Abstract
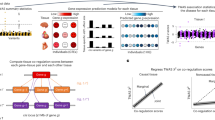
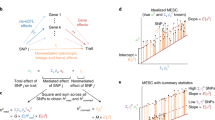
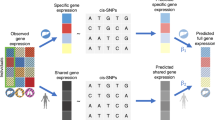
Biological interpretation of genome-wide association study data frequently involves assessing whether SNPs linked to a biological process, for example, binding of a transcription factor, show unsigned enrichment for disease signal. However, signed annotations quantifying whether each SNP allele promotes or hinders the biological process can enable stronger statements about disease mechanism. We introduce a method, signed linkage disequilibrium profile regression, for detecting genome-wide directional effects of signed functional annotations on disease risk. We validate the method via simulations and application to molecular quantitative trait loci in blood, recovering known transcriptional regulators. We apply the method to expression quantitative trait loci in 48 Genotype-Tissue Expression tissues, identifying 651 transcription factor-tissue associations including 30 with robust evidence of tissue specificity. We apply the method to 46 diseases and complex traits (average n = 290 K), identifying 77 annotation-trait associations representing 12 independent transcription factor-trait associations, and characterize the underlying transcriptional programs using gene-set enrichment analyses. Our results implicate new causal disease genes and new disease mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Cowper-Sal lari, R. et al. Breast cancer risk-associated SNPs modulate the affinity of chromatin for FOXA1 and alter gene expression. Nat. Genet. 44, 1191–1198 (2012).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Trynka, G. et al. Chromatin marks identify critical cell types for fine mapping complex trait variants. Nat. Genet. 45, 124–130 (2013).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: from polygenic to omnigenic. Cell 169, 1177–1186 (2017).
Zhu, X. & Stephens, M. A large-scale genome-wide enrichment analysis identifies new trait-associated genes, pathways and tissues across 31 human phenotypes. bioRxiv 160770 (2017).
Karczewski, K. J. et al. Systematic functional regulatory assessment of disease-associated variants. Proc. Natl Acad. Sci., USA 110, 9607–9612 (2013).
Mathelier, A., Shi, W. & Wasserman, W. W. Identification of altered cis-regulatory elements in human disease. Trends Genet. 31, 67–76 (2015).
Price, A. L., Spencer, C. C. A. & Donnelly, P. Progress and promise in understanding the genetic basis of common diseases. Proc. R. Soc. B 282, 20151684 (2015).
Whitington, T. et al. Gene regulatory mechanisms underpinning prostate cancer susceptibility. Nat. Genet. 48, 387–397 (2016).
Liu, Y. et al. Identification of breast cancer associated variants that modulate transcription factor binding. PLoS Genet. 13, e1006761 (2017).
Pique-Regi, R. et al. Accurate inference of transcription factor binding from DNA sequence and chromatin accessibility data. Genome Res. 21, 447–455 (2011).
Lee, D. et al. A method to predict the impact of regulatory variants from DNA sequence. Nat. Genet. 47, 955–961 (2015).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Zeng, H., Hashimoto, T., Kang, D. D. & Gifford, D. K. GERV: a statistical method for generative evaluation of regulatory variants for transcription factor binding. Bioinformatics 32, 490–496 (2016).
Kelley, D. R., Snoek, J. & Rinn, J. Basset: Learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Res. 26, 990–999 (2016).
Pasaniuc, B. & Price, A. L. Dissecting the genetics of complex traits using summary association statistics. Nat. Rev. Genet. 18, 117–127 (2017).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414.e24 (2016).
GTEx Consortium et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci., USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Yang, W. et al. Genome-wide association study in Asian populations identifies variants in ETS1 and WDFY4 associated with systemic lupus erythematosus. PLoS Genet. 6, e1000841 (2010).
Arbiza, L. et al. Genome-wide inference of natural selection on human transcription factor binding sites. Nat. Genet. 45, 723–729 (2013).
Ernst, J. et al. Genome-scale high-resolution mapping of activating and repressive nucleotides in regulatory regions. Nat. Biotechnol. 34, 1180–1190 (2016).
Bodine, D. M. Introduction to a review series on transcription factors in hematopoiesis and hematologic disease. Blood 129, 2039 (2017).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Sharrocks, A. D., Brown, A. L., Ling, Y. & Yates, P. R. The ETS-domain transcription factor family. Int. J. Biochem. Cell Biol. 29, 1371–1387 (1997).
Kimura, T. et al. Involvement of the IRF-1 transcription factor in antiviral responses to interferons. Science 264, 1921–1924 (1994).
Kakizuka, A. et al. Chromosomal translocation t(15;17) in human acute promyelocytic leukemia fuses RARα with a novel putative transcription factor, PML. Cell 66, 663–674 (1991).
Wright, F. A. et al. Heritability and genomics of gene expression in peripheral blood. Nat. Genet. 46, 430–437 (2014).
Friedman, J. S. et al. The minimal transactivation domain of the basic motif-leucine zipper transcription factor NRL interacts with TATA-binding protein. J. Biol. Chem. 279, 47233–47241 (2004).
Bell, A. C., West, A. G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Xie, X. et al. Systematic discovery of regulatory motifs in conserved regions of the human genome, including thousands of CTCF insulator sites. Proc. Natl Acad. Sci., USA 104, 7145–7150 (2007).
Gao, N. et al. Dynamic regulation of Pdx1 enhancers by Foxa1 and Foxa2 is essential for pancreas development. Genes Dev. 22, 3435–3448 (2008).
Song, Y., Washington, M. K. & Crawford, H. C. Loss of FOXA1/2 is essential for the epithelial-to-mesenchymal transition in pancreatic cancer. Cancer Res. 70, 2115–2125 (2010).
Gao, N. et al. Foxa1 and Foxa2 maintain the metabolic and secretory features of the mature β-cell. Mol. Endocrinol. 24, 1594–1604 (2010).
Hagman, J., Ramírez, J. & Lukin, K. B lymphocyte lineage specification, commitment and epigenetic control of transcription by early B cell factor 1. Curr. Top. Microbiol. Immunol. 356, 17–38 (2012).
Somasundaram, R., Prasad, M. A. J., Ungerbäck, J. & Sigvardsson, M. Transcription factor networks in B-cell differentiation link development to acute lymphoid leukemia. Blood 126, 144–152 (2015).
Odom, D. T. et al. Control of pancreas and liver gene expression by HNF transcription factors. Science 303, 1378–1381 (2004).
Bonzo, J. A., Ferry, C. H., Matsubara, T., Kim, J.-H. & Gonzalez, F. J. Suppression of hepatocyte proliferation by hepatocyte nuclear factor 4α in adult mice. J. Biol. Chem. 287, 7345–7356 (2012).
Wolff, L. & Ruscetti, S. The spleen focus-forming virus (SFFV) envelope gene, when introduced into mice in the absence of other SFFV genes, induces acute erythroleukemia. J. Virol. 62, 2158–2163 (1988).
Angel, P. E. & Herrlich, P. The FOS and JUN Families of Transcription Factors. (CRC Press, Boca Raton, FL, USA 1994).
Bullitt, E. Expression of c-fos-like protein as a marker for neuronal activity following noxious stimulation in the rat. J. Comp. Neurol. 296, 517–530 (1990).
Velazquez, F. N. et al. Brain development is impaired in c-fos -/- mice. Oncotarget 6, 16883–16901 (2015).
Zhang, J. et al. c-fos regulates neuronal excitability and survival. Nat. Genet. 30, 416–420 (2002).
Nischan, J. et al. Binding sites for ETS family of transcription factors dominate the promoter regions of differentially expressed genes in abdominal aortic aneurysms. Circ. Genomic Precis. Med. 2, 565–572 (2009).
Triarhou, L. C. Dopamine and Parkinson’s Disease. (Landes Bioscience, Austin, TX, USA, 2013).
Aneichyk, T. et al. Dissecting the causal mechanism of X-linked dystonia-parkinsonism by integrating genome and transcriptome assembly. Cell 172, 897–909.e21 (2018).
Davis, F. P. & Eddy, S. R. Transcription factors that convert adult cell identity are differentially Polycomb repressed. PLoS One 8, e63407 (2013).
Popov, D. V., Lysenko, E. A., Makhnovskii, P. A., Kurochkina, N. S. & Vinogradova, O. L. Regulation of PPARGC1A gene expression in trained and untrained human skeletal muscle. Physiol. Rep. 5, e13543 (2017).
Kim, S., Yu, N.-K. & Kaang, B.-K. CTCF as a multifunctional protein in genome regulation and gene expression. Exp. Mol. Med. 47, e166 (2015).
Kleiman, E., Jia, H., Loguercio, S., Su, A. I. & Feeney, A. J. YY1 plays an essential role at all stages of B-cell differentiation. Proc. Natl Acad. Sci., USA 113, E3911–E3920 (2016).
Hwang, S. S. et al. YY1 inhibits differentiation and function of regulatory T cells by blocking Foxp3 expression and activity. Nat. Commun. 7, 10789 (2016).
Kwon, H.-K., Chen, H.-M., Mathis, D. & Benoist, C. Different molecular complexes that mediate transcriptional induction and repression by FoxP3. Nat. Immunol. 18, 1238–1248 (2017).
Gabriele, M. et al. YY1 haploinsufficiency causes an intellectual disability syndrome featuring transcriptional and chromatin dysfunction. Am. J. Hum. Genet. 100, 907–925 (2017).
Weintraub, A. S. et al. YY1 is a structural regulator of enhancer-promoter loops. Cell 171, 1573–1588.e28 (2017).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Basak, A. et al. BCL11A deletions result in fetal hemoglobin persistence and neurodevelopmental alterations. J. Clin. Invest. 125, 2363–2368 (2015).
Funnell, A. P. W. et al. 2p15-p16.1 microdeletions encompassing and proximal to BCL11A are associated with elevated HbF in addition to neurologic impairment. Blood 126, 89–93 (2015).
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
Dias, C. et al. BCL11A haploinsufficiency causes an intellectual disability syndrome and dysregulates transcription. Am. J. Hum. Genet. 99, 253–274 (2016).
Lipton, J. O. & Sahin, M. The neurology of mTOR. Neuron 84, 275–291 (2014).
Reijnders, M. R. F. et al. Variation in a range of mTOR-related genes associates with intracranial volume and intellectual disability. Nat. Commun. 8, 1052 (2017).
Laplante, M. & Sabatini, D. M. An emerging role of mTOR in lipid biosynthesis. Curr. Biol. 19, R1046–R1052 (2009).
Mathews, E. S. & Appel, B. Cholesterol biosynthesis supports myelin gene expression and axon ensheathment through modulation of P13K/Akt/mTor signaling. J. Neurosci. 36, 7628–7639 (2016).
Koudinov, A. R. & Koudinova, N. V. Cholesterol homeostasis failure as a unifying cause of synaptic degeneration. J. Neurol. Sci. 229, 233–240 (2005).
Zhang, J. & Liu, Q. Cholesterol metabolism and homeostasis in the brain. Protein Cell 6, 254–264 (2015).
Macari, E. R., Schaeffer, E. K., West, R. J. & Lowrey, C. H. Simvastatin and t-butylhydroquinone suppress KLF1 and BCL11A gene expression and additively increase fetal hemoglobin in primary human erythroid cells. Blood 121, 830–839 (2013).
TANG, L. et al. BCL11A gene DNA methylation contributes to the risk of type 2 diabetes in males. Exp. Ther. Med. 8, 459–463 (2014).
Li, S. et al. Transcription factor CTIP1/BCL11A regulates epidermal differentiation and lipid metabolism during skin development. Sci. Rep. 7, 13427 (2017).
Franke, A. et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat. Genet. 42, 1118–1125 (2010).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Lange, K. Mde et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Rioux, J. D. et al. Genetic variation in the 5q31 cytokine gene cluster confers susceptibility to Crohn disease. Nat. Genet. 29, 223–228 (2001).
Silverberg, M. S. OCTNs: Will the real IBD5 gene please stand up? World J. Gastroenterol. 12, 3678–3681 (2006).
Brant, S. R. IBD5: the second Crohn’s disease gene? Inflamm. Bowel Dis. 8, 371–372 (2002).
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Wainberg, M. et al. Vulnerabilities of transcriptome-wide association studies. bioRxiv 206961 (2017).
Mancuso, N. et al. Integrating gene expression with summary association statistics to identify genes associated with 30 complex traits. Am. J. Hum. Genet. 100, 473–487 (2017).
Romeo, G. et al. IRF-1 as a negative regulator of cell proliferation. J. Interferon Cytokine Res. 22, 39–47 (2002).
Honda, K., Takaoka, A. & Taniguchi, T. Type I interferon gene induction by the interferon regulatory factor family of transcription factors. Immunity 25, 349–360 (2006).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Linterman, M. A. et al. IL-21 acts directly on B cells to regulate Bcl-6 expression and germinal center responses. J. Exp. Med. 207, 353–363 (2010).
Chevrier, S., Kratina, T., Emslie, D., Tarlinton, D. M. & Corcoran, L. M. IL4 and IL21 cooperate to induce the high Bcl6 protein level required for germinal center formation. Immunol. Cell Biol. 95, 925–932 (2017).
Hurtz, C. et al. BCL6-mediated repression of p53 is critical for leukemia stem cell survival in chronic myeloid leukemia. J. Exp. Med. 208, 2163–2174 (2011).
Hatzi, K. et al. A hybrid mechanism of action for BCL6 in B cells defined by formation of functionally distinct complexes at enhancers and promoters. Cell Rep. 4, 578–588 (2013).
Huang, C., Hatzi, K. & Melnick, A. Lineage-specific functions of Bcl-6 in immunity and inflammation are mediated by distinct biochemical mechanisms. Nat. Immunol. 14, 380–388 (2013).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed-model association for biobank-scale datasets. Nat. Genet. (2018).
Ek, W. E., Rask-Andersen, M., Karlsson, T. & Johansson, A. Genome-wide association analysis identifies 26 novel loci for asthma, hay fever and eczema. bioRxiv 195933 (2017).
Portelli, M. A., Hodge, E. & Sayers, I. Genetic risk factors for the development of allergic disease identified by genome-wide association. Clin. Exp. Allergy 45, 21–31 (2015).
Boraska, V. et al. A genome-wide association study of anorexia nervosa. Mol. Psychiatry 19, 1085–1094 (2014).
Ben-Shachar, D. & Karry, R. Sp1 expression is disrupted in schizophrenia; a possible mechanism for the abnormal expression of mitochondrial complex I genes, NDUFV1 and NDUFV2. PLoS One 2, e817 (2007).
Fusté, M. et al. Reduced expression of SP1 and SP4 transcription factors in peripheral blood mononuclear cells in first-episode psychosis. J. Psychiatr. Res. 47, 1608–1614 (2013).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Striegel-Moore, R. H. et al. Gender difference in the prevalence of eating disorder symptoms. Int. J. Eat. Disord. 42, 471–474 (2009).
Colman, R. J. et al. Caloric restriction delays disease onset and mortality in rhesus monkeys. Science 325, 201–204 (2009).
Pan, X., Solomon, S. S., Borromeo, D. M., Martinez-Hernandez, A. & Raghow, R. Insulin deprivation leads to deficiency of Sp1 transcription factor in H-411E hepatoma cells and in streptozotocin-induced diabetic ketoacidosis in the rat. Endocrinology 142, 1635–1642 (2001).
Yasui, D., Peedicayil, J. & Grayson, D. R. Neuropsychiatric Disorders and Epigenetics. (Academic Press, Cambridge, MA, USA 2016).
Zhang, X. et al. Hypermethylation of Sp1 binding site suppresses hypothalamic POMC in neonates and may contribute to metabolic disorders in adults: impact of maternal dietary CLAs. Diabetes 63, 1475–1487 (2014).
Yang, G. et al. FoxO1 inhibits leptin regulation of pro-opiomelanocortin promoter activity by blocking STAT3 interaction with specificity protein 1. J. Biol. Chem. 284, 3719–3727 (2009).
Moreno-Aliaga, M. J. et al. Sp1-mediated transcription is involved in the induction of leptin by insulin-stimulated glucose metabolism. J. Mol. Endocrinol. 38, 537–546 (2007).
Audet-Walsh, É. et al. Nuclear mTOR acts as a transcriptional integrator of the androgen signaling pathway in prostate cancer. Genes Dev. 31, 1228–1242 (2017).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23, R89–R98 (2014).
Verbanck, M., Chen, C.-Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Michelson, A. M. Deciphering genetic regulatory codes: A challenge for functional genomics. Proc. Natl Acad. Sci., USA 99, 546–548 (2002).
Deplancke, B., Alpern, D. & Gardeux, V. The genetics of transcription factor DNA binding variation. Cell 166, 538–554 (2016).
Zeng, H., Edwards, M. D., Liu, G. & Gifford, D. K. Convolutional neural network architectures for predicting DNA-protein binding. Bioinformatics 32, i121–i127 (2016).
Kumar, S., Ambrosini, G. & Bucher, P. SNP2transcription factorBS – a database of regulatory SNPs affecting predicted transcription factor binding site affinity. Nucleic Acids Res. 45, D139–D144 (2017).
Yevshin, I., Sharipov, R., Valeev, T., Kel, A. & Kolpakov, F. GTRD: a database of transcription factor binding sites identified by ChIP-seq experiments. Nucleic Acids Res. 45, D61–D67 (2017).
Kulakovskiy, I. V. et al. HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-Seq analysis. Nucleic Acids Res. 46, D252–D259 (2018).
Venkataraman, A. et al. A toolbox of immunoprecipitation-grade monoclonal antibodies to human transcription factors. Nat. Methods (2018).
Berisa, T. & Pickrell, J. K. Approximately independent linkage disequilibrium blocks in human populations. Bioinformatics 32, 283–285 (2016).
Schoech, A. et al. Quantification of frequency-dependent genetic architectures and action of negative selection in 25 UK Biobank traits. bioRxiv 188086 (2017).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Banda, Y. et al. Characterizing race/ethnicity and genetic ancestry for 100,000 subjects in the Genetic Epidemiology Research on Adult Health and Aging (GERA) cohort. Genetics 200, 1285–1295 (2015).
1000 Genomes Project Consortium. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol 57, 289–300 (1995).
Hormozdiari, F. et al. Leveraging molecular quantitative trait loci to understand the genetic architecture of diseases and complex traits. Nat. Genet. 50, 1041–1047 (2017).
Carroll, R. J. Measurement Error in Epidemiologic Studies. in Wiley StatsRef: Statistics Reference Online (Wiley, Hoboken, NJ, USA, 2014).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Acknowledgements
We thank C. de Boer, L. Dicker, J. Engreitz, T. Finucane, N. Friedman, R. Gumpert, M. Kanai, S. Kim, X. Liu, M. Mitzenmacher, J. Perry, S. Reilly, D. Reshef, S. Raychaudhuri, A. Schoech, P. Sabeti, R. Tewhey, O. Troyanskaya, P. Turley, O. Weissbrod, J. Zhou, and the CGTA discussion group for helpful discussions. This research was conducted using the UK Biobank Resource under Application No. 16549 and was supported by US National Institutes of Health grants No. U01 HG009379, R01 MH101244, R01 MH109978, and R01 MH107649. Y.A.R. was supported by award No. T32GM007753 from the National Institute of General Medical Sciences, the National Defense Science and Engineering Graduate Fellowship, and the Paul and Daisy Soros Foundation. H.K.F. was supported by the Fannie and John Hertz Foundation and by Eric and Wendy Schmidt. F.H. is supported by National Institute of Health award No. T32 DK110919. L.P. is supported by National Institutes of Health award No. R00HG008399. R.P.A. is supported by NSF grant No. IIS-1421780. Computational analyses were performed on the Orchestra High Performance Compute Cluster at Harvard Medical School, which is partially supported by grant No. NCRR 1S10RR028832-01. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Y.A.R. and A.L.P. designed the study. Y.A.R., H.K.F., D.R.K., A.G., F.H., J.N., and P.-R.L. analyzed data. Y.A.R. and A.L.P. wrote the manuscript with assistance from H.K.F., D.R.K., A.G., D.K., J.C.U., F.H., J.N., L.O., B.v.d.G., P.-R.L., S.R.G., G.B., S.G., P.F.P., L.P., N.P., and R.P.A.
Corresponding authors
Ethics declarations
Competing interests
D.R.K. is employed by the Calico Life Sciences LLC. The rest of the authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14, Supplementary Tables 1, 13, 14, 16, 20 and 21, and Supplementary Note
Supplementary Table 2
Numerical results for Fig. 1
Supplementary Table 3
Numerical results for Fig. 2
Supplementary Table 4
List of traits analyzed in BLUEPRINT/NTR analysis
Supplementary Table 5
Details of results of BLUEPRINT/NTR analysis
Supplementary Table 6
List of GTEx traits analyzed
Supplementary Table 7
Results of GTEx analysis
Supplementary Table 8
List of diseases and complex traits analyzed
Supplementary Table 9
Results of SLDP analysis of 46 diseases and complex traits
Supplementary Table 10
Results of enrichment analysis of signed LD profile regression disease/complex trait analysis
Supplementary Table 11
Numerical results for Fig. 6
Supplementary Table 12
Numerical results for Fig. 7
Supplementary Table 15
Numerical results for Supplementary Fig. 9
Supplementary Table 17
Results of signed LD profile regression using DeepSEA-based annotations
Supplementary Table 18
Results of signed LD profile regression using GTRD-based annotations
Supplementary Table 19
Results of signed LD profile regression using HOCOMOCO motif-based annotations
Rights and permissions
About this article
Cite this article
Reshef, Y.A., Finucane, H.K., Kelley, D.R. et al. Detecting genome-wide directional effects of transcription factor binding on polygenic disease risk. Nat Genet 50, 1483–1493 (2018). https://doi.org/10.1038/s41588-018-0196-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0196-7
This article is cited by
-
Benchmarking of deep neural networks for predicting personal gene expression from DNA sequence highlights shortcomings
Nature Genetics (2023)
-
Modeling tissue co-regulation estimates tissue-specific contributions to disease
Nature Genetics (2023)
-
Epigenomic charting and functional annotation of risk loci in renal cell carcinoma
Nature Communications (2023)
-
A sequence-based global map of regulatory activity for deciphering human genetics
Nature Genetics (2022)
-
Identifying interpretable gene-biomarker associations with functionally informed kernel-based tests in 190,000 exomes
Nature Communications (2022)