Abstract
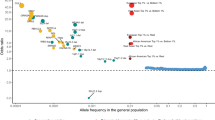
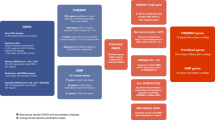
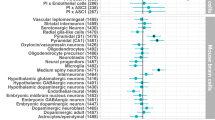
With few exceptions, the marked advances in knowledge about the genetic basis of schizophrenia have not converged on findings that can be confidently used for precise experimental modeling. By applying knowledge of the cellular taxonomy of the brain from single-cell RNA sequencing, we evaluated whether the genomic loci implicated in schizophrenia map onto specific brain cell types. We found that the common-variant genomic results consistently mapped to pyramidal cells, medium spiny neurons (MSNs) and certain interneurons, but far less consistently to embryonic, progenitor or glial cells. These enrichments were due to sets of genes that were specifically expressed in each of these cell types. We also found that many of the diverse gene sets previously associated with schizophrenia (genes involved in synaptic function, those encoding mRNAs that interact with FMRP, antipsychotic targets, etc.) generally implicated the same brain cell types. Our results suggest a parsimonious explanation: the common-variant genetic results for schizophrenia point at a limited set of neurons, and the gene sets point to the same cells. The genetic risk associated with MSNs did not overlap with that of glutamatergic pyramidal cells and interneurons, suggesting that different cell types have biologically distinct roles in schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sullivan, P. F., Daly, M. J. & O’Donovan, M. Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat. Rev. Genet. 13, 537–551 (2012).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Genovese, G. et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat. Neurosci. 19, 1433–1441 (2016).
Singh, T. et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat. Neurosci. 19, 571–577 (2016).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Marshall, C. R. et al. Contribution of copy-number variants to schizophrenia from a genome-wide study of 41,321 subjects. Nat. Genet. 49, 27–35 (2016).
Finucane, H. K. et al. Partitioning heritability by functional category using GWAS summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Lips, E. S. et al. Functional gene group analysis identifies synaptic gene groups as risk factor for schizophrenia. Mol. Psychiatry 17, 996–1006 (2012).
Darnell, J. C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Goudriaan, A. et al. Specific glial functions contribute to schizophrenia susceptibility. Schizophr. Bull. 40, 925–935 (2014).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Pers, T. H. et al. Comprehensive analysis of schizophrenia-associated loci highlights ion channel pathways and biologically plausible candidate causal genes. Hum. Mol. Genet. 25, 1247–1254 (2016).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Romanov, R. A. et al. Molecular interrogation of hypothalamic organization reveals distinct dopamine neuronal subtypes. Nat. Neurosci. 20, 176–188 (2017).
La Manno, G. et al. Molecular diversity of midbrain development in mouse, human and stem cells. Cell 167, 566–580 (2016).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
GTEx Consortium. The Genotype–Tissue Expression (GTEx) pilot analysis: multi-tissue gene regulation in humans. Science 348, 648–660 (2015).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Gokce, O. et al. Cellular taxonomy of the mouse striatum as revealed by single-cell RNA-seq. Cell Rep. 16, 1126–1137 (2016).
Habib, N. et al. Div-Seq: single-nucleus RNA-seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single-cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Abdelmoez, M.N. et al. Correlation of gene expressions between nucleus and cytoplasm reflects single-cell physiology. Preprint at bioRxiv https://www.biorxiv.org/content/early/2017/10/20/206672 (2017).
Cajigas, I. J. et al. The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 74, 453–466 (2012).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single-cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Skene, N. G. & Grant, S. G. Identification of vulnerable cell types in major brain disorders using single-cell transcriptomes and expression-weighted cell-type enrichment. Front. Neurosci. 10, 16 (2016).
Gaspar, H. A. & Breen, G. Drug enrichment and discovery from schizophrenia genome-wide association results: an analysis and visualisation approach. Sci. Rep. 7, 12460 (2017).
Anttila, V. et al. Analysis of shared heritability in common disorders of the brain. Science (in the press).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Bertram, L., McQueen, M. B., Mullin, K., Blacker, D. & Tanzi, R. E. Systematic meta-analyses of Alzheimer disease genetic association studies: the AlzGene database. Nat. Genet. 39, 17–23 (2007).
Patsopoulos, N. et al. The Multiple Sclerosis Genomic Map: role of peripheral immune cells and resident microglia in susceptibility. Preprint at bioRxiv https://www.biorxiv.org/content/early/2017/07/13/143933 (2017).
Yang, H., Robinson, P. N. & Wang, K. Phenolyzer: phenotype-based prioritization of candidate genes for human diseases. Nat. Methods 12, 841–843 (2015).
Burow, D. A. et al. Dynamic regulation of mRNA decay during neural development. Neural Dev. 10, 11 (2015).
Akbarian, S. et al. The PsychENCODE project. Nat. Neurosci. 18, 1707–1712 (2015).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
de Leeuw, C. A., Neale, B. M., Heskes, T. & Posthuma, D. The statistical properties of gene-set analysis. Nat. Rev. Genet. 17, 353–364 (2016).
Brown, M. B. A method for combining non-independent, one-sided tests of significance. Biometrics 31, 987–992 (1975).
Lun, A. T., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res 5, 2122 (2016).
Vu, T. N. et al. Beta-Poisson model for single-cell RNA-seq data analyses. Bioinformatics 32, 2128–2135 (2016).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Sniekers, S. et al. Genome-wide association meta-analysis of 78,308 individuals identifies new loci and genes influencing human intelligence. Nat. Genet. 49, 1107–1112 (2017).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014).
Wood, A. R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Paternoster, R., Brame, R., Mazerolle, P. & Piquero, A. Using the correct statistical test for the equality of regression coefficients. Criminology 36, 859–866 (1998).
Wagnon, J. L. et al. CELF4 regulates translation and local abundance of a vast set of mRNAs, including genes associated with regulation of synaptic function. PLoS Genet. 8, e1003067 (2012).
Collins, M. O. et al. Molecular characterization and comparison of the components and multiprotein complexes in the postsynaptic proteome. J. Neurochem. 97, 16–23 (2006). (Suppl. 1).
Bayés, A. et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nat. Neurosci. 14, 19–21 (2011).
Fernández, E. et al. Targeted tandem affinity purification of PSD95 recovers core postsynaptic complexes and schizophrenia susceptibility proteins. Mol. Syst. Biol. 5, 269 (2009).
Weyn-Vanhentenryck, S. M. et al. HITS–CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Rep. 6, 1139–1152 (2014).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Brozzi, A., Urbanelli, L., Germain, P. L., Magini, A. & Emiliani, C. hLGDB: a database of human lysosomal genes and their regulation. Database 2013, bat024 (2013).
Calvo, S. E., Clauser, K. R. & Mootha, V. K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, D1251–D1257 (2016). (D1).
Shigeoka, T. et al. Dynamic axonal translation in developing and mature visual circuits. Cell 166, 181–192 (2016).
Boyken, J. et al. Molecular profiling of synaptic vesicle docking sites reveals novel proteins but few differences between glutamatergic and GABAergic synapses. Neuron 78, 285–297 (2013).
Takamori, S. et al. Molecular anatomy of a trafficking organelle. Cell 127, 831–846 (2006).
Acknowledgements
J.H.-L. was funded by the Swedish Research Council (Vetenskapsrådet, award 2014-3863), StratNeuro, the Wellcome Trust (108726/Z/15/Z) and the Swedish Brain Foundation (Hjärnfonden). P.F.S. gratefully acknowledges support from the Swedish Research Council (Vetenskapsrådet, award D0886501). N.G.S. was supported by the Wellcome Trust (108726/Z/15/Z). J.B. was supported by the Swiss National Science Foundation. The PGC has received major funding from the US National Institute of Mental Health (U01 MH109528 and U01 MH109532). H.A.G. was supported by PGC grant 1U01MH109514-01. Primary schizophrenia GWAS data were generated with support from the Medical Research Council (MRC) Centre (MR/L010305/1), program grant G0800509 and project grant MR/L011794/1, and funding from the European Union’s Seventh Framework Programme for research, technological development and demonstration under grant agreement no. 279227 (CRESTAR Consortium).
Author information
Authors and Affiliations
Consortia
Contributions
N.G.S., J.B., P.F.S. and J.H.-L. designed the study and wrote and reviewed the manuscript; N.G.S. performed the LDSC analyses; J.B. performed the MAGMA analyses; T.E.B., R.D.H., J.A.M. and E.S.L. generated the human mid-temporal cortex data; A.B.M.-M., J.R., S.L. and J.H.-L. generated the KI single-cell data; the Major Depressive Disorder Working Group of the PGC performed the MDD GWAS; J.T.R.W., J.J.C., P.G.-R., M.C.O., M.J.O. and A.F.P. performed the schizophrenia CLOZUK GWAS; G.B. and H.A.G. analyzed the antipsychotic drug targets; and all authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
P.F.S. is on the advisory committee at Lundbeck, is a Scientific Advisory Board member at Pfizer and has received speaker reimbursement and grant funding from Roche. J.H.-L. is a Scientific Advisor at Cartana and has received grant funding from Roche.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–21, Supplementary Note and Supplementary Tables 1–3
Supplementary Table 4
Specificity values for Karolinska scRNA-seq superset
Rights and permissions
About this article
Cite this article
Skene, N.G., Bryois, J., Bakken, T.E. et al. Genetic identification of brain cell types underlying schizophrenia. Nat Genet 50, 825–833 (2018). https://doi.org/10.1038/s41588-018-0129-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0129-5
This article is cited by
-
Systematic immune cell dysregulation and molecular subtypes revealed by single-cell RNA-seq of subjects with type 1 diabetes
Genome Medicine (2024)
-
The genetic architecture of youth anxiety: a study protocol
BMC Psychiatry (2024)
-
A concerted neuron–astrocyte program declines in ageing and schizophrenia
Nature (2024)
-
rworkflows: automating reproducible practices for the R community
Nature Communications (2024)
-
Integrative cross-omics and cross-context analysis elucidates molecular links underlying genetic effects on complex traits
Nature Communications (2024)