Abstract
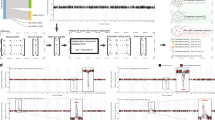
Although cancer genomes are replete with noncoding mutations, the effects of these mutations remain poorly characterized. Here we perform an integrative analysis of 930 tumor whole genomes and matched transcriptomes, identifying a network of 193 noncoding loci in which mutations disrupt target gene expression. These ‘somatic eQTLs’ (expression quantitative trait loci) are frequently mutated in specific cancer tissues, and the majority can be validated in an independent cohort of 3,382 tumors. Among these, we find that the effects of noncoding mutations on DAAM1, MTG2 and HYI transcription are recapitulated in multiple cancer cell lines and that increasing DAAM1 expression leads to invasive cell migration. Collectively, the noncoding loci converge on a set of core pathways, permitting a classification of tumors into pathway-based subtypes. The somatic eQTL network is disrupted in 88% of tumors, suggesting widespread impact of noncoding mutations in cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Cancer Genome Atlas Research Network et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
International Cancer Genome Consortium. International network of cancer genome projects. Nature 464, 993–998 (2010).
Hofree, M. et al. Challenges in identifying cancer genes by analysis of exome sequencing data. Nat. Commun. 7, 12096 (2016).
Iorio, F. et al. A landscape of pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Khurana, E. et al. Role of non-coding sequence variants in cancer. Nat. Rev. Genet. 17, 93–108 (2016).
Melton, C., Reuter, J. A., Spacek, D. V. & Snyder, M. Recurrent somatic mutations in regulatory regions of human cancer genomes. Nat. Genet. 47, 710–716 (2015).
Weinhold, N., Jacobsen, A., Schultz, N., Sander, C. & Lee, W. Genome-wide analysis of noncoding regulatory mutations in cancer. Nat. Genet. 46, 1160–1165 (2014).
Fredriksson, N. J., Ny, L., Nilsson, J. A. & Larsson, E. Systematic analysis of noncoding somatic mutations and gene expression alterations across 14 tumor types. Nat. Genet. 46, 1258–1263 (2014).
Huang, F. W. et al. Highly recurrent TERT promoter mutations in human melanoma. Science 339, 957–959 (2013).
Hoffman, M. M. et al. Integrative annotation of chromatin elements from ENCODE data. Nucleic Acids Res. 41, 827–841 (2013).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Ernst, J. & Kellis, M. Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues. Nat. Biotechnol. 33, 364–376 (2015).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Van der Auwera, G. A. et al. From FastQ data to high-confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Fishilevich, S. et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database 2017, https://doi.org/10.1093/database/bax028 (2017).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Li, Q. et al. Integrative eQTL-based analyses reveal the biology of breast cancer risk loci. Cell 152, 633–641 (2013).
Futreal, P. A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Habas, R., Kato, Y. & He, X. Wnt/Frizzled activation of Rho regulates vertebrate gastrulation and requires a novel Formin homology protein Daam1. Cell 107, 843–854 (2001).
Liu, W. et al. Mechanism of activation of the Formin protein Daam1. Proc. Natl Acad. Sci. USA 105, 210–215 (2008).
Zhu, Y. et al. Dvl2-dependent activation of Daam1 and RhoA regulates Wnt5a-induced breast cancer cell migration. PLoS One 7, e37823 (2012).
Fraley, S. I. et al. A distinctive role for focal adhesion proteins in three-dimensional cell motility. Nat. Cell Biol. 12, 598–604 (2010).
Fraley, S. I. et al. Three-dimensional matrix fiber alignment modulates cell migration and MT1-MMP utility by spatially and temporally directing protrusions. Sci. Rep. 5, 14580 (2015).
Kumar, S. & Weaver, V. M. Mechanics, malignancy, and metastasis: the force journey of a tumor cell. Cancer Metastasis Rev. 28, 113–127 (2009).
Velez, D. O. et al. 3D collagen architecture induces a conserved migratory and transcriptional response linked to vasculogenic mimicry. Nat. Commun. 8, 1651 (2017).
Hofree, M., Shen, J. P., Carter, H., Gross, A. & Ideker, T. Network-based stratification of tumor mutations. Nat. Methods 10, 1108–1115 (2013).
Cancer Genome Atlas Research Network. Integrated genomic characterization of papillary thyroid carcinoma. Cell 159, 676–690 (2014).
Wu, G., Feng, X. & Stein, L. A human functional protein interaction network and its application to cancer data analysis. Genome Biol. 11, R53 (2010).
Fatyol, K. & Szalay, A. A. The p14ARF tumor suppressor protein facilitates nucleolar sequestration of hypoxia-inducible factor-1α (HIF-1α) and inhibits HIF-1-mediated transcription. J. Biol. Chem. 276, 28421–28429 (2001).
Nishi, H. et al. Hypoxia-inducible factor 1 mediates upregulation of telomerase (hTERT). Mol. Cell. Biol. 24, 6076–6083 (2004).
Fan, Q.-W. et al. EGFR signals to mTOR through PKC and independently of Akt in glioma. Sci. Signal. 2, ra4 (2009).
Kawauchi, K., Ihjima, K. & Yamada, O. IL-2 increases human telomerase reverse transcriptase activity transcriptionally and posttranslationally through phosphatidylinositol 3′-kinase/Akt, heat shock protein 90, and mammalian target of rapamycin in transformed NK cells. J. Immunol. 174, 5261–5269 (2005).
Li, Y., Cheng, H. S., Chng, W. J. & Tergaonkar, V. Activation of mutant TERT promoter by RAS–ERK signaling is a key step in malignant progression of BRAF-mutant human melanomas. Proc. Natl Acad. Sci. USA 113, 14402–14407 (2016).
Cooper, M. T. & Bray, S. J. Frizzled regulation of Notch signalling polarizes cell fate in the Drosophila eye. Nature 397, 526–530 (1999).
Spruck, C. et al. A CDK-independent function of mammalian Cks1: targeting of SCFSkp2 to the CDK inhibitor p27Kip1. Mol. Cell 7, 639–650 (2001).
Lee, S.-H. et al. IFN-γ/IRF-1-induced p27kip1 down-regulates telomerase activity and human telomerase reverse transcriptase expression in human cervical cancer. FEBS Lett. 579, 1027–1033 (2005).
Warner, D. R., Roberts, E. A., Greene, R. M. & Pisano, M. M. Identification of novel Smad binding proteins. Biochem. Biophys. Res. Commun. 312, 1185–1190 (2003).
Okuyama, S. et al. Excessive activity of apolipoprotein B mRNA editing enzyme catalytic polypeptide 2 (APOBEC2) contributes to liver and lung tumorigenesis. Int. J. Cancer 130, 1294–1301 (2012).
Hirano, Y., Ohniwa, R. L., Wada, C., Yoshimura, S. H. & Takeyasu, K. Human small G proteins, ObgH1, and ObgH2, participate in the maintenance of mitochondria and nucleolar architectures. Genes Cells 11, 1295–1304 (2006).
Ashiuchi, M. & Misono, H. Biochemical evidence that Escherichia coli hyi (orfb0508, gip) gene encodes hydroxypyruvate isomerase. Biochim. Biophys. Acta 1435, 153–159 (1999).
Bush, W. S. & Moore, J. H. Chapter 11: genome-wide association studies. PLoS Comput. Biol. 8, e1002822 (2012).
Price, A. L., Zaitlen, N. A., Reich, D. & Patterson, N. New approaches to population stratification in genome-wide association studies. Nat. Rev. Genet. 11, 459–463 (2010).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Michaelson, J. J., Alberts, R., Schughart, K. & Beyer, A. Data-driven assessment of eQTL mapping methods. BMC Genomics 11, 502 (2010).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Hansen, R. S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl Acad. Sci. USA 107, 139–144 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Wu, P.-H., Giri, A., Sun, S. X. & Wirtz, D. Three-dimensional cell migration does not follow a random walk. Proc. Natl Acad. Sci. USA 111, 3949–3954 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Roberts, A. & Pachter, L. Streaming fragment assignment for real-time analysis of sequencing experiments. Nat. Methods 10, 71–73 (2013).
Cao, Y., Bryan, T. M. & Reddel, R. R. Increased copy number of the TERT and TERC telomerase subunit genes in cancer cells. Cancer Sci. 99, 1092–1099 (2008).
Xie, H. et al. TERT promoter mutations and gene amplification: promoting TERT expression in Merkel cell carcinoma. Oncotarget 5, 10048–10057 (2014).
Von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 17, 395–416 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Andersen, P. K. & Gill, R. D. Cox’s regression model for counting processes: a large sample study. Ann. Stat. 10, 1100–1120 (1982).
Acknowledgements
The results published here are in whole or part based upon data generated by the TCGA Research Network (see URLs). We would also like to acknowledge the clinical contributors and the data producers from the ICGC who have generated the particular datasets and made them available for public analysis. This work was supported by NIH grants to T.I. (U24CA184427, U54CA209891, P50GM085764, P41GM103504 and R01HG009979) and H.C. (DP5OD017937). G.X. is supported by a UCSD CTRI grant (UL1TR001442). S.I.F. and D.O.V. are supported by a Burroughs Wellcome Fund Career Award at the Scientific Interface (1012027), an NSF CAREER Award (1651855), and UCSD CTRI and FISP pilot grants. We would like to thank members of the Ideker laboratory for valuable comments and critical reading of the manuscript. Finally, we wish to thank the patients and their families for their contributions of valuable data without which this project would not have been possible.
Author information
Authors and Affiliations
Contributions
W.Z. and T.I. conceived the study. W.Z. designed and performed most of the analyses. G.X. performed mutation calling of 358 tumors. C.M. and M.S. provided mutation calling of 572 tumors. A.B.-G., K.S.S., J.P.S., K.M.O. and E.K.F. performed the somatic eQTL reporter assays. A.B.-G. and J.F.K. analyzed the flow cytometry and luciferase assay data. A.B.-G., J.P.S. and K.L. performed protein electropherogram analysis. D.O.V., K.C. and S.I.F. performed 3D cell culture assays. M.K.Y. and H.C. helped W.Z. in designing the somatic eQTL analysis. J.K.H. helped W.Z. in network analysis. T.I., J.F.K. and W.Z. wrote the manuscript and formulated all figures.
Corresponding authors
Ethics declarations
Competing interests
T.I. is cofounder of Data4Cure, Inc., and has an equity interest. T.I. has an equity interest in Ideaya BioSciences, Inc. The terms of this arrangement have been reviewed and approved by the University of California, San Diego, in accordance with its conflict-of-interest policies. No potential conflicts of interest were disclosed by the other authors.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Statistics of recurrently mutated loci.
a–e, Distribution (mean ± s.d.) of loci, enhancers and genes in somatic eQTL analysis. a, The sizes of recurrently mutated loci are typically <200 bp (52.0 ± 48.9 bp; n = 8800 loci). b,c, Most loci contain <10 SNV sites (4.0 ± 2.3; n = 8800 loci; b) and <40 patients with mutations (12.7 ± 10.4; n = 8800 loci; c). d, The length of enhancers (1,397 ± 1,934 bp; maximum = 86 kb; n = 247,345 enhancers) from the GeneHancer database17. e, Number of loci tested per gene in the somatic eQTL analysis (2.6 ± 3.1; maximum = 53; n = 9445 genes). f, Enrichment of mutations at each consecutive step of locus selection.
Supplementary Figure 2 Global transcriptional network of all significant somatic eQTLs.
a, Associations for which significance was better than a threshold (20% FDR) define a global network of somatic eQTLs. This network has been deposited in NDEx (http://www.ndexbio.org/; UUID: bd9e210f-dc84-11e7-adc1-0ac135e8bacf). b, The quantile–quantile plot shows the observed P values (F test, n = 783 tumors) versus random expectation for associations between somatic eQTLs and the expression levels of their target genes. FDR is calculated using the Storey approach50. c, Power analysis. All locus–gene pairs were plotted, evaluated by the number of patients with mutations (x axis) versus the change in gene expression given the mutation (y axis; one unit represents 1 s.d. of change in residual gene expression). Power was defined as 1 – P(type II error) at a significance level of P(type I error) = 0.0085, which is approximately at 20% FDR (n = 783 tumors).
Supplementary Figure 3 Examples of transcriptional regulation by somatic eQTLs.
a,b, A somatic eQTL on the promoter of TERT is associated with upregulation of TERT gene expression (a), with three mutation sites leading to the creation of binding motifs for Ets family members (b). c,d, A somatic eQTL in the 5′ UTR of KCNJ5 is associated with downregulation of this gene (c), with a gain of a Smad4 motif by a G-to-C mutation (d). e,f, Another somatic eQTL in the 5′ UTR of IQUB is associated with its transcriptional downregulation (e), by an A-to-G substitution creating an N-Myc binding motif (f). a,c,e, Violin plots were generated using the residual expression, which is the z-score-standardized RNA-seq data subtracting fitted values by all known and hidden covariates. P values were calculated using the F test (without multiple-testing correction; n = 783 tumors). Box plot elements are defined as in Fig. 1b. The 95% confidence intervals for the mean are (0.26, 0.45) and (–0.10, 0.00) (a), (–0.98, –0.26) and (–0.03, 0.06) (c), and (–0.56, –0.05) and (–0.03, 0.05) (e) for mutant and wild-type sequences, respectively.
Supplementary Figure 4 DAAM1 promoter activity assays in three additional cell lines.
a, Flow cytometry analysis of MDA-MB-231 breast cancer cells 48 h after transient transfection with the different GFP reporter constructs. A ‘live’ gate was drawn based on forward scatter versus side scatter. The first of three triplicate samples is shown here. b, For all three of the triplicates, the polygon delineated by black lines shows the gated region used to define GFP+ cells. The first row corresponds to the samples depicted in a. d,f, Flow cytometry analysis of U2OS osteosarcoma cells (d) and RPMI-7951 metastatic melanoma cells (f). These plots are representative of three independent cell culture experiments. c,e,g, Bar graphs (average ± s.d. across three cell culture replicates; P values are from two-tailed t tests with three wild-type and three mutant samples) showing the percentage of GFP+ cells and the median fluorescence intensity of GFP+ events in MDA-MB-231 (c), U2OS (e) and RPMI-7951 (g). Individual data points are available in Supplementary Table 5.
Supplementary Figure 5 3D collagen cell migration assay with RNA-seq analysis.
a, Representative trajectories of breast cancer cells migrating through 3D culture conditions. In low-density collagen (2.5 mg/ml) mimicking the stiffness of normal tissue, both MDA-MB-231 breast cancer cells and HT-1080 fibrosarcoma cells display a less invasive phenotype, whereas in high-density collagen (6 mg/ml) mimicking the stiffness of tumor tissue, cells display a highly invasive phenotype26, 27. Trajectories shown are representative of observations made in at least 60 cells from three independent experiments. b, RNA-seq analysis reveals a marked upregulation of DAAM1 in the high-density collagen condition where enhanced invasion is observed. DAAM1 is one of the most upregulated transcripts associated with this invasive migration phenotype across the genome28 (mean ± s.e.m. across three independent cell cultures; P values from two-tailed t tests; the 95% confidence intervals of the mean are (–2.0, 39.4) and (13.9, 63.3) in HT-1080 cells and (–0.4, 28.0) and (22.6, 32.9) in MDA-MB-231 cells for lowly and highly invasive phenotypes, respectively).
Supplementary Figure 6 3D collagen cell migration assay with DAAM1 overexpression.
a, Persistence of the motility of wild-type and DAAM1-overexpressing MDA-MB-231 cells, defined as the ratio between the total invasion distance and the path length (P = 0.008, two-tailed Mann–Whitney U test; the 95% confidence intervals of the mean are (0.07, 0.10) and (0.10, 0.14) for wild-type and DAAM1-overexpressing cells, respectively). b, Length of the trajectories travelled by the cells. c, Single-cell mean velocity. In a–c, 74 and 83 cells were imaged in the wild-type and DAAM1 overexpression groups, respectively. d, Total invasion distance travelled by individual cells with additional Wnt5a signaling, defined as the distance from start to end of the cell’s trajectory (P = 0.0002, two-tailed Mann–Whitney U test; the 95% confidence intervals of the mean are (11.4 µm, 17.5 µm) and (18.8 µm, 60.3 µm) for wild-type and DAAM1-overexpressing cells, respectively). Here 63 and 15 cells were imaged in the wild-type and DAAM1 overexpression groups, respectively. In a–d, box plot elements are defined as in Fig. 1b. e, Uncropped pictures of protein the electropherogram presented in the main figure Fig. 3e. Bands were cropped to save space in the main figure as indicated by dashed boxes.
Supplementary Figure 7 Association between NBS subtypes and cancer tissues.
a, The number of patients (y axis) is plotted against cancer tissues (x axis). NBS subtypes are represented by colors. b, Pairwise one-sided Fisher’s exact test between cancer tissues and molecular subtypes (n = 810 tumors). c, Percentage of patients shared between the subtypes derived from both coding and noncoding mutations and those derived from coding mutations only.
Supplementary Figure 8 Association of NBS subtypes with patient disease-free survival times.
Log-likelihood ratio test P values (n = 793 patients) of significance (y axis) are plotted against the numbers of subtypes (x axis). Each blue bar represents a P value of a simple Cox proportional-hazard model, where survival time is a function of subtypes. Each orange bar represents a P value by comparing a complete model, which takes into account subtypes and tissues, against a null model that includes tissues only.
Supplementary Figure 9 Network views of the TERT–BRAF–IDH1 pathway subtype.
a, Pathways characterizing the TERT–BRAF–IDH1 subtype, defined as subnetwork regions extracted from ReactomeFI by NBS. b, Mutation matrix of the TERT–BRAF–IDH1 pathway subtype showing individual tumors (columns; ordered by cancer tissues) with indicated types of mutations on signature genes (rows).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Table 4 and Supplementary Note
Supplementary Tables
Supplementary Tables 1–3, 5 and 6
Rights and permissions
About this article
Cite this article
Zhang, W., Bojorquez-Gomez, A., Velez, D.O. et al. A global transcriptional network connecting noncoding mutations to changes in tumor gene expression. Nat Genet 50, 613–620 (2018). https://doi.org/10.1038/s41588-018-0091-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0091-2
This article is cited by
-
A CRISPR-drug perturbational map for identifying compounds to combine with commonly used chemotherapeutics
Nature Communications (2023)
-
Functional analysis of recurrent CDC20 promoter variants in human melanoma
Communications Biology (2023)
-
Artificial intelligence reveals dysregulation of osteosarcoma and cuproptosis-related biomarkers, PDHA1, CDKN2A and neutrophils
Scientific Reports (2023)
-
Interpretation of the role of germline and somatic non-coding mutations in cancer: expression and chromatin conformation informed analysis
Clinical Epigenetics (2022)
-
PROBER identifies proteins associated with programmable sequence-specific DNA in living cells
Nature Methods (2022)