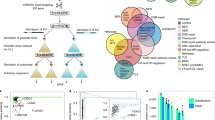
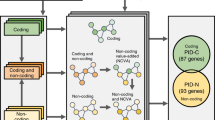
Abstract
Functional redundancy shared by paralog genes may afford protection against genetic perturbations, but it can also result in genetic vulnerabilities due to mutual interdependency1,2,3,4,5. Here, we surveyed genome-scale short hairpin RNA and CRISPR screening data on hundreds of cancer cell lines and identified MAGOH and MAGOHB, core members of the splicing-dependent exon junction complex, as top-ranked paralog dependencies6,7,8. MAGOHB is the top gene dependency in cells with hemizygous MAGOH deletion, a pervasive genetic event that frequently occurs due to chromosome 1p loss. Inhibition of MAGOHB in a MAGOH-deleted context compromises viability by globally perturbing alternative splicing and RNA surveillance. Dependency on IPO13, an importin-β receptor that mediates nuclear import of the MAGOH/B-Y14 heterodimer9, is highly correlated with dependency on both MAGOH and MAGOHB. Both MAGOHB and IPO13 represent dependencies in murine xenografts with hemizygous MAGOH deletion. Our results identify MAGOH and MAGOHB as reciprocal paralog dependencies across cancer types and suggest a rationale for targeting the MAGOHB-IPO13 axis in cancers with chromosome 1p deletion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Diss, G. et al. Gene duplication can impart fragility, not robustness, in the yeast protein interaction network. Science 355, 630–634 (2017).
Muller, F. L., Aquilanti, E. A. & DePinho, R. A. Collateral lethality: a new therapeutic strategy in oncology. Trends Cancer 1, 161–173 (2015).
Frei, E. Gene deletion: a new target for cancer chemotherapy. Lancet 342, 662–664 (1993).
McDonald, E. R. 3rd. et al. Project DRIVE: A compendium of cancer dependencies and synthetic lethal relationships uncovered by large-scale, deep RNAi screening. Cell 170, 577–592.e10 (2017).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e16 (2017).
Boehm, V. & Gehring, N. H. Exon junction complexes: supervising the gene expression assembly line. Trends Genet. 32, 724–735 (2016).
Chang, Y.-F., Imam, J. S. & Wilkinson, M. F. The nonsense-mediated decay RNA surveillance pathway. Annu. Rev. Biochem. 76, 51–74 (2007).
Singh, K. K., Wachsmuth, L., Kulozik, A. E. & Gehring, N. H. Two mammalian MAGOH genes contribute to exon junction complex composition and nonsense-mediated decay. RNA Biol. 10, 1291–1298 (2013).
Mingot, J.-M., Kostka, S., Kraft, R., Hartmann, E. & Görlich, D. Importin 13: a novel mediator of nuclear import and export. EMBO J. 20, 3685–3694 (2001).
Helming, K. C. et al. ARID1B is a specific vulnerability in ARID1A-mutant cancers. Nat. Med. 20, 251–254 (2014).
Hoffman, G. R. et al. Functional epigenetics approach identifies BRM/SMARCA2 as a critical synthetic lethal target in BRG1-deficient cancers. Proc. Natl Acad. Sci. USA 111, 3128–3133 (2014).
D'Antonio, M. et al. Recessive cancer genes engage in negative genetic interactions with their functional paralogs. Cell Rep. 5, 1519–1526 (2013).
Dey, P. et al. Genomic deletion of malic enzyme 2 confers collateral lethality in pancreatic cancer. Nature 542, 119–123 (2017).
Cowley, G. S. et al. Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies. Sci. Data 1, 140035 (2014).
Meyers, R. Computational correction of copy-number effect improves specificity of CRISPR-Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Veitia, R. A. Gene duplicates: agents of robustness or fragility? Trends Genet. 33, 377–379 (2017).
Bono, F. & Gehring, N. H. Assembly, disassembly and recycling. RNA Biol. 8, 24–29 (2011).
Caron, H. et al. Allelic loss of chromosome 1p as a predictor of unfavorable outcome in patients with neuroblastoma. N. Engl. J. Med. 334, 225–230 (1996).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Ni, J. Z. et al. Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay. Genes Dev. 21, 708–718 (2007).
Lareau, L. F., Inada, M., Green, R. E., Wengrod, J. C. & Brenner, S. E. Unproductive splicing of SR genes associated with highly conserved and ultraconserved DNA elements. Nature 446, 926–929 (2007).
Lau, C.-K., Diem, M. D., Dreyfuss, G. & Van Duyne, G. D. Structure of the Y14-Magoh core of the exon junction complex. Curr. Biol. 13, 933–941 (2003).
Ren, Y. et al. Targeted tumor-penetrating siRNA nanocomplexes for credentialing the ovarian cancer oncogene ID4. Sci. Transl. Med. 4, 147ra112 (2012).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Baudis, M. Genomic imbalances in 5918 malignant epithelial tumors: an explorative meta-analysis of chromosomal CGH data. BMC Cancer 7, 226 (2007).
Nijhawan, D. et al. Cancer vulnerabilities unveiled by genomic loss. Cell 150, 842–854 (2012).
Paolella, B. R. et al. Copy-number and gene dependency analysis reveals partial copy loss of wild-type SF3B1 as a novel cancer vulnerability. eLife 6, e23268 (2017).
Lee, S. C.-W. et al. Modulation of splicing catalysis for therapeutic targeting of leukemia with mutations in genes encoding spliceosomal proteins. Nat. Med. 22, 672–678 (2016).
Obeng, E. A. et al. Physiologic expression of Sf3b1(K700E) causes impaired erythropoiesis, aberrant splicing, and sensitivity to therapeutic spliceosome modulation. Cancer Cell 30, 404–417 (2016).
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2014).
Krönke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014).
Hwang, S.-Y. et al. Direct targeting of β-catenin by a small molecule stimulates proteasomal degradation and suppresses oncogenic Wnt/β-catenin signaling. Cell Rep. 16, 28–36 (2016).
Kerres, N. et al. Chemically induced degradation of the oncogenic transcription factor BCL6. Cell Rep. 20, 2860–2875 (2017).
Han, T. et al. Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science 356, eaal3755 (2017).
Soderholm, J. F. et al. Importazole, a small molecule inhibitor of the transport receptor importin-β. ACS Chem. Biol. 6, 700–708 (2011).
Hintersteiner, M. et al. Identification of a small molecule inhibitor of importin beta mediated nuclear import by confocal on-bead screening of tagged one-bead one-compound libraries. ACS Chem. Biol. 5, 967–979 (2010).
Wagstaff, K. M., Sivakumaran, H., Heaton, S. M., Harrich, D. & Jans, D. A. Ivermectin is a specific inhibitor of importin α/β-mediated nuclear import able to inhibit replication of HIV-1 and dengue virus. Biochem. J. 443, 851–856 (2012).
Yang, X. et al. A public genome-scale lentiviral expression library of human ORFs. Nat. Methods 8, 659–661 (2011).
Lippa, M. S. et al. Expression of anti-apoptotic factors modulates Apo2L/TRAIL resistance in colon carcinoma cells. Apoptosis 12, 1465–1478 (2007).
Brown, C. Y. et al. Robust, reversible gene knockdown using a single lentiviral short hairpin RNA vector. Hum. Gene Ther. 21, 1005–1017 (2010).
Root, D. E., Hacohen, N., Hahn, W. C., Lander, E. S. & Sabatini, D. M. Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat. Methods 3, 715–719 (2006).
Kwon, E. J., Dudani, J. S. & Bhatia, S. N. Ultrasensitive tumour-penetrating nanosensors of protease activity. Nat. Biomed. Eng. 1, 0054 (2017).
McLachlan, G. J. & Krishnan, T. The EM Algorithm and Extensions 2nd edn (Wiley-Interscience, Hoboken, NJ, 2008).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017)..
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Wickham, H. ggplot2 https://doi.org/10.1007/978-0-387-98141-3 (Springer, New York, 2009).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33, 676–689 (2018).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. JMLR 12, 2825–2830 (2011).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Vilella, A. J. et al. EnsemblCompara GeneTrees: complete, duplication-aware phylogenetic trees in vertebrates. Genome Res. 19, 327–335 (2008).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Aken, B. L. et al. Ensembl 2017. Nucleic Acids Res. 45, D635–D642 (2017).
Pimentel, H. J., Bray, N., Puente, S., Melsted, P. & Pachter, L. Differential analysis of RNA-seq incorporating quantification uncertainty. Nat. Methods 14, 687–690 (2016).
The Gene Ontology Consortium. Gene Ontology Consortium: going forward. Nucleic Acids Res. 43, D1049–D1056 (2015).
Mi, H., Muruganujan, A., Casagrande, J. T. & Thomas, P. D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 8, 1551–1566 (2013).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-seq data. Proc. Natl Acad. Sci. USA 111, E5593–E5601 (2014).
Durinck, S. et al. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
Kim, J. W. et al. Characterizing genomic alterations in cancer by complementary functional associations. Nat. Biotechnol. 34, 539–546 (2016).
Sveen, A., Kilpinen, S., Ruusulehto, A., Lothe, R. A., & Skotheim, R. I. Aberrant RNA splicing in cancer; expression changes and driver mutations of splicing factor genes. Oncogene 35, 2413–2427 (2015).
Acknowledgements
S.R.V. was supported by a Young Investigator Award from the American Society of Clinical Oncology. This work was supported by a National Cancer Institute grant 1R35CA197568 and an American Cancer Society Research Professorship to M.M. P.T. was supported by NIH grants U01CA217885 and R01HG009285. W.C.H. was supported by U01CA176058. C.G.B. and S.N.B. were supported by a Koch Institute Support Grant (P30-CA14051) from the National Cancer Institute (Swanson Biotechnology Center) and a Core Center Grant (P30-ES002109) from the National Institute of Environmental Health Sciences, and the Ludwig Center for Molecular Oncology. C.G.B. was supported by the National Science Foundation Graduate Research Fellowship Program. S.N.B. is a Howard Hughes Medical Institute Investigator. P.S.C. was supported by an NIH Pathway to Independence Award (K99 CA208028). E.M. would like to thank H. Horn for help with the expectation maximization algorithm. The authors thank the Koch Institute Swanson Biotechnology Center for technical support, specifically K. Cormier in the Hope Babette Tang Histology Facility.
Author information
Authors and Affiliations
Contributions
S.R.V. and M.M. conceived of the research and wrote the manuscript. S.R.V. and M.F.N. performed experiments. C.G.B. performed in vivo xenograft experiments. M.J.W., S.R.V., and P.S.C. performed data analysis on RNA sequencing data. J.M.K.-B. and S.R.V. performed data analysis on shRNA and CRISPR screening data. A.C.B., A.M.T., and J.S. performed copy number analysis on TCGA and cell line data. C.A.S. assisted in generation of shRNA reagents. P.T., A.D.C., and C.S.P. performed or oversaw data analysis. B.T., K.L., S.A.C., E.M., and M.S. performed mass spectrometry or were involved in downstream data analysis. S.N.B. oversaw in vivo xenograft experiments. F.V., A.T., and W.C.H. directed shRNA and CRISPR screening efforts. M.M. directed the overall project.
Corresponding author
Ethics declarations
Competing interests
A.C.B., A.D.C., C.A.S., and M.M. receive research support from Bayer Pharmaceuticals. M.M. is a scientific advisory board member of, consultant for, and holds equity in OrigiMed. The content of this manuscript is the subject of a pending patent application (S.V., M.M.).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 MAGOH and MAGOHB score as reciprocal paralog dependencies in both genome-scale CRISPR and shRNA screening datasets.
a, Plot of the percentage DNA sequence identity between gene pairs versus Pearson correlation for their co-expression across the CCLE cell lines (n = 93,433 gene pairs plotted). The solid line shows the mean; the shaded region shows the standard error. b, Correlation between MAOGHB expression (y axis) and MAGOH dependency in CRISPR data (x axis). Pearson’s r2 = 0.2235 (n = 336 cell lines plotted). c, Correlation between MAGOH gene dependencies in shRNA (y axis) and CRISPR (x axis) data. Pearson’s r2 = 0.1763 (n = 215 cell lines plotted). d, Correlation between MAGOH expression (y axis) and MAGOHB dependency in CRISPR data (x axis). Pearson’s r2 = 0.1113 (n = 336 cell lines plotted). e, Correlation between MAGOHB gene dependencies in shRNA (y axis) and CRISPR (x axis) data. Pearson’s r2 = 0.3068 (n = 215 cell lines plotted).
Supplementary Figure 2 Validation of reciprocal MAGOH/MAGOHB dependencies.
a, Validation of selective MAGOH and MAGOHB knockdown using doxycycline-inducible shRNAs against MAGOH or MAGOHB in Heya8 cells. Expression levels of each paralog are shown relative to the no-knockdown (–Dox) condition in each case. Data are presented as mean ± s.d., n = 4 technical replicates. b, Cell viability was measured in Heya8 cells (which do not have deletions in either MAGOH or MAGOHB) on knockdown of MAGOH, MAGOHB, or both. c, Cell viability measured on MAGOH knockdown (using Dox-inducible shRNA) in hemizygous MAGOHB-deleted Huh1 cells with or without reconstitution of MAGOHB-V5. d, Cell viability measured on MAGOH knockout (using sgRNA) in MAGOHB-deleted Huh1 cells with or without reconstitution of MAGOHB-V5. e, Validation of MAGOHB dependency in MAGOH-deleted H1437 cells using either control sgRNA (sgGFP) or two different sgRNAs against MAGOHB. For b–e, error bars show mean ± s.d., n = 3 replicates from a representative experiment repeated twice; P value by two-tailed, two-sample t test.
Supplementary Figure 3 MAGOH and MAGOHB dependencies are associated with deletion/low expression of the partner paralog.
Radial plots showing association of the top copy number alterations (top row) or gene expression changes (bottom row) correlated to MAGOH (left column) or MAGOHB (right column) dependency, among splicing and NMD-related genes. The radial plot depicts both the association of copy number/gene expression correlates with the target dependency profile (either MAGOH or MAGOHB dependency) and with each other. The radial distance of each feature (blue dot) from the dependency target (red) is inversely proportional to the information coefficient score between that feature and the target. The angular distances between features are based on the association matrix of information coefficient scores generated when comparing the top features with each other. If two features are close in angular distance, they have a higher information coefficient value with each other as compared to features that are father away in angular distance. MAGOH/MAGOHB dependencies are uniquely predicted by low expression or copy number loss of the partner paralog (circled in green).
Supplementary Figure 4 Additional validation of MAGOHB dependency in cells with hemizygous MAGOH loss.
a, Validation of MAGOHB knockdown with multiple MAGOHB shRNAs. H460 cells were transduced with lentivirus expressing either control shRNA (shGFP) or shRNAs against MAGOHB in the pLKO.1 vector. MAGOHB knockdown was assessed 72 h after selection using quantitative PCR. Expression levels are shown as normalized to control (shGFP) condition (dotted line). Data are presented as mean ± s.d., n = 4 technical replicates. b, Validation of selective MAGOHB dependency in MAGOH-deleted cells. Hemizygous MAGOH-deleted H460 cells or Kuramochi cells (no MAGOH deletion) were transduced with lentivirus encoding either control shRNA (shGFP) or shRNAs against MAGOHB in the pLKO.1 vector. Cell viability was assessed by CellTiterGlo 9 d after infection. Data are presented as mean ± s.d., n = 3 replicates (Kuramochi), n = 5 replicates (H460); P value by two-tailed two-sample t test. c, Quantification of colony formation assay (related to Fig. 1e). Following crystal violet staining, wells were destained and measurement of absorbance at 595 nm was performed for quantification of colony number. Data are presented as mean ± s.d., n = 3 replicates; P value by two-tailed, two-sample t test. d, Validation of ectopic MAGOH-V5 expression in ChagoK1 cells (related to Fig. 1f). Cells were transduced with lentivirus encoding V5-tagged MAGOH and western blotting was performed using an anti-MAGOH/B antibody. The experiment was repeated twice with similar results. e, Cell viability measured on MAGOHB knockdown in ChagoK1 cells with or without ectopic expression of untagged MAGOH. Data are presented as mean ± s.d., n = 6 replicates. Center wells analyzed in each condition; the experiment was repeated using two separate untagged MAGOH overexpression constructs with similar results. f, Validation of ectopic untagged MAGOH expression by western blotting in ChagoK1 cells. Results from a single experiment, repeated once in H1437 cells with similar results.
Supplementary Figure 5 Knockout of MAGOH confers MAGOHB dependency.
a, Validation of MAGOH-deleted clonal Heya8 cell lines. Heya8 cells, which are copy neutral for MAGOH, were transiently transfected with either a control sgRNA (sgGFP) or an sgRNA targeting MAGOH. Single-cell clones were derived and MAGOH expression was measured by quantitative PCR. Two single-cell clones (sgMAGOH_Clone 1 and sgMAGOH_Clone 5) demonstrated ablation of MAGOH expression as compared with control clones. Data are presented as relative MAGOH expression, normalized to expression in sgGFP_Clone 4. Points are shown from n = 2 technical replicates. b, Single-cell Heya8 clones were transduced with lentivirus encoding a doxycycline-inducible shRNA against MAGOHB. Cell viability was assessed 7 d after shRNA induction by CellTiterGlo. Data are presented as viability normalized to the –Dox condition for each cell line. Error bars show mean ± s.d., n = 3 replicates from a representative experiment repeated twice; P value by two-tailed, two-sample t test.
Supplementary Figure 6 Validation of sgRNAs used for MAGOH knockout in Heya8 cells.
a, Schematic showing the sgRNA target and PAM site, which lies near the intron 1–exon 2 junction of MAGOH. b, Sanger sequencing traces around the MAGOH sgRNA cut site, from single-cell Heya8 clones derived from cells transduced with either non-targeting guide (sgGFP) or guide against MAGOH. In both MAGOH-deleted clones (clones 1 and 5), an indel at the indicated position (black underline) occurs in a similar location and could cause a frameshift. In clone 5, mixed peaks may suggest distinct editing events on each allele. Ablation of MAGOH expression in these clones is shown in Supplementary Fig. 5.
Supplementary Figure 7 MAGOHB dependency in cells with chromosome 1p loss.
a, Distribution of MAGOHB CERES dependency scores in screened cell lines with chromosome 1p loss (–1, n = 46), 1p neutral status (0, n = 177), or 1p gain (+1, n = 14). 1p-deleted cell lines are significantly more dependent on MAGOHB than are 1p neutral cell lines; P value calculated by two-sample, one-sided Welch’s t test. Horizontal lines indicate means. b, Validation of selective MAGOHB dependency in 1p-deleted neuroblastoma cells. CHP212 (1p-deleted, left) or SKNDZ (1p-neutral, right) neuroblastoma cells were transduced with lentivirus expressing a doxycycline-inducible shRNA against MAGOHB and seeded for colony formation assay in either the absence of doxycycline or the presence of 100 ng/ml doxycycline. Colony formation was quantified by destaining of crystal violet and measurement of absorbance at 595 nm. Data are presented as mean ± s.d., n = 3 replicates; P value by two-tailed, two-sample t test.
Supplementary Figure 8 MAGOH dependency in cells with chromosome 12p loss.
a, Frequency of chromosome 12p and 1p loss across TCGA cohorts. b, Heat map of arm-level deletion events co-occurring with chromosome 1p loss across TCGA cohorts. Cohorts in which chromosome 1p co-deletion with chromosome 12p are significantly mutually exclusive (as compared with all possible 1p co-deletion events) are indicated by an asterisk. *P <0.05 by one-sided binomial test. Range of TCGA cohort sizes analyzed, n = 36 (CHOL) to n = 1,030 (BRCA). c, Distribution of MAGOH CERES dependency scores in screened cell lines with chromosome 12p loss (–1, n = 37), 12p neutral status (0, n = 169), or 12p gain (1, n = 69). 12p-deleted cell lines are significantly more dependent on MAGOH than are 12p-neutral cell lines. Horizontal lines indicate means; P value by Welch’s two-sample one-sided t test.
Supplementary Figure 9 RNA sequencing reveals multiple splicing events with potential effects on AS-NMD loops on MAGOHB knockdown in ChagoK1 cells, but not MAGOH-V5-reconstituted ChagoK1 cells.
a–c, Sashimi plots around ultraconserved elements reported to regulate AS-NMD loops in either the absence (gray) or presence (red) of MAGOHB knockdown in ChagoK1 cells (left panels) or MAGOH-V5-reconstituted ChagoK1 cells (right panels) for the genes HNRNPH1 (a), SRSF7 (b), and SRSF2 (c). Schematics depict splice-site usage events that result in the creation of premature termination codons. Numbers of junction-spanning reads averaged over three replicates for each condition are as indicated.
Supplementary Figure 10 Alterations in splice isoform usage on MAGOHB knockdown in a MAGOH-deleted context and predicted changes in protein levels of affected genes.
a,b, Isoform abundances (i) and predicted expression changes on the protein level (ii) for the genes PSPC1 (a) and HNRNPH1 (b) in ChagoK1 cells and MAGOH-reconstituted ChagoK1 cells with (red) or without (gray) MAGOHB knockdown. To predict effects on protein expression, isoforms were grouped by predicted protein-coding size as described in the Methods. Protein expression levels were assessed by western blotting (iii). a (iii), Western blotting for PSPC1 shows a dominant band presumed to correspond to the 523-aa/58-kDa predicted protein isoform. Western blots are from a representative experiment repeated twice. b (iii), Western blotting for HNRNPH1 shows a dominant truncated isoform for HNRNPH1 (~35 kDa) observed on MAGOHB knockdown in a MAGOH-deleted context that does not exactly correspond in size to what is expected based on RNA-sequencing data; this truncated species has not been fully characterized. Western blots are from a representative experiment repeated three times.
Supplementary Figure 11 MAGOH and MAGOHB dependencies are correlated with dependencies on other members of the MAGOH/MAGOHB interactomes.
a, Gene set enrichment analysis reveals enrichment of nonsense-mediated decay/exon junction complex gene sets among gene dependencies (n = 6,304) correlated with MAGOH copy number. Significance was assessed by running preranked GSEA using the gene dependency list as described (Methods). b, Immunoprecipitation–mass spectrometry was performed on MAGOH-V5- or MAGOHB-V5-expressing 293T or parental 293T cells (‘control’) using an anti-V5 antibody. Fold enrichment of interacting proteins over control for MAGOH-V5 (x axis) or MAGOHB-V5 (y axis) IPs is shown. Open circles indicate interactors with log2 (fold change) enrichment > 0.5 over control. Baits are not shown on the plot. c, Volcano plot for MAGOH-V5- (left) or MAGOHB-V5- (right) interacting proteins. The color scale depicts proteins in the following classes (from light pink to maroon): RNA binding, S ribosomal, EJC/NMD pathway member. Labeled proteins highlight selected EJC/NMD members found in the IPs. Significance was determined by moderated t test to calculate P values for the n = 1,995 genes from the IP, corrected for multiple hypotheses (FDR). d, MAGOH-V5 (left) or MAGOHB-V5 (right) interactomes overlaid with gene dependencies correlated with MAGOH (left) or MAGOHB (right) dependency. Dependency–dependency correlations with z scores >1.5 or <–1.5 are colored according to the scale. Gene dependencies correlated to MAGOH dependency (z > 1.5) are enriched among MAGOH-V5 interactors (log2 FC > 0.7) (P = 1.5 × 10–4) and gene dependencies correlated to MAGOHB dependency (z > 1.5) are enriched among MAGOHB-V5 interactors (log2 FC > 0.7) (P = 0.051). P values were calculated by one-sided Fisher exact test on the population of n = 380 genes (the overlap set of IP genes with log2 FC > 0.7 and the dependency data). Labels indicate proteins with z > 1.5 and log2 FC > 1 for the MAGOH-V5 plot and z > 1.5 and log2 FC > 0.5 for the MAGOHB-V5 plot.
Supplementary Figure 12 Additional validation of IPO13 dependency in MAGOH-deleted cells.
a, IPO13 knockdown was verified in H460 cells by quantitative PCR. H460 cells were transduced with lentivirus expressing either control shRNA (shGFP) or shRNAs against IPO13 in the pLKO.1 vector. IPO13 expression levels were assessed 72 h after selection using quantitative PCR. Expression levels are shown as normalized to control (shGFP) condition. Data are presented as mean ± s.d., n = 4 technical replicates. b, Quantification of the colony formation assay (related to Fig. 3d). Following crystal violet staining, wells were destained and measurement of absorbance at 595 nm was performed for quantification of colony number; data are normalized to absorbance in the shGFP condition. Data are presented as mean ± s.d., n = 3 replicates; P value by two-tailed two-sample t test. c, Clonogenic capacity was measured in MAGOH-deleted H1437 cells on IPO13 knockdown in either the absence (left) or presence (right) of MAGOH-V5 reconstitution. Destaining and measurement of absorbance at 595 nm was performed for quantification of colonies formed; data are normalized to absorbance in the shGFP condition. Data are presented as mean ± s.d., n = 3 replicates; P value by two-tailed two-sample t test. d, Cell viability was measured in MAGOH-deleted H1437 cells on IPO13 knockdown in either the absence (left) or presence (right) of MAGOH-V5 reconstitution. Data represent relative cell viability normalized to the shGFP condition. Data are presented as mean ± s.d., n = 3 replicates; P value by two-tailed two-sample t test.
Supplementary Figure 13 IPO13 knockdown in the setting of MAGOH deletion disrupts normal MAGOH/B localization and may interfere with EJC shuttling.
a, Representative fields of immunofluoresence staining of MAGOH/B in H460 cells in either the absence (top) or presence (bottom) of siRNA-mediated IPO13 knockdown. IPO13 knockdown in the setting of MAGOH deletion leads to decreased nuclear (black arrowhead) signal in favor of cytoplasmic (green arrowhead) signal. b, Quantification of fluorescence signal (nuclear/cytoplasmic ratio) in H460 cells transfected with either control siRNA or si-IPO13-2. Quantification was performed on three random fields per condition (n = 26 cells for control; n = 30 cells for si-IPO13-2) by ImageJ. Data are presented as mean ± s.d.; P value by two-tailed two-sample t test.
Supplementary Figure 14 H1437 xenografts can be targeted by TPNCs carrying siRNA against MAGOHB or IPO13.
a, Individual tumor volumes for the xenograft experiment shown in Fig. 4b (n = 6 tumors per condition). The solid line connects the mean tumor volume at each time point. b, Individual tumor volumes for the xenograft experiment shown in Fig. 4e (n = 10 tumors per condition). Solid line overlays show mean ± s.e.m. for each condition at each time point. c, Surface expression for p32 (receptor for LyP-1-containing nanocomplexes) in H1437 cells, as assessed by flow cytometry. Results are from single experiment. d, H1437 tumor xenograft experiment performed by intra-tumoral injection of LyP-1-containing nanocompexes carrying either siGFP or siMAGOHB cargo. Thin lines show tumor growth in individual mice (n = 10 tumors per group). Solid lines show mean ± s.e.m. for each condition. e, Representative staining for cleaved caspase-3 in an siGFP-treated mouse (top) and an siMAGOHB-treated mouse (bottom) using LyP-1-based tumor-penetrating nanocomplexes. Staining was performed on n = 6 tumors per group (randomly selected). f, Quantification of staining levels in six randomly selected tumors from each group. Data are presented as mean ± s.d.; P value by two-tailed, two-sample t test.
Supplementary Figure 15 A model for MAGOH/B and IPO13 dependency.
a,b, Cells without alterations in MAGOH/B and IPO13 have adequate MAGOH/B and IPO13 to tolerate partial inhibition of any of these factors. c, However, hemizygous loss of MAGOH and/or IPO13, as occurs in chromosome 1p loss, can sensitize cells to MAGOHB or IPO13 inhibition (top panel). An increased dependency on EJC function/shuttling may also occur in other states perhaps driven by an increased need for splicing or by somatic alterations in as-yet-unknown factors (bottom panel).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Note
Supplementary Table 1
Analysis of reciprocal paralog dependencies in genome-scale shRNA screening data
Supplementary Table 2
Analysis of reciprocal paralog dependencies in genome-scale CRISPR–Cas9 screening data
Supplementary Table 3
Gene ontology enrichment among genes found to be symmetric or asymmetric paralog dependencies in analysis of shRNA and CRISPR screening data
Supplementary Table 4
MAGOH hemizygous deletion status across CCLE cell lines used for PARIS analysis
Supplementary Table 5
Results of PARIS analysis to identify gene dependencies correlated with MAGOH deletion across CCLE cell lines
Supplementary Table 6
Significance of co-occurrence between chromosome 1p loss and hemizygous MAGOH deletion by TCGA tumor type
Supplementary Table 7
Downregulated protein-coding isoforms (b < –1) for genes that had concomitant upregulation of NMD isoforms (b > 1) upon MAGOHB knockdown in ChagoK1 cells
Supplementary Table 8
Gene ontology enrichment among genes displaying both an increase in expression level of NMD substrate isoform(s) and a proportional decrease in expression level of coding isoform(s) upon MAGOHB knockdown in ChagoK1 cells
Supplementary Table 9
Peptide report generated after IP–MS of MAGOH-V5 or MAGOHB-V5 and used as input into the expectation maximization algorithm
Supplementary Table 10
Results of the expectation maximization algorithm showing enriched interactors with MAGOH-V5
Supplementary Table 11
Results of the expectation maximization algorithm showing enriched interactors with MAGOHB-V5
Supplementary Table 12
Gene dependencies correlated to MAGOH or MAGOHB dependency
Supplementary Table 13
DEMETER score for MAGOH, MAGOHB, and IPO13 across screened cell lines as well as hemizygous deletion status (0, no deletion; 1, deletion) for each of these genes for each cell line
Supplementary Table 14
Oligonucleotide sequences used in experimental validation experiments (see Methods)
Rights and permissions
About this article
Cite this article
Viswanathan, S.R., Nogueira, M.F., Buss, C.G. et al. Genome-scale analysis identifies paralog lethality as a vulnerability of chromosome 1p loss in cancer. Nat Genet 50, 937–943 (2018). https://doi.org/10.1038/s41588-018-0155-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0155-3
This article is cited by
-
MAGOH promotes gastric cancer progression via hnRNPA1 expression inhibition-mediated RONΔ160/PI3K/AKT signaling pathway activation
Journal of Experimental & Clinical Cancer Research (2024)
-
Nuclear transport proteins: structure, function, and disease relevance
Signal Transduction and Targeted Therapy (2023)
-
Multifaceted roles of MAGOH Proteins
Molecular Biology Reports (2023)
-
Identification of key gene signatures for the overall survival of ovarian cancer
Journal of Ovarian Research (2022)
-
A pan-CRISPR analysis of mammalian cell specificity identifies ultra-compact sgRNA subsets for genome-scale experiments
Nature Communications (2022)