Abstract
We demonstrate that a Drosophila Golgi protein, Gorab, is present not only in the trans-Golgi but also in the centriole cartwheel where, complexed to Sas6, it is required for centriole duplication. In addition to centriole defects, flies lacking Gorab are uncoordinated due to defects in sensory cilia, which lose their nine-fold symmetry. We demonstrate the separation of centriole and Golgi functions of Drosophila Gorab in two ways: first, we have created Gorab variants that are unable to localize to trans-Golgi but can still rescue the centriole and cilia defects of gorab null flies; second, we show that expression of C-terminally tagged Gorab disrupts Golgi functions in cytokinesis of male meiosis, a dominant phenotype overcome by mutations preventing Golgi targeting. Our findings suggest that during animal evolution, a Golgi protein has arisen with a second, apparently independent, role in centriole duplication.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Gönczy, P. Towards a molecular architecture of centriole assembly. Nat. Rev. Mol. Cell Biol. 13, 425–435 (2012).
Goshima, G. et al. Genes required for mitotic spindle assembly in Drosophila S2 cells. Science 316, 417–421 (2007).
Dobbelaere, J. et al. A genome-wide RNAi screen to dissect centriole duplication and centrosome maturation in Drosophila. PLoS Biol. 6, e224 (2008).
Dzhindzhev, N. S. et al. Two-step phosphorylation of Ana2 by Plk4 is required for the sequential loading of Ana2 and Sas6 to initiate procentriole formation. Open Biol. 7, 170247 (2017).
Dzhindzhev, N. S. et al. Plk4 phosphorylates Ana2 to trigger Sas6 recruitment and procentriole formation. Curr. Biol. 24, 2526–2532 (2014).
Ohta, M. et al. Direct interaction of Plk4 with STIL ensures formation of a single procentriole per parental centriole. Nat. Commun. 5, 5267 (2014).
Hiraki, M., Nakazawa, Y., Kamiya, R. & Hirono, M. Bld10p constitutes the cartwheel-spoke tip and stabilizes the 9-fold symmetry of the centriole. Curr. Biol. 17, 1778–1783 (2007).
Jerka-Dziadosz, M. et al. Basal body duplication in Paramecium: the key role of Bld10 in assembly and stability of the cartwheel. Cytoskeleton https://doi.org/10.1002/cm.20433 (2010).
Lin, Y.-C. et al. Human microcephaly protein CEP135 binds to hSAS-6 and CPAP, and is required for centriole assembly. EMBO J. 32, 1141–1154 (2013).
Tang, C.-J. C., Fu, R.-H., Wu, K.-S., Hsu, W.-B. & Tang, T. K. CPAP is a cell-cycle regulated protein that controls centriole length. Nat. Cell Biol. 11, 825–831 (2009).
Schmidt, T. I. et al. Control of centriole length by CPAP and CP110. Curr. Biol. 19, 1005–1011 (2009).
Kohlmaier, G. et al. Overly long centrioles and defective cell division upon excess of the SAS-4-related protein CPAP. Curr. Biol. 19, 1012–1018 (2009).
Wang, W.-J., Soni, R. K., Uryu, K. & Tsou, M. F. The conversion of centrioles to centrosomes: essential coupling of duplication with segregation. J. Cell Biol. 193, 727–739 (2011).
Izquierdo, D., Wang, W.-J., Uryu, K. & Tsou, M.-F. B. Stabilization of cartwheel-less centrioles for duplication requires CEP295-mediated centriole-to-centrosome conversion. Cell Rep. 8, 957–965 (2014).
Fu, J. et al. Conserved molecular interactions in centriole-to-centrosome conversion. Nat. Cell Biol. 18, 87–99 (2016).
Efimov, A. et al. Asymmetric CLASP-dependent nucleation of noncentrosomal microtubules at the trans-Golgi network. Dev. Cell 12, 917–930 (2007).
Rivero, S., Cardenas, J., Bornens, M. & Rios, R. M. Microtubule nucleation at the cis-side of the Golgi apparatus requires AKAP450 and GM130. EMBO J. 28, 1016–1028 (2009).
Rios, R. M. The centrosome-Golgi apparatus nexus. Phil. Trans. R. Soc. Lond. B 369, 369 (2014).
Emmer, B. T., Maric, D. & Engman, D. M. Molecular mechanisms of protein and lipid targeting to ciliary membranes. J. Cell Sci. 123, 529–536 (2010).
Kim, H. et al. Ciliary membrane proteins traffic through the Golgi via a Rabep1/GGA1/Arl3-dependent mechanism. Nat. Commun. 5, 5482 (2014).
Stoetzel, C. et al. A mutation in VPS15 (PIK3R4) causes a ciliopathy and affects IFT20 release from the cis-Golgi. Nat. Commun. 7, 13586 (2016).
Follit, J. A. et al. The Golgin GMAP210/TRIP11 anchors IFT20 to the Golgi complex. PLoS Genet. 4, e1000315 (2008).
Broekhuis, J. R., Rademakers, S., Burghoorn, J. & Jansen, G. SQL-1, homologue of the Golgi protein GMAP210, modulates intraflagellar transport in C. elegans. J. Cell Sci. 126, 1785–1795 (2013).
Hilbert, M. et al. SAS-6 engineering reveals interdependence between cartwheel and microtubules in determining centriole architecture. Nat. Cell Biol. 18, 393–403 (2016).
Wang, W.-J. et al. De novo centriole formation in human cells is error-prone and does not require SAS-6 self-assembly. eLife 4, e10586 (2015).
Hennies, H. C. et al. Gerodermia osteodysplastica is caused by mutations in SCYL1BP1, a Rab-6 interacting golgin. Nat. Genet. 40, 1410–1412 (2008).
Di, Y. et al. Cloning and characterization of a novel gene which encodes a protein interacting with the mitosis-associated kinase-like protein NTKL. J. Hum. Genet. 48, 315–321 (2003).
Burman, J. L. et al. Scyl1, mutated in a recessive form of spinocerebellar neurodegeneration, regulates COPI-mediated retrograde traffic. J. Biol. Chem. 283, 22774–22786 (2008).
Burman, J. L., Hamlin, J. N. R. & McPherson, P. S. Scyl1 regulates Golgi morphology. PLoS One 5, e9537 (2010).
Egerer, J. et al. GORAB Missense Mutations Disrupt RAB6 and ARF5 Binding and Golgi Targeting. J. Invest. Dermatol. 135, 2368–2376 (2015).
Ripoche, J., Link, B., Yucel, J. K., Tokuyasu, K. & Malhotra, V. Location of Golgi membranes with reference to dividing nuclei in syncytial Drosophila embryos. Proc. Natl Acad. Sci. USA 91, 1878–1882 (1994).
Papoulas, O., Hays, T. S. & Sisson, J. C. The golgin Lava lamp mediates dynein-based Golgi movements during Drosophila cellularization. Nat. Cell Biol. 7, 612–618 (2005).
Frescas, D., Mavrakis, M., Lorenz, H., Delotto, R. & Lippincott-Schwartz, J. The secretory membrane system in the Drosophila syncytial blastoderm embryo exists as functionally compartmentalized units around individual nuclei. J. Cell Biol. 173, 219–230 (2006).
Blachon, S. et al. A proximal centriole-like structure is present in Drosophila spermatids and can serve as a model to study centriole duplication. Genetics 182, 133–144 (2009).
Basto, R. et al. Flies without centrioles. Cell 125, 1375–1386 (2006).
Gottardo, M. et al. Loss of centrobin enables daughter centrioles to form sensory cilia in Drosophila. Curr. Biol. 25, 2319–2324 (2015).
Rodrigues-Martins, A. et al. DSAS-6 organizes a tube-like centriole precursor, and its absence suggests modularity in centriole assembly. Curr. Biol. 17, 1465–1472 (2007).
Al-Dosari, M. & Alkuraya, F. S. A novel missense mutation in SCYL1BP1 produces geroderma osteodysplastica phenotype indistinguishable from that caused by nullimorphic mutations. Am. J. Med. Genet. A 149A, 2093–2098 (2009).
Gillingham, A. K. & Munro, S. Finding the Golgi: golgin coiled-coil proteins show the way. Trends Cell Biol. 26, 399–408 (2016).
Kitazawa, D., Yamaguchi, M., Mori, H. & Inoue, Y. H. COPI-mediated membrane trafficking is required for cytokinesis in Drosophila male meiotic divisions. J. Cell Sci. 125, 3649–3660 (2012).
Wong, M. & Munro, S. Membrane trafficking. The specificity of vesicle traffic to the Golgi is encoded in the golgin coiled-coil proteins. Science 346, 1256898 (2014).
Hilbert, M. et al. Caenorhabditis elegans centriolar protein SAS-6 forms a spiral that is consistent with imparting a ninefold symmetry. Proc. Natl Acad. Sci. USA 110, 11373–11378 (2013).
Liu, Y. et al. Gorab is required for dermal condensate cells to respond to Hedgehog signals during hair follicle morphogenesis. J. Invest. Dermatol. 136, 378–386 (2016).
Strnad, P. et al. Regulated HsSAS-6 levels ensure formation of a single procentriole per centriole during the centrosome duplication cycle. Dev. Cell 13, 203–213 (2007).
Fong, C. S., Kim, M., Yang, T. T., Liao, J.-C. & Tsou, M.-F. B. SAS-6 assembly templated by the lumen of cartwheel-less centrioles precedes centriole duplication. Dev. Cell 30, 238–245 (2014).
Lambrus, B. G. et al. p53 protects against genome instability following centriole duplication failure. J. Cell Biol. 210, 63–77 (2015).
Lambrus, B. G. et al. A USP28–53BP1-p53-p21 signaling axis arrests growth after centrosome loss or prolonged mitosis. J. Cell Biol. 214, 143–153 (2016).
Raj, K., Ogston, P. & Beard, P. Virus-mediated killing of cells that lack p53 activity. Nature 412, 914–917 (2001).
D’Avino, P. P. et al. Isolation of protein complexes involved in mitosis and cytokinesis from Drosophila cultured cells. Methods Mol. Biol. 545, 99–112 (2009).
Lipinszki, Z. et al. Affinity purification of protein complexes from Drosophila embryos in cell cycle studies. Methods Mol. Biol. 1170, 571–588 (2014).
Fu, J. & Glover, D. M. Structured illumination of the interface between centriole and peri-centriolar material. Open Biol. 2, 120104 (2012).
Port, F., Chen, H.-M., Lee, T. & Bullock, S. L. Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proc. Natl Acad. Sci. USA 111, E2967–E2976 (2014).
Groth, A. C., Fish, M., Nusse, R. & Calos, M. P. Construction of transgenic Drosophila by using the site-specific integrase from phage phiC31. Genetics 166, 1775–1782 (2004).
Bischof, J., Maeda, R. K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific phiC31 integrases. Proc. Natl Acad. Sci. USA 104, 3312–3317 (2007).
Riedel, F., Gillingham, A. K., Rosa-Ferreira, C., Galindo, A. & Munro, S. An antibody toolkit for the study of membrane traffic in Drosophila melanogaster. Biol. Open 5, 987–992 (2016).
Heuer, J. G., Li, K. & Kaufman, T. C. The Drosophila homeotic target gene centrosomin (cnn) encodes a novel centrosomal protein with leucine zippers and maps to a genomic region required for midgut morphogenesis. Development 121, 3861–3876 (1995).
Dzhindzhev, N. S. et al. Asterless is a scaffold for the onset of centriole assembly. Nature 467, 714–718 (2010).
Sechi, S. et al. Rab1 interacts with GOLPH3 and controls Golgi structure and contractile ring constriction during cytokinesis in Drosophila melanogaster. Open Biol. 7, 160257 (2017).
Zhong, L. & Belote, J. M. The testis-specific proteasome subunit Prosalpha6T of D. melanogaster is required for individualization and nuclear maturation during spermatogenesis. Development 134, 3517–3525 (2007).
Chen, J. V. et al. Rootletin organizes the ciliary rootlet to achieve neuron sensory function in Drosophila. J. Cell Biol. 211, 435–453 (2015).
Acknowledgements
We acknowledge P. Deák (Department of Genetics, University of Szeged, Szeged, Hungary) for his encouragement and support of L.K. and M. Pál (Department of Genetics, University of Szeged, Szeged, Hungary) for injection of CRISPR–Cas9 guide-RNAs for Gorab mutagenesis; and thank S. Chang, K. Oras, and A. Madich (Deparment of Genetics, University of Cambridge, Cambridge, UK) for injection of Gorab transgenes. We also greatly appreciate the advice of M. Richter and A. Fatalska in studies of protein–protein interactions. We thank T. Megraw (Department of Biomedical Sciences, Florida State University, Tallahassee, FL, USA) for GFP-rootletin flies, C. Gonzalez (Institute for Research in Biomedicine, The Barcelona Institute of Science and Technology, Barcelona, Spain) for YFP-centrobin flies, S. Munro (MRC Laboratory of Molecular Biology, Cambridge, UK) for anti-golgin antibodies, K. Raj (Radiation Effects Department, Public Health England, Didcot, UK) for the U2OSp53DD cell line, and J. Debski for advice in mass-spectrometry. D.M.G. is grateful for a Wellcome Investigator Award, which supported this work. The study was initiated with support from Cancer Research UK.
Author information
Authors and Affiliations
Contributions
L.K. contributed to planning experiments, fluorescence microscopy, Drosophila genetics, super-resolution microscopy, cell culture work, data analysis, and writing the manuscript; J.C.-C. contributed to antibody generation, mass spectrometry, Drosophila genetics, and fluorescence microscopy; S.S. contributed to mass spectrometry, in vitro interaction studies, protein structure analysis, and cell-culture work; M.G. contributed to electron microscopy; G.T. contributed to super-resolution microscopy; N.S.D. contributed to planning experiments and data analysis; M.G.R. contributed to electron microscopy; G.C. contributed to electron microscopy; and D.M.G. contributed to the conception and supervision of the study, planning experiments, and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Localization of Gorab in mitotic cells.
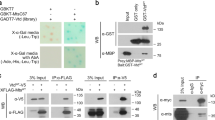
(a) D-Mel2 cells in metaphase were immunostained to reveal Gorab, dPLP (centrosome), and Golgin245 (trans-Golgi). Arrowheads indicate centrosomes. Experiment repeated 3 times with similar results.Main scale bar, 5 µm; inset scale bar, 0.5 µm.(b) Wing discs from larvae expressing N-terminally GFP-tagged Gorab were immunostained for dPLP (red) and Golgin245 (grey). Dashed lines highlight an interphase (left) and a mitotic cell (right). Arrowheads indicate centrosomes of mitotic cells. Experiement repeated 3 times with similar results.Main scale bar,5 µm.Inset scale bar,0.5µm.(c) Brains from larvae expressing N-terminally GFP-tagged Gorab immunostained for dPLP (red) and Golgin245 (grey). Dashed lines highlight an interphase (left) and a mitotic cell (right). Arrowheads indicate centrosomes of a mitotic neuroblast. Experiment repeated with similar result. Main scale bar,5 µm, Inset scale bar, 1µm. (d) Fluorescent micrographs of control and Gorab depleted D-Mel2 cells stained by DAPI (blue) and anti-Gorab (green) to demonstrate the specificity of Gorab antibody. Cells subjected to 4 times 4 days siRNA treatment with control or Gorab specific siRNAs followed by fixation in methanol and immunostaning. Experiment independently repeated twice with similar results. Scale bar: 10 µm.
Supplementary Figure 2 Gorab interaction with Sas6.
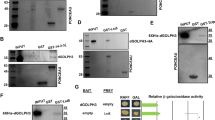
Autoradiograms of GST-tagged Gorab’s interaction with the 35S-Methionine-labelled Sas6 protein fragments schematically illustrated in Fig 2b. The length of the Sas6 fragments are indicated. Experiments were repeated twice by different investigators with similar results.
Supplementary Figure 3 Sas6 interaction with Gorab.
(a)Autoradiogram of GST-Sas6 interaction with 35S-Methionine-labelled Gorab protein fragments, schematically illustrated in Fig 5b. The experiments were repeated once by a different investigator with similar result. (b) Sas6 and Gorab interaction in vivo. D-Mel2 cells were transiently transfected with myc-tagged Sas6 and GFP-tagged wild type or ΔSID Gorab. Samples were subjected to GFP-trap immunoprecipitation and western blot using anti-myc and anti-GFP antibodies. Experiments were repeated once by a different investigator with similar result. Anti-alpha tubulin is loading control.
Supplementary Figure 4 Localization of V266P mutant Gorab to basal bodies.
Ciliated neurons from femoral chordotonal organs of flies expressing GFP-gorabV266P in a gorab1 mutant background immunostained with dPLP to reveal basal bodies and with phalloidin to visualize actin in scolopale rods. Experiment was repeated once with similar result. Arrowheads, GFP-Gorab at basal bodies. Scale bar, 10 µm.
Supplementary Figure 5 C-terminally GFP-tagged wild-type Gorab rescues gorab1 mutant phenotypes.
(a) Fertility of wild type, gorab1mutant and rescued females. Wild type C-terminally GFP tagged gorab cDNA was expressed under the control of the ubiqutin promoter in gorab1 to rescue (Ubq-gorabwt-GFP, gorab1). Individual females were mated with wild type males and allowed to lay eggs at 25 °C for 6 days.Data points represent the number of progeny of individual females. Means ± s.e.m are shown for n=15 females per genotype. p values of two tailed unpaired t-tests are shown. p value in blue indicates significant difference (99% confidence interval). Experiment repeated once with similar result.(b) Climbing ability of wild type, gorab1mutant and rescued flies. Wild type C-terminally GFP tagged gorab cDNA was expressed under the control of the ubiqutin promoter in gorab1 to give rescue (Ubq-gorabwt-GFP, gorab1). Flies were raised at 29 °C and subjected to a climbing assay. Each data point represents the percentage of flies from a group of 15 flies. Means ± s.e.m are shown for N=3 independent experiments, n= 15 flies/genotype were investigated in each experiment. p values of two tailed unpaired t-tests are shown. p value in blue indicates significant difference (99% confidence interval).
Supplementary Figure 6 Centrosomal localization of human GORAB.
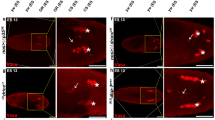
(a) Centrosomal localization of wild type and A220P mutant GORAB. U-2 OS cells transiently transfected with constructs constitutively expressing N terminally GFP-tagged wild type (left panel) or A220P mutant human GORAB. Pericentrin (PCNT, red) stains centrosomes, Golgin-97 (grey) highlights trans-Golgi. Main scale bar,2 µm. Inset scale bar, 0.2 µm. Experiments repeated twice with similar results.(b)GORAB localization inside the centrioles.3D-SIM micrographs of U-2 OS centrosomes immunostained for Pericentrin (blue), SAS6 (red) and GORAB (green). Centrosomes of top (upper panel) and side view (lower panel) are not identical. Experiments repeated twice with similar results. Asterisk, site of procentriole formation where SAS6 is recruited. Arrowheads indicate SAS6, which is initially recruited to the proximal lumen of the cartwheel-less mother centriole and which moves to the site of procentriole formation45. Scale bar, 200 nm. (c) Synergistic effect of SAS6 and GORAB on loss of centrosomes. U-2OSp53DDcells subjected to 3 times 72h siRNA treatments against the indicated genes. Fixed cells were immunostained with a combination of CENP-J and gamma-Tubulin antibodies to reveal centrioles and centrosomes..Each data point represent the percentage of cells with the given centrosome number from an independent experiment (n=100 cells). Means ± s.e.m are shown for N=4 independent experiments (n=100 cells/experiment)p values of two tailed unpaired t-tests are shown(99% confidence interval). (d)Representative images of U-2OS cells subjected to siRNAs against GFP (control) or GORAB and treated with a combination of 4 µM aphidicolin and 1.5 mM hydroxyurea (A/HU). Anti-gamma-tubulin immunostaning reveals centrosomes (red) and DAPI staining (blue) reveals nuclei. Experiment repeated twice with similar results. Scale bar, 5 µm. (e) Quantification of centrosome numbers in U-2OS cell shown on d. Each data point represents the percentage of cells with a given number from an independent experiment (n=100 cells). Mean ± s.e.m are shown for N=3 independent experiments (n=100 cells/experiment).p values of two tailed unpaired t-tests are shown (99% confidence interval). (f) Anti-Gorab Western blot on GORAB depleted U-2 OSp53DD cell lysates. Cells were subjected to3 siRNA treatments, each for 72h, with control or GORAB specific siRNAs to check the specificity of the GORAB antibody (Atlas, #HPA027250). Subsequent Coomassie staining of the membrane (CBB) proves a loading control. Experiment independently repeated once with similar results.(g) Fluorescent micrographs of control and GORAB-depleted U-2 OSp53DD cells stained with DAPI (blue) and anti-GORAB (green) to demonstrate the specificity of GORAB antibody (Atlas, #HPA027250). Cells were subjected to3 siRNA treatments, each for 72h, with control or GORAB specific siRNAs followed by fixation in 4% formaldehyde and immunostaning. Experiment independently repeated once with similar results. Scale bar,5 µm.
Supplementary information
Supplementary Figures
Supplementary Figures 1–6
Supplementary Tables
Supplementary Tables 1–3
Supplementary Movie 1
Startle response assay of gorab1 mutant. Flies were placed in 3cm diameter Petri dishes and left for 1 h to acclimatize. Movies were recorded using a LifeCam Cinema HD (Microsoft) web camera
Rights and permissions
About this article
Cite this article
Kovacs, L., Chao-Chu, J., Schneider, S. et al. Gorab is a Golgi protein required for structure and duplication of Drosophila centrioles. Nat Genet 50, 1021–1031 (2018). https://doi.org/10.1038/s41588-018-0149-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0149-1
This article is cited by
-
New insights into the role of the Golgi apparatus in the pathogenesis and therapeutics of human diseases
Archives of Pharmacal Research (2022)
-
GORAB scaffolds COPI at the trans-Golgi for efficient enzyme recycling and correct protein glycosylation
Nature Communications (2019)