Abstract
Neuronal migration defects, including pachygyria, are among the most severe developmental brain defects in humans. Here, we identify biallelic truncating mutations in CTNNA2, encoding αN-catenin, in patients with a distinct recessive form of pachygyria. CTNNA2 was expressed in human cerebral cortex, and its loss in neurons led to defects in neurite stability and migration. The αN-catenin paralog, αE-catenin, acts as a switch regulating the balance between β-catenin and Arp2/3 actin filament activities1. Loss of αN-catenin did not affect β-catenin signaling, but recombinant αN-catenin interacted with purified actin and repressed ARP2/3 actin-branching activity. The actin-binding domain of αN-catenin or ARP2/3 inhibitors rescued the neuronal phenotype associated with CTNNA2 loss, suggesting ARP2/3 de-repression as a potential disease mechanism. Our findings identify CTNNA2 as the first catenin family member with biallelic mutations in humans, causing a new pachygyria syndrome linked to actin regulation, and uncover a key factor involved in ARP2/3 repression in neurons.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Drees, F., Pokutta, S., Yamada, S., Nelson, W. J. & Weis, W. I. α-Catenin is a molecular switch that binds E-cadherin-β-catenin and regulates actin-filament assembly. Cell 123, 903–915 (2005).
Leventer, R. J., Guerrini, R. & Dobyns, W. B. Malformations of cortical development and epilepsy. Dialogues Clin. Neurosci. 10, 47–62 (2008).
Ayala, R., Shu, T. & Tsai, L.-H. Trekking across the brain: the journey of neuronal migration. Cell 128, 29–43 (2007).
Meyer, G. & Feldman, E. L. Signaling mechanisms that regulate actin-based motility processes in the nervous system. J. Neurochem. 83, 490–503 (2002).
Luo, L. Actin cytoskeleton regulation in neuronal morphogenesis and structural plasticity. Annu. Rev. Cell Dev. Biol. 18, 601–635 (2002).
Rocca, D. L., Martin, S. P., Jenkins, E. L. & Hanley, J. G. Inhibition of Arp2/3-mediated actin polymerization by PICK1 regulates neuronal morphology and AMPA receptor endocytosis. Nat. Cell Biol. 10, 259–271 (2008).
Choi, M. et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc. Natl. Acad. Sci. USA 106, 19096–19101 (2009).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Magi, A. et al. H3M2: detection of runs of homozygosity from whole-exome sequencing data. Bioinformatics 30, 2852–2859 (2014).
Seelow, D., Schuelke, M., Hildebrandt, F. & Nurnberg, P. HomozygosityMapper—an interactive approach to homozygosity mapping. Nucleic Acids Res. 37, W593–W599 (2009).
Ajioka, I. & Nakajima, K. Switching of alpha-catenin from alphaE-catenin in the cortical ventricular zone to alphaN-catenin II in the intermediate zone. Brain Res. Dev. Brain Res. 160, 106–111 (2005).
Stocker, A. M. & Chenn, A. Differential expression of alpha-E-catenin and alpha-N-catenin in the developing cerebral cortex. Brain Res. 1073–1074, 151–158 (2006).
Park, C., Falls, W., Finger, J. H., Longo-Guess, C. M. & Ackerman, S. L. Deletion in Catna2, encoding αN-catenin, causes cerebellar and hippocampal lamination defects and impaired startle modulation. Nat. Genet. 31, 279–284 (2002).
Park, C., Longo, C. M. & Ackerman, S. L. Genetic and physical mapping of the cerebellar deficient folia (cdf) locus on mouse chromosome 6. Genomics 69, 135–138 (2000).
Park, C., Finger, J. H., Cooper, J. A. & Ackerman, S. L. The cerebellar deficient folia (cdf) gene acts intrinsically in Purkinje cell migrations. Genesis 32, 32–41 (2002).
Togashi, H. et al. Cadherin regulates dendritic spine morphogenesis. Neuron 35, 77–89 (2002).
Abe, K., Chisaka, O., van Roy, F. & Takeichi, M. Stability of dendritic spines and synaptic contacts is controlled by alpha N-catenin. Nat. Neurosci. 7, 357–363 (2004).
Uemura, M. & Takeichi, M. αN-catenin deficiency causes defects in axon migration and nuclear organization in restricted regions of the mouse brain. Dev. Dyn. 235, 2559–2566 (2006).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Katakowski, M., Zhang, Z., deCarvalho, A. C. & Chopp, M. EphB2 induces proliferation and promotes a neuronal fate in adult subventricular neural precursor cells. Neurosci. Lett. 385, 204–209 (2005).
Bershteyn, M. et al. Human iPSC-derived cerebral organoids model cellular features of lissencephaly and reveal prolonged mitosis of outer radial glia. Cell Stem Cell 20, 435–449.e4 (2017).
Torres, M. et al. An α-E-catenin gene trap mutation defines its function in preimplantation development. Proc. Natl. Acad. Sci. USA 94, 901–906 (1997).
Lein, W.-H., Klezovitch, O., Fernandez, T. E., Delrow, J. & Vasioukhin, V. αE-catenin controls cerebral cortical size by regulating the hedgehog signaling pathway. Science 311, 1609–1612 (2006).
Pokutta, S. & Weis, W. I. Structure of the dimerization and β-catenin-binding region of α-catenin. Mol. Cell 5, 533–543 (2000).
Hansen, S. D. et al. αE-catenin actin-binding domain alters actin filament conformation and regulates binding of nucleation and disassembly factors. Mol. Biol. Cell 24, 3710–3720 (2013).
Wang, P.-S. et al. Crucial roles of the Arp2/3 complex during mammalian corticogenesis. Development 143, 2741–2752 (2016).
Rouiller, I. et al. The structural basis of actin filament branching by the Arp2/3 complex. J. Cell Biol. 180, 887–895 (2008).
Hetrick, B., Han, M. S., Helgeson, L. A. & Nolen, B. J. Small molecules CK-666 and CK-869 inhibit actin-related protein 2/3 complex by blocking an activating conformational change. Chem. Biol. 20, 701–712 (2013).
Wynshaw-Boris, A., Pramparo, T., Youn, Y. H. & Hirotsune, S. Lissencephaly: mechanistic insights from animal models and potential therapeutic strategies. Semin. Cell Dev. Biol. 21, 823–830 (2010).
Moon, H. M. & Wynshaw-Boris, A. Cytoskeleton in action: lissencephaly, a neuronal migration disorder. Wiley Interdiscip. Rev. Dev. Biol. 2, 229–245 (2013).
Solecki, D. J., Model, L., Gaetz, J., Kapoor, T. M. & Hatten, M. E. Par6α signaling controls glial-guided neuronal migration. Nat. Neurosci. 7, 1195–1203 (2004).
Strasser, G. A., Rahim, N. A., VanderWaal, K. E., Gertler, F. B. & Lanier, L. M. Arp2/3 is a negative regulator of growth cone translocation. Neuron 43, 81–94 (2004).
Kholmanskikh, S. S., Dobrin, J. S., Wynshaw-Boris, A., Letourneau, P. C. & Ross, E. M. Disregulated RhoGTPases and actin cytoskeleton contribute to the migration defect in Lis1-deficient neurons. J. Neurosci. 23, 8673–8681 (2003).
Chai, X., Forster, E., Zhao, S., Bock, H. H. & Frotscher, M. Reelin stabilizes the actin cytoskeleton of neuronal processes by inducing n-cofilin phosphorylation at serine3. J. Neurosci. 29, 288–299 (2009).
Zhou, F.-Q., Waterman-Storer, C. M. & Cohan, C. S. Focal loss of actin bundles causes microtubule redistribution and growth cone turning. J. Cell Biol. 157, 839–849 (2002).
Winkelman, J. D. et al. Fascin- and α-actinin-bundled networks contain intrinsic structural features that drive protein sorting. Curr. Biol. 26, 2697–2706 (2016).
Hoffmann, K. & Lindner, T. H. easyLINKAGE-Plus—automated linkage analyses using large-scale SNP data. Bioinformatics 21, 3565–3567 (2005).
Lardelli, R. M. et al. Bi-allelic mutations in the 3′ exonuclease TOE1 cause pontocerebellar hypoplasia and uncover a role in snRNA processing. Nat. Genet. 49, 457–464 (2017).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 8, 409–412 (2011).
Marchetto, M. C. N. et al. A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 143, 527–539 (2010).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Youn, Y. H., Pramparo, T., Hirotsune, S. & Wynshaw-Boris, A. Distinct dose-dependent cortical neuronal migration and neurite extension defects in Lis1 and Ndel1 mutant mice. J. Neurosci. 29, 15520–15530 (2009).
Yeung, Y. G., Wang, Y., Einstein, D. B., Lee, P. S. & Stanley, E. R. Colony-stimulating factor-1 stimulates the formation of multimeric cytosolic complexes of signaling proteins and cytoskeletal components in macrophages. J. Biol. Chem. 273, 17128–17137 (1998).
Gleeson, J. G., Lin, P. T., Flanagan, L. A. & Walsh, C. A. Doublecortin is a microtubule-associated protein and is expressed widely by migrating neurons. Neuron 23, 257–271 (1999).
Acknowledgements
We thank the patients and their families for participation. We thank A. Wynshaw-Boris for generous scientific and editorial input. The research was supported by NIH R01NS041537, R01NS048453, R01NS052455, P01HD070494, P30NS047101, Qatar National Research Fund number 6-1463-351, the Simons Foundation Autism Research Initiative, and the Howard Hughes Medical Institute (to J.G.G). A.E.S. is a recipient of an A.P. Giannini Fellowship and an NIH Pathway to Independence Award, R00HD082337. S.T.B. is supported by a 2014 NARSAD Young Investigator Grant from the Brain and Behavior Research Foundation. We thank the Broad Institute and Yale Center for Mendelian Disorders (UMIHG008900 to D. MacArthur and H. Rehm, and UMIHG006504 to R. Lifton and M.G.), and the Gregory M. Kiez and Mehmet Kutman Foundation (to M.G). We acknowledge M. Gerstein, S. Mane, A. B. Ekici, and S. Uebe for sequencing support and analysis, the Yale Biomedical High Performance Computing Center for data analysis and storage, the Yale Program on Neurogenetics, and the Yale Center for Human Genetics and Genomics. Exome data have been deposited into the database of Genotypes and Phenotypes (phs000288).
Author information
Authors and Affiliations
Contributions
N.A-S., H.Y.A-A., H.Kaymakçalan, C.Y., R.O.R., V.S., K.N.J., M.S.Z., S.E., T.B-O., A.Karminejad, H.Kayserili, F.M., M.K., E.Fenercioglu, B.K., H.M., F.I., S.D., Y.A., E.Faqeih, G.M., B.A.B., L.M., I.M., B.S., J.C., W.B.D., M.G., J.G.G., and R.A.J. performed the patient recruitment and phenotyping. A.E.S., A.O.C., S.T.B., B.C., D.M., E.C.S., K.B., J.L.S., M.G., and J.G.G. supported the sequencing and variant interpretation. A.E.S., N.C., R.W., and R.N. performed the tissue culture experiments. A.E.S. and A.Kalur generated recombinant proteins and performed the actin assays. A.E.S., M.W.B., and J.G.G. wrote and edited the manuscript. J.G.G. directed the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Confirmation of CTNNA2-truncating mutation.
a, Intervals of homozygosity from families 1101, 1263, and 4727. The patient mutations in CTNNA2 (red arrow) were found within an overlapping interval on chromosome 2 in each family. b, Potentially deleterious variants identified from WES in families 1101, 1263, and 4727. Chr., chromosome; ref, reference allele; alt, alternative allele.
Supplementary Fig. 2 Expression of CTNNA2 encoding αN-catenin in developing human cortex and codistribution with markers of migration.
a, qRT–PCR analysis of CTNNA2 abundance across human tissues. The strongest expression is in fetal and mature brain, with lower levels in other tissues. GAPDH was used as a loading control. Quantitative expression of CTNNA2 was normalized to GAPDH and then to average CTNNA2 expression across all tissues. Experiments were repeated in triplicate, quantified at the right and plotted as the mean (bar) with each data point (red dot). Error bars, s.e.m. b, Location of αN-catenin relative to Nestin and codistribution with markers of neuronal migration including Dcx and Tuj1 (arrows) at embryonic day (E) 13.5 in mouse. CP, cortical plate; SVZ, subventricular zone; VZ, ventricular zone; DAPI (blue). Repeated in three biological replicates. Scale bar, 50 μm. c, Localization of αN-catenin (brown) in human 20-week gestation fetal cortex in CP and marginal zone (MZ), compared with TUJ1 and DCX, counterstained with Nissl. Bottom, αN-catenin (brown) shows signal along the apical surface of the VZ, similar to Nestin. Repeated in duplicate. Scale bar, 100 μm.
Supplementary Fig. 3 Generation of control, MDS, and CTNNA2-mutant stem cells and NPCs.
a, iPSCs and NPCs derived from patient cells are indistinguishable. Patient-derived iPSCs express stem cell markers OCT4, TRA1-81, NANOG, LIN28, and TRA1-60 and form EBs and neural rosettes upon neural induction. Isolated NPCs are PAX6 and NESTIN co-positive. Representative images shown, repeated in three independently generated clones per patient. Scale bar, 50 μm. b, Karyotyping of iPSCs from family 1263 shows normal karyotype after reprogramming. Patients with MDS show a deletion of chromosome 17p11.3 as expected. Karyotyping was performed once for each independently generated clone, for each patient. c, Schematic of CRISPR–Cas9-mediated CTNNA2 mutagenesis and cell morphology and marker expression during neural differentiation. d, CTNNA2KO stem cells form EBs and neural rosettes upon neural induction. Isolated NPCs are PAX6 and NESTIN co-positive. Representative images shown, repeated in three independently generated CTNNA2KO clones. Scale bar, 50 μm. e, Chromatograms of the Cas9 cleavage site from three independent CTNNA2KO stem cell lines used for experiments. Homozygous (clones 1 and 2) or compound heterozygous (clone 3) Cas9-induced frameshift mutations in exon 12 are detected.
Supplementary Fig. 4 CTNNA2-mutant NPCs have less αN-catenin.
Western blot of αN-catenin in control, 1263 affected (A), and CTNNA2KO NPCs show the patient, and CRISPR–SpCas9-mediated mutations reduce the levels of αN-catenin protein. GAPDH was a loading control. Densitometry quantification (at right) is shown from three independent iPSC clones per patient and three CTNNA2KO clones. Top box, Q3; bottom box, Q1; median displayed. The whiskers represent the minimum and maximum values observed in the dataset. **, significant P values (see “Statistics and reproducibility”).
Supplementary Fig. 5 Migrating cells of the neuronal migration assay are postmitotic neurons.
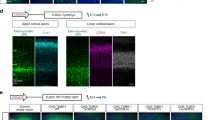
a, Low-magnification immunofluorescent staining of cells exiting the neuronal progenitor cell aggregate over the course of the migration assay. Migrating cells express the immature neuron marker DCX. Representative images shown, repeated in three independently generated patient-derived iPSC or CTNNA2KO hESC clones. b, High magnification of cells assessed by the migration assay. Shown are immunofluorescent staining for the postmitotic markers Tuj1, MAP2, and DCX, the progenitor markers SOX2, Nestin, PAX6, BLBP, and GFAP, and the oligodentrocyte marker OLIG2. In addition, nuclei were visualized by DAPI staining. Arrowheads indicate representative cells that would have been counted in our migration assay. Control cells were positive for neuronal markers, but not however for any of the progenitor markers or OLIG2. Note that the migration-deficient cell lines (i.e., the CTNNA2KO line 1263A and the MDS-derived line) are similar to control and suggest that the observed migration effect is due to defective neuronal migration and not differences in differentiation potential. Representative images shown, repeated in three independently generated patient-derived iPSC or CTNNA2KO hESC clones. Scale bar, 20 μm.
Supplementary Fig. 6 Distribution of neuronal cells following an assay of neuronal migration.
Migration assay from human iPSC-derived neural progenitor cells. The distance neuronal cell bodies migrate from the edge of a neuronal aggregate after 48 h is measured and binned into 30-μm increments, plotted as the average ± s.e.m. The graph of MDS, 1263A, and CTNNA2KO stem cell–derived neurons shows a left-shifted pattern, as neurons from patients with Miller–Dieker syndrome or CTNNA2 mutations do not migrate as far as control neurons. Treatment with lentivirus encoding full-length CTNNA2 or the actin-binding domain (ABD) of αN-catenin (CTNNA2ABD), but not αN-catenin lacking the ABD (CTNNA2ΔABD), was sufficient to restore the distribution of neurons in the assay. Results shown from three different iPSC clones per patient or three CTNNA2KO clones, 1,360 cells scored. The data are also displayed as a whisker box plot in Fig. 2a,c.
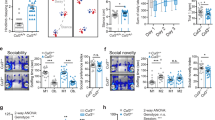
Supplementary Fig. 7 Neuronal cell body migration distance and neurite length as displayed by stem cell clone identifier.
Neurosphere neuronal migration assays from human iPSC-derived neuronal progenitor cells were performed with three different iPSCs or H9 hESC CTNNA2-mutant stem cell lines for each experiment. The distribution of migrating neurons is shown as a box plot: top box, Q3; bottom box, Q1; and median. The whiskers represent the minimum and maximum values observed within each of the datasets. 2,194 cells were quantified. During migration, the length of the leading process from the edge of the neuronal cell body to the tip of the growth cone was measured. The distribution of neurite length is shown, depicted as a box plot: top box, Q3; bottom box, Q1; median. The whiskers represent the minimum and maximum values observed for each dataset. 1,574 cells were quantified.
Supplementary Fig. 8 Expression of human CTNNA2 in control and CTNNA2-deficient NPCs.
a, Long and short exposures of a western blot for αN-catenin in control, control + CTNNA2, 1263A + CTNNA2, and CTNNA2KO + CTNNA2 NPCs show similar levels of forced CTNNA2-GFP expression in the lentivirus-transduced NPC lines. A cleavage project was observed but not further investigated. Representative image shown, repeated in three independently generated patient-derived iPSC or CTNNA2KO hESC clones. b, Long and short exposures of a western blot for αN-catenin in control, control + CTNNA2ΔABD, 1263A + CTNNA2ΔABD, and CTNNA2KO + CTNNA2ΔABD NPCs showed nearly endogenous levels of forced CTNNA2ΔABD-GFP expression in lentivirus-transduced NPC lines. GAPDH loading control. Representative image shown, repeated in three independently generated patient-derived iPSC or CTNNA2KO hESC clones. c. Long and short exposure of Western blot for GFP in Control, Control + CTNNA2ABD, 1263A + CTNNA2ABD, and CTNNA2KO + CTNNA2ABD NPCs show forced CTNNA2ABD-GFP expression in lentivirus transduced NPC lines. GAPDH loading control. Representative image shown, repeated in three independently generated patient-derived iPSC or CTNNA2KO hESC clones.
Supplementary Fig. 9 Loss of CTNNA2 does not disrupt neuroepithelial polarity in neural rosettes.
a, Immunofluorescence for αN-catenin in patient-derived neural rosettes. DAPI counterstain shows the cell nucleus. Patient rosettes showed absent αN-catenin staining (bottom). The white dashed line outlines the apical lumen. Representative image shown, repeated in three independently generated patient-derived iPSC clones. Scale bar, 25 μm. b, Immunofluorescence for apically localized markers in patient-derived neural rosettes. Shown is the primary cilia marker ARL13B in combinations with the tight junction marker ZO-1 and the adherens junction markers N-cadherin and β-catenin. DAPI counterstain shows the cell nucleus. Both control and patient-derived rosettes form polarized domains that are positive for apical markers and localize the primary cilium to this domain. Representative image shown, repeated in three independently generated patient-derived iPSC clones. Scale bar, 20 μm.
Supplementary Fig. 10 Loss of αN-catenin does not alter gene expression.
a, High Pearson correlation values across RNA-sequencing biological replicates of neural progenitor cells showing consistent expression profiles between individuals. Repeated in two independently generated iPSC-derived clones per patient. Pearson correlation was calculated by using the filtered (median FPKM >1) and log2-transformed gene expression values from each dataset (n = 12,769 genes). b, RNA-sequencing results indicate that <4% of all detected transcripts were differentially expressed in CTNNA2-mutant patient-derived NPCs, suggesting αN-catenin loss does not affect global transcription rates. c, Heat map of differentially expressed transcripts in control, MDS, and 1263A NPCs show few similarities between MDS or CTNNA2-mutated patients. d. αN-catenin does not measurably affect Wnt signaling in NPCs. qRT–PCR of Wnt target genes AXIN2, ID2, and LEF1 showed similar expression levels in CTNNA2-mutant and control NPCs. The distribution of expression is shown depicted as a box plot: top box, Q3; bottom box, Q1; median. The whiskers represent the minimum and maximum values observed for each dataset. Repeated in two independently generated iPSC clones, in triplicate. Two-tailed Student’s t test was used to calculate significance: AXIN2, P = 0.814; ID2, P = 0.525; LEF1, P = 0.319.
Supplementary Fig. 11 Generation of recombinant αN-catenin protein.
GST-tagged full-length αN-catenin, and a fragment of αN-catenin containing the actin-binding domain (ABD), was produced in BL21 bacterial cells. Cell lysates were run on an SDS–PAGE gel alongside BSA standards and Coomassie stained to show full-length and ABD αN-catenin expression (black arrows) following affinity purification. Repeated in triplicate.
Supplementary Fig. 12 αN-catenin mutations do not affect actin ratios in patient neuronal progenitor cells.
Western blot of fractionated G- and F-actin from control and affected individuals with CTNNA2 mutation. Densitometry quantification on right. Top box, Q3; bottom box, Q1; median. The whiskers represent the minimum and maximum values observed in the dataset. Repeated with three biological replicates. Two-tailed Student’s t test was used to calculate significance: P = 0.354.
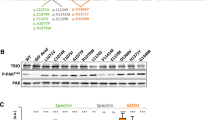
Supplementary Fig. 13 The actin-binding domain of αN-catenin is sufficient to repress Arp2/3-mediated actin polymerization.
a, Full-length western blots corresponding to the cropped western blots in Fig. 4a. Repeated in three biological replicates. b, Full-length western blots corresponding to the cropped western blots in Fig. 4c. Repeated in two biological replicates. c, Actin polymerization assay showing dose-dependent inhibition of Arp2/3 + VCA domain of WASP mediated actin polymerization by recombinant αN-catenin ABD, measured by relative fluorescent units. In the absence of Arp2/3 (black), there was minimal polymerization, but when Arp2/3 + VCA was added (green) polymerization increased substantially, the effect of which was reversed upon dose escalation of αN-catenin ABD. Repeated in duplicate.
Supplementary Fig. 14 Pharmacological inhibition of ARP2/3 restores neurite length in CTNNA2-mutant, but not MDS, neurons.
ARP2/3 inhibition by CK-869 rescued the neurite length defect in CTNNA2-mutant dissociated neurons, but not in wild-type and MDS mutant neurons. Quantification shown as a box plot: top box, Q3; bottom box, Q1, median. The whiskers represent the minimum and maximum values observed in the dataset. Repeated in three independent iPSC clones per patient or three CTNNA2KO clones, 581 cells were scored. ***, significant P values (see “Statistics and reproducibility”). Scale bar, 50 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Table 1
Supplementary Video 1
Control neurons develop bipolar morphology and migrate unidirectionally. Time-lapse imaging shows immature neurons derived from H9 hESCs generate a stable, leading process and predominantly migrate in a single direction over time. Elapsed time ~4 h. Repeated in triplicate
Supplementary Video 2
Neurons derived from patient 1263A have reduced neurite stability and velocity during migration. Neurons derived from patient 1263A show frequent collapse of the primary neurite, with abnormal cellular movement driven by disorganized lamelipodia. Elapsed time ~4 h. Repeated in three independently generated iPSC-derived clones
Supplementary Video 3
CTNNA2KO neurons display unstable primary neurites and aberrant neuronal migration. Similar to neurons derived from patient 1263A, neurons from CTNNA2-knockout hESCs showed loss of bipolar morphology due to frequent collapse of the primary neurite and erratic cellular movement. Elapsed time ~4 h. Repeated in three independently generated CTNNA2KO hESC clones
Rights and permissions
About this article
Cite this article
Schaffer, A.E., Breuss, M.W., Caglayan, A.O. et al. Biallelic loss of human CTNNA2, encoding αN-catenin, leads to ARP2/3 complex overactivity and disordered cortical neuronal migration. Nat Genet 50, 1093–1101 (2018). https://doi.org/10.1038/s41588-018-0166-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0166-0
This article is cited by
-
Non-coding de novo mutations in chromatin interactions are implicated in autism spectrum disorder
Molecular Psychiatry (2022)
-
Cul3 regulates cytoskeleton protein homeostasis and cell migration during a critical window of brain development
Nature Communications (2021)
-
Association between maternal depression during pregnancy and newborn DNA methylation
Translational Psychiatry (2021)
-
RNA m6A modification orchestrates a LINE-1–host interaction that facilitates retrotransposition and contributes to long gene vulnerability
Cell Research (2021)
-
Infant inhibited temperament in primates predicts adult behavior, is heritable, and is associated with anxiety-relevant genetic variation
Molecular Psychiatry (2021)