Abstract
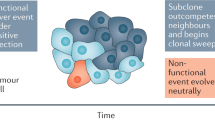
A major challenge to personalized oncology is that driver mutations vary among cancer cells inhabiting the same tumor. Whether this reflects principally disparate patterns of Darwinian evolution in different tumor regions has remained unexplored1,2,3,4,5. We mapped the prevalence of genetically distinct clones over 250 regions in 54 childhood cancers. This showed that primary tumors can simultaneously follow up to four evolutionary trajectories over different anatomic areas. The most common pattern consists of subclones with very few mutations confined to a single tumor region. The second most common is a stable coexistence, over vast areas, of clones characterized by changes in chromosome numbers. This is contrasted by a third, less frequent, pattern where a clone with driver mutations or structural chromosome rearrangements emerges through a clonal sweep to dominate an anatomical region. The fourth and rarest pattern is the local emergence of a myriad of clones with TP53 inactivation. Death from disease was limited to tumors exhibiting the two last, most dynamic patterns.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mengelbier, L. H. et al. Intratumoral genome diversity parallels progression and predicts outcome in pediatric cancer. Nat. Commun. 6, 6125 (2015).
Eleveld, T. F. et al. Relapsed neuroblastomas show frequent RAS-MAPK pathway mutations. Nat. Genet. 47, 864–871 (2015).
Cresswell, G. D. et al. Intra-tumor genetic heterogeneity in Wilms tumor: clonal evolution and clinical implications. EBioMedicine 9, 120–129 (2016).
Schramm, A. et al. Mutational dynamics between primary and relapse neuroblastomas. Nat. Genet. 47, 872–877 (2015).
Padovan-Merhar, O. M. et al. Enrichment of targetable mutations in the relapsed neuroblastoma genome. PLoS Genet. 12, e1006501 (2016).
McGranahan, N. & Swanton, C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168, 613–628 (2017).
Gillies, R. J., Verduzco, D. & Gatenby, R. A. Evolutionary dynamics of carcinogenesis and why targeted therapy does not work. Nat. Rev. Cancer 12, 487–493 (2012).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Notta, F. et al. A renewed model of pancreatic cancer evolution based on genomic rearrangement patterns. Nature 538, 378–382 (2016).
Kovac, M. et al. Recurrent chromosomal gains and heterogeneous driver mutations characterise papillary renal cancer evolution. Nat. Commun. 6, 6336 (2015).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015).
Williams, M. J., Werner, B., Barnes, C. P., Graham, T. A. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48, 238–244 (2016).
Alves, J. M., Prieto, T. & Posada, D. Multiregional tumor trees are not phylogenies. Trends Cancer 3, 546–550 (2017).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Molenaar, J. J. et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature 483, 589–593 (2012).
Walz, A. L. et al. Recurrent DGCR8, DROSHA, and SIX homeodomain mutations in favorable histology Wilms tumors. Cancer Cell 27, 286–297 (2015).
Wegert, J. et al. Mutations in the SIX1/2 pathway and the DROSHA/DGCR8 miRNA microprocessor complex underlie high-risk blastemal type Wilms tumors. Cancer Cell 27, 298–311 (2015).
Kohsaka, S. et al. A recurrent neomorphic mutation in MYOD1 defines a clinically aggressive subset of embryonal rhabdomyosarcoma associated with PI3K-AKT pathway mutations. Nat. Genet. 46, 595–600 (2014).
Staaf, J. et al. Segmentation-based detection of allelic imbalance and loss-of-heterozygosity in cancer cells using whole genome SNP arrays. Genome Biol. 9, R136 (2008).
Gisselsson, D. et al. Generation of trisomies in cancer cells by multipolar mitosis and incomplete cytokinesis. Proc. Natl Acad. Sci. USA 107, 20489–20493 (2010).
Vujanic, G. M. et al. Revised International Society of Paediatric Oncology (SIOP) working classification of renal tumors of childhood. Med. Pediatr. Oncol. 38, 79–82 (2002).
Chagtai, T. et al. Gain of 1q as a prognostic biomarker in Wilms tumors (WTs) treated with preoperative chemotherapy in the International Society of Paediatric Oncology (SIOP) WT 2001 Trial: a SIOP Renal Tumours Biology Consortium Study. J. Clin. Oncol. 34, 3195–3203 (2016).
Gratias, E. J. et al. Association of chromosome 1q gain with inferior survival in favorable-histology Wilms tumor: a report from the Children’s Oncology Group. J. Clin. Oncol. 34, 3189–3194 (2016).
Ooms, A. H. et al. Significance of TP53 mutation in Wilms tumors with diffuse anaplasia: a report from the Children’s Oncology Group. Clin. Cancer Res. 22, 5582–5591 (2016).
Caren, H. et al. High-risk neuroblastoma tumors with 11q-deletion display a poor prognostic, chromosome instability phenotype with later onset. Proc. Natl Acad. Sci. USA 107, 4323–4328 (2010).
Caron, H. et al. Allelic loss of chromosome 1p as a predictor of unfavorable outcome in patients with neuroblastoma. N. Engl. J. Med. 334, 225–230 (1996).
Brodeur, G. M., Seeger, R. C., Schwab, M., Varmus, H. E. & Bishop, J. M. Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science 224, 1121–1124 (1984).
Marusyk, A. et al. Non-cell-autonomous driving of tumour growth supports sub-clonal heterogeneity. Nature 514, 54–58 (2014).
Baker, A. M. et al. Robust RNA-based in situ mutation detection delineates colorectal cancer subclonal evolution. Nat. Commun. 8, 1998 (2017).
Mayrhofer, M., Viklund, B. & Isaksson, A. Rawcopy: Improved copy number analysis with Affymetrix arrays. Sci. Rep. 6, 36158 (2016).
Rasmussen, M. et al. Allele-specific copy number analysis of tumor samples with aneuploidy and tumor heterogeneity. Genome Biol. 12, R108 (2011).
Koster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv, https://arxiv.org/abs/1303.3997v2 (2013).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Narzisi, G. et al. Accurate de novo and transmitted indel detection in exome-capture data using microassembly. Nat. Methods 11, 1033–1036 (2014).
Cibulskis, K. et al. ContEst: estimating cross-contamination of human samples in next-generation sequencing data. Bioinformatics 27, 2601–2602 (2011).
Okonechnikov, K., Conesa, A. & Garcia-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Ewels, P., Magnusson, M., Lundin, S. & Kaller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Karlsson, A. et al. Mutational and gene fusion analyses of primary large cell and large cell neuroendocrine lung cancer. Oncotarget 6, 22028–22037 (2015).
Lindquist, K. E. et al. Clinical framework for next generation sequencing based analysis of treatment predictive mutations and multiplexed gene fusion detection in non-small cell lung cancer. Oncotarget 8, 34796–34810 (2017).
Gao, R. et al. Punctuated copy number evolution and clonal stasis in triple-negative breast cancer. Nat. Genet. 48, 1119–1130 (2016).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Barde, I., Salmon, P. & Trono, D. Production and titration of lentiviral vectors. Curr. Protoc. Neurosci. 53, 4.21.1–4.21.23 (2010).
Kabadi, A. M., Ousterout, D. G., Hilton, I. B. & Gersbach, C. A. Multiplex CRISPR/Cas9-based genome engineering from a single lentiviral vector. Nucleic Acids Res. 42, e147 (2014).
Holm, A., Baldetorp, B., Olde, B., Leeb-Lundberg, L. M. & Nilsson, B. O. The GPER1 agonist G-1 attenuates endothelial cell proliferation by inhibiting DNA synthesis and accumulating cells in the S and G2 phases of the cell cycle. J. Vasc. Res. 48, 327–335 (2011).
Gisselsson, D. Classification of chromosome segregation errors in cancer. Chromosoma 117, 511–519 (2008).
Gisselsson, D. et al. When the genome plays dice: circumvention of the spindle assembly checkpoint and near-random chromosome segregation in multipolar cancer cell mitoses. PLoS One 3, e1871 (2008).
Mengelbier, L. H. et al. Deletions of 16q in Wilms tumors localize to blastemal-anaplastic cells and are associated with reduced expression of the IRXB renal tubulogenesis gene cluster. Am. J. Pathol. 177, 2609–2621 (2010).
Cohn, S. L. et al. The International Neuroblastoma Risk Group (INRG) classification system: an INRG Task Force report. J. Clin. Oncol. 27, 289–297 (2009).
Dasgupta, R., Fuchs, J. & Rodeberg, D. Rhabdomyosarcoma. Semin. Pediatr. Surg. 25, 276–283 (2016).
Acknowledgements
This study was supported by grants to D.G. from the Swedish Research Foundation (2016-01022), the Swedish Cancer Society (CAN2015/284), the Swedish Childhood Cancer Foundation (PR2016-024, NCP2015-0035), the Crafoord Foundation, the Royal Physiographic Society and the Medical Faculty of Lund University Sweden. We also acknowledge technical support from the Science for Life Laboratory, the Knut and Alice Wallenberg Foundation, the National Genomics Infrastructure founded by the Swedish Research Council and Uppsala Multidisciplinary Center for Advanced Computational Science for assistance with massively parallel sequencing and access to the UPPMAX computational infrastructure. We also thank the Swegene Centre for Integrative Biology at Lund University for assistance.
Author information
Authors and Affiliations
Contributions
J.K., A.V. and D.G. conceived and designed the project. J.K., A.V., L.H.M., J.S., B.V., T.J. and A.I. coordinated and analyzed whole-genome genotyping and sequencing data, while A.V. performed phylogenetic analysis. S.B., I.Ø., A.B., T.B., B.S., N.P., B.G.L., N.B., D.B. and D.G. performed clinical correlation studies and contributed tumor material. L.C. prepared knockout cell lines. C.J. and A.W. performed technical work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19 and Supplementary Tables 1 and 2
Supplementary Data 1
Overview of patients and analytical platforms
Supplementary Data 2
Segmented allelic imbalances
Supplementary Data 3
Mutations detected by whole-exome sequencing
Supplementary Data 4
Variants identified with deep resequencing
Supplementary Data 5
Analysis performed on each sample
Supplementary Data 6
Genome profiles from whole-genome genotyping
Rights and permissions
About this article
Cite this article
Karlsson, J., Valind, A., Holmquist Mengelbier, L. et al. Four evolutionary trajectories underlie genetic intratumoral variation in childhood cancer. Nat Genet 50, 944–950 (2018). https://doi.org/10.1038/s41588-018-0131-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0131-y
This article is cited by
-
Hallmark discoveries in the biology of Wilms tumour
Nature Reviews Urology (2024)
-
Spatial biology of cancer evolution
Nature Reviews Genetics (2023)
-
Neuroblastoma arises in early fetal development and its evolutionary duration predicts outcome
Nature Genetics (2023)
-
Clonal evolution during metastatic spread in high-risk neuroblastoma
Nature Genetics (2023)
-
Gene expression-based dissection of inter-histotypes, intra-histotype and intra-tumor heterogeneity in pediatric tumors
Scientific Reports (2022)