard Hughes Medical Institute Medical Research Fellows Program. This research was also supported by the Center for Immuno-Oncology at the Dana-Farber Cancer Institute and a Stand Up To Cancer–American Cancer Society Lung Cancer Dream Team Translational Research Grant (SU2C-AACR-DT17-15). Stand Up To Cancer (SU2C) is a program of the Entertainment Industry Foundation. Research grants are administered by the American Association for Cancer Research, the scientific partner of SU2C.
Author information
Authors and Affiliations
Contributions
D.M., C.A.M., N.I.V., A.T.-W., D.L., D.A., P.P., G.G., and D.K. performed the analyses. B.S., M.M., M.M.A., N.G.C., G.J.H., R.H. and S.M.W. provided clinical annotations. L.M.S., S.S., and S.J.R. contributed to immunohistochemical profiling. K.-K.W., J.A.E., M.M.A., D.A.B., R.I.H., D.S., F.S.H., T.K.C., J.B., P.A.J., R.H., A.T., P.H., and E.M.V.A. contributed to sample acquisition. E.M.V.A. supervised the study. D.M., C.A.M., N.I.V., D.L., and E.M.V.A. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
A.T., D.L., D.M., M.M., N.I.V., C.A.M., D.A., D.K., S.M.W., L.M.S., A.T.-W., P.P., K.-K.W., S.J.R., J.B., P.A.J., N.G.C., R.H., and M.M.A. declare no conflicts of interest. T.K.C. has advisory roles with AstraZeneca, Bayer, Bristol-Myers Squibb, Cerulean, Elsa, Foundation Medicine, Genentech, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, Prometheus Labs, Roche, and Elsai. T.K.C. receives research funding from AstraZeneca, Bristol-Myers Squibb, Exelixis, Genentech, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, Roche, Tracon, and Eisai. G.J.H. receives institutional support from Bristol-Myers Squibb and EMD Serono. B.S. is on the advisory board or has received honoraria from Novartis, Roche, Bristol-Myers Squibb, and MSD Sharp & Dohme, research funding from Bristol-Myers Squibb and MSD Sharp & Dohme, and travel support from Novartis, Roche, Bristol-Myers Squibb, and Amgen. R.I.H. has advisory roles with Bristol-Myers Squibb, Pfizer, Merck, AstraZeneca, Genentech, and Celgene. R.I.H. receives research funding from Bristol-Myers Squibb, Merck, Genentech, and Pfizer. S.S. is a consultant for AstraZeneca and Merck and receives research funding from AstraZeneca, Bristol-Myers Squibb, Exelixis, and Roche. G.G. has an advisory role with MD Anderson and receives research funding from IBM and Bayer. G.G. is listed as an inventor on patent applications regarding MuTect, ABSOLUTE, and Polysolver. D.A.B. is a consultant for N of One. D.S. receives consulting fees from Amgen, GlaxoSmithKline, BMS, Novartis, Roche, Amgen, Merck, AstraZeneca, Merck-Serono, and Pfizer. P.H. and J.A.E. are employees of Novartis. F.S.H. is a consultant to Bristol-Myers Squibb, Merck, Novartis, EMD Serono, Sanofi, and Genentech and receives institutional research support from Bristol-Myers Squibb. E.M.V.A. holds consulting roles with Tango Therapeutics, Invitae, and Genome Medical and receives research support from Bristol-Myers Squibb and Novartis.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Tumor purity, mutational burden, and response to immune checkpoint therapy.
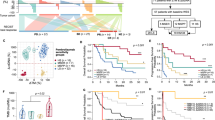
a, Called mutations by estimated tumor purity (ABSOLUTE) in 297 tumors with adequate sequencing quality (i.e., not excluded owing to sample contamination, low sequencing coverage, or other quality issues). Fifteen samples with high purity and outlying high mutational burdens are not shown. Purity <10% was associated with decreased ability to call mutations in most samples. The two samples with purity <10% and >500 called mutations/exome had unusually high sequencing coverage. However, inference of clonal versus subclonal mutational architecture was not possible by ABSOLUTE given low confidence in calling somatic CNAs, and a high rate of false negative mutation calls may have occurred, so these samples were excluded from analysis. Box plots show the median, first and third quartiles, whiskers extend to 1.5 times the interquartile range, and outlying points are plotted individually. b, Bar plot showing the frequency of estimated tumor purities, colored by patient response status. Tumor purity did not strongly predict clinical benefit from immune checkpoint therapies in 249 tumors included in the final analysis. c, Scatterplot showing subclonal mutational burden by estimated tumor purity in 249 included samples. Even after excluding tumors with estimated purity <0.1, subclonal mutations were likely undercalled in tumors with purity <0.2.
Supplementary Figure 2 Clinical response categories and mutational burden across four cancer types.
Box plots showing mutational burden on the y axis (all mutations, all nonsynonymous mutations, and all clonal nonsynonymous mutations) are displayed in an array, organized by tumor type (in columns) and response categorization used (in rows). The range of mutational load varied greatly by cancer type. Additionally, the choice of response metric influenced both the number of patients designated as OR or NR per cancer type (Supplementary Table 3) and the strength of the relationship between mutational burden and response. Two-sided Mann–Whitney U test: *P < 0.05, **P < 0.005. Box plots show the median, first and third quartiles, whiskers extend to 1.5 times the interquartile range, and outlying points are plotted individually.
Supplementary Figure 3 Differences in mutational load and clonality by response category and treatment.
a, Mutational burden among response groups after stratifying by treatment category. There was a significant difference in mutational burden across all three response categories in patients treated with PD-1/PD-L1 inhibitors. In anti-CTLA-4-treated patients, mutational burden differed only between CR/PR patients and PD patients. b, All mutations (synonymous and nonsynonymous), nonsynonymous, and clonal nonsynonymous mutational burden among the response groups after stratifying SD and PD by duration of overall survival (OS). Across all mutation categories, mutational burden was significantly higher in CR/PR patients than in those with SD and OS <1 year and those with PD and OS <2 years. However, there was no statistically significant difference in mutation load between CR/PR patients and those with SD and OS >1 year nor between those with PD and OS >2 years (P > 0.05). Two-sided Mann–Whitney U test: *P < 0.05, **P < 0.005. Box plots show the median, first and third quartiles, whiskers extend to 1.5 times the interquartile range, and outlying points are plotted individually.
Supplementary Figure 4 Receiver operating characteristic analysis of mutational load as a univariate whole-exome-based response predictor.
The red line represents the accuracy of prediction of CR/PR versus PD using nonsynonymous mutational burden as a continuous variable alone (area under the ROC curve (AUC) = 0.66). The ROC curve was generated using data from 193 (70 CR/PR, 123 PD) patient samples.
Supplementary Figure 5 Enrichment of nonsynonymous mutations in all mutated genes in the cohort by response category.
The y axis shows –log10 (P) for enrichment of mutations in OR (above the dashed line) or in NR (below the dashed line) by two-sided Fisher’s exact test. The x axis shows the prevalence of each mutation across the cohort. Colors denote the significance of enrichment, with dark gray points representing genes that are significantly mutated across the entire cohort (MutSig2CV q < 0.1), blue representing genes that are mutated significantly more frequently in either OR or NR but are not mutated at significant levels across the entire cohort (P < 0.05), and red representing genes that were significantly enriched in mutations in OR or NR and mutated frequently across the cohort by MutSig2CV. All other genes are shown in light gray. Circle size is proportional to the number of patients with a mutation in a given gene. Three different methods of stratifying patients into OR versus NR are shown.
Supplementary Figure 6 PIK3CA driver mutations in the APOBEC mutational signature spectrum.
Mutational context of mutations in PIK3CA observed in this cohort overlaid on the APOBEC-associated mutational spectrum. Mutations in dark red text are known hotspots in PIK3CA. Hotspots occurring because of C>T mutations have been previously associated with APOBEC-associated mutational processes in HNSCC (Henderson et al. Cell Rep. 2014).
Supplementary Figure 7 Smoking status, tumor mutational burden, and clinical benefit from immune checkpoint therapy.
a, Proportion of patients with the smoking dominant mutational signature (S4, brown), APOBEC (S2/S13, teal), aging (S1, light tan), or unknown (S5, dark tan) by response group. b, Differences in mutational burden by response group in patients with a self-reported current/former smoking history (>5 pack-years). All P values reported are from two-sided Mann–Whitney U tests. CR/PR versus PD, P = 0.0002; CR/PR versus SD, P = 0.003; SD versus PD, P = 0.085. **P < 0.005 (two-sided Mann–Whitney U test). Box plots show the median, first and third quartiles, whiskers extend to 1.5 times the interquartile range, and outlying points are plotted individually.
Supplementary Figure 8 Mutational burden, mutational signatures, tumor purity, histological subtype, and response to immune checkpoint therapy in melanoma.
a, Stacked plots showing tumor mutational burden (histogram, top, blue and purple), estimated tumor purity by ABSOLUTE (tile plot, top, purple), presence of BRAF p.V600E mutation (tile plot, top, red), response to immune checkpoint therapy (tile plot, middle), mutational signatures (filled histogram, bottom), and tumor subtype (bottom). UV-dominant (S7), alkylating-dominant (S11), and non-UV-/non-alkylating-dominant (S1, S5, S17, or S6) tumors are shown separately. b, Dominant (most represented) mutational signature by tumor subtype. Cutaneous melanomas were largely UV associated, while acral lentiginous melanomas and non-cutaneous melanomas generally had few UV-associated mutations, although exceptions occurred.
Supplementary Figure 9 Comparison of multiple definitions of genomic segment-level deletion or amplification and association with purity.
The proportion of the exome affected by amplifications or deletions is shown on the y axis, with estimated tumor purity by ABSOLUTE on the x axis. a, Defining copy number alterations as segments with |log2 (copy ratio)| > 0.5 leads to results that are highly dependent on tumor purity. b, Correcting for tumor purity and ploidy while using a similar threshold for calling a segment amplified or deleted yields copy number calls that are more sensitive and less dependent on tumor purity. c, A more stringent definition of copy number alterations designed to detect only major amplifications and homozygous deletions to specifically identify events expected to lead to major changes in gene expression incorporates both tumor purity and ploidy as well as a measurement of segment focality, as further described in the Methods.
Supplementary Figure 10 Copy number alterations affecting interferon signaling by drug class.
a, Proportion of samples harboring a CNA disrupting interferon-γ signaling by drug class using a definition of clinical benefit from Roh et al. (2017). For the anti-CTLA-4 condition, n = 145; two-sided Fisher’s exact test P = 0.021. For the anti-PD-1/PD-L1 condition, n = 94; two-sided Fisher’s exact test P = 0.038. b, Proportion of samples harboring a CNA disrupting interferon signaling by drug class using a definition of clinical benefit from Van Allen et al. (2015). For the anti-CTLA-4 condition, n = 133; two-sided Fisher’s exact test P = 0.008. For the anti-PD-1/PD-L1 condition, n = 92; two-sided Fisher’s exact test P = 0.073. Two-sided Fisher’s exact test: *P < 0.05. Error bars represent standard error above and below the group proportion.
Supplementary Figure 11 Copy number alterations affecting interferon signaling by cancer type.
All P values listed are from two-sided Fisher’s exact tests. a, Proportion of samples harboring a CNA disrupting interferon-γ signaling by cancer type using a definition of clinical benefit from Roh et al. (2017). All, n = 249 and P = 0.0002; bladder, n = 27 and P = 0.33; HNSCC, n = 12 and P = 1; lung, n = 57 and P = 0.26; melanoma, n = 151 and P = 0.02. b, Proportion of samples harboring a CNA disrupting interferon signaling by cancer type using a definition of clinical benefit from Van Allen et al. (2015). All, n = 234 and P = 0.002; bladder, n = 26 and P = 0.31; HNSCC, n = 12 and P = 1; lung, n = 56 and P = 0.46; melanoma, n = 138 and P = 0.004. Two-sided Fisher’s exact test: *P < 0.05, **P < 0.005. Error bars represent standard error above and below the group proportion.
Supplementary Figure 12 Putative biallelic PTEN loss in progressing lesions from two patients with CB to anti-CTLA-4 therapy.
a, Copy number plot (top) and allelic copy number plot (bottom) showing a PTEN nonsense mutation (p.V119fs) in a progressing lesion from a patient with overall PR to anti-CTLA-4 monotherapy. The estimated tumor purity in this sample was 0.14, which was too low to accurately evaluate the presence or absence of a heterozygous deletion at this locus. Thus, this patient has at least monoallelic and potentially biallelic PTEN loss. b, Copy number plot (top) and allelic copy number plot (bottom) with the arrow indicating the PTEN locus, which occurs in an area of copy-neutral loss of heterozygosity on chromosome 10. This sample was derived from a progressing lesion from a patient with a long duration of stable disease on anti-CTLA-4 therapy. The tumor harbored a PTEN splice-site mutation, a PTEN p.P89L missense mutation, and loss of heterozygosity over the PTEN locus, suggesting biallelic PTEN loss.
Supplementary Figure 13 Truncating mutations and biallelic deletions of genes encoding SWI/SNF subunits.
All samples with at least one truncating mutation in PBRM1 or ARID2 are shown. Colors represent mutational classification (nonsense, missense, etc.). The bottom tile plot indicates patient response to immune checkpoint therapy. Cells containing asterisks indicate patients with more than one truncating mutation in a given gene. Phasing of these mutations could not be determined. Mutations occurring in the context of heterozygous deletion (light blue) likely lead to complete loss of protein function.
Supplementary Figure 14 Biallelic loss of genes involved in JAK–STAT signaling.
a, Samples with biallelic loss (truncating mutation with deletion of the wild-type allele) of at least one gene in the JAK–STAT pathway are shown. Color indicates mutation class. The bottom tile plot indicates clinical response to immune checkpoint therapy. b, Samples with mutations or copy number alterations leading to biallelic loss in antigen-presentation pathway genes are shown. c, Copy number plot demonstrating biallelic loss of β2-microglobulin via a homozygous deletion in a patient with melanoma.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Tables 3, 6 and 8
Supplementary Table 1
Sequencing metrics and inclusion/exclusion criteria for whole-exome sequencing from 314 clinically annotated patient tumors
Supplementary Table 2
Clinical outcomes and clinical covariates for 249 patients
Supplementary Table 4
Oncogenes and tumor suppressors assessed for driver mutations
Supplementary Table 5
All mutations (somatic single-nucleotide variants and small insertions and deletions) across 249 tumors included in analysis
Supplementary Table 7
Mutational signature activity across all tumors
Supplementary Table 9
Copy number alterations across all tumors
Supplementary Table 10
Predicted HLA-restricted neoantigens across all tumors
Rights and permissions
About this article
Cite this article
Miao, D., Margolis, C.A., Vokes, N.I. et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat Genet 50, 1271–1281 (2018). https://doi.org/10.1038/s41588-018-0200-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0200-2
This article is cited by
-
MRI-derived radiomics assessing tumor-infiltrating macrophages enable prediction of immune-phenotype, immunotherapy response and survival in glioma
Biomarker Research (2024)
-
Mutation status of the KMT2 family associated with immune checkpoint inhibitors (ICIs) therapy and implicating diverse tumor microenvironments
Molecular Cancer (2024)
-
Biomarker discovery with quantum neural networks: a case-study in CTLA4-activation pathways
BMC Bioinformatics (2024)
-
Molecular patterns of resistance to immune checkpoint blockade in melanoma
Nature Communications (2024)
-
Assessment of human leukocyte antigen-based neoantigen presentation to determine pan-cancer response to immunotherapy
Nature Communications (2024)