Abstract
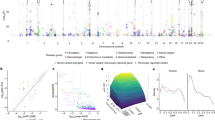
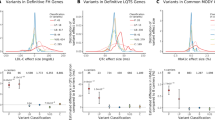
We aggregated coding variant data for 81,412 type 2 diabetes cases and 370,832 controls of diverse ancestry, identifying 40 coding variant association signals (P < 2.2 × 10−7); of these, 16 map outside known risk-associated loci. We make two important observations. First, only five of these signals are driven by low-frequency variants: even for these, effect sizes are modest (odds ratio ≤1.29). Second, when we used large-scale genome-wide association data to fine-map the associated variants in their regional context, accounting for the global enrichment of complex trait associations in coding sequence, compelling evidence for coding variant causality was obtained for only 16 signals. At 13 others, the associated coding variants clearly represent ‘false leads’ with potential to generate erroneous mechanistic inference. Coding variant associations offer a direct route to biological insight for complex diseases and identification of validated therapeutic targets; however, appropriate mechanistic inference requires careful specification of their causal contribution to disease predisposition.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kooner, J. S. et al. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat. Genet. 43, 984–989 (2011).
Cho, Y. S. et al. Meta-analysis of genome-wide association studies identifies eight new loci for type 2 diabetes in east Asians. Nat. Genet. 44, 67–72 (2011).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Mahajan, A. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Ng, M. C. et al. Meta-analysis of genome-wide association studies in African Americans provides insights into the genetic architecture of type 2 diabetes. PLoS. Genet. 10, e1004517 (2014).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Walter, K. et al. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
Gaulton, K. J. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat. Genet. 47, 1415–1425 (2015).
Horikoshi, M. et al. Transancestral fine-mapping of four type 2 diabetes susceptibility loci highlights potential causal regulatory mechanisms. Hum. Mol. Genet. 25, 2070–2081 (2016).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS. Med. 12, e1001779 (2015).
Cook, J. P. & Morris, A. P. Multi-ethnic genome-wide association study identifies novel locus for type 2 diabetes susceptibility. Eur. J. Hum. Genet. 24, 1175–1180 (2016).
Estrada, K. et al. Association of a low-frequency variant in HNF1A with type 2 diabetes in a Latino population. J. Am. Med. Assoc. 311, 2305–2314 (2014).
Sveinbjornsson, G. et al. Weighting sequence variants based on their annotation increases power of whole-genome association studies. Nat. Genet. 48, 314–317 (2016).
Liu, D. J. et al. Meta-analysis of gene-level tests for rare variant association. Nat. Genet. 46, 200–204 (2014).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Steinthorsdottir, V. et al. Identification of low-frequency and rare sequence variants associated with elevated or reduced risk of type 2 diabetes. Nat. Genet. 46, 294–298 (2014).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Maller, J. B. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Flannick, J. et al. Loss-of-function mutations in SLC30A8 protect against type 2 diabetes. Nat. Genet. 46, 357–363 (2014).
Beer, N. L. et al. The P446L variant in GCKR associated with fasting plasma glucose and triglyceride levels exerts its effect through increased glucokinase activity in liver. Hum. Mol. Genet. 18, 4081–4088 (2009).
Murphy, R., Ellard, S. & Hattersley, A. T. Clinical implications of a molecular genetic classification of monogenic beta-cell diabetes. Nat. Clin. Pract. Endocrinol. Metab. 4, 200–213 (2008).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Kozlitina, J. et al. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 46, 352–356 (2014).
Kulzer, J. R. et al. A common functional regulatory variant at a type 2 diabetes locus upregulates ARAP1 expression in the pancreatic beta cell. Am. J. Hum. Genet. 94, 186–197 (2014).
Carrat, G. R. et al. Decreased STARD10 expression is associated with defective insulin secretion in humans and mice. Am. J. Hum. Genet. 100, 238–256 (2017).
Deeb, S. S. et al. A Pro12Ala substitution in PPARγ2 associated with decreased receptor activity, lower body mass index and improved insulin sensitivity. Nat. Genet. 20, 284–287 (1998).
Majithia, A. R. et al. Rare variants in PPARG with decreased activity in adipocyte differentiation are associated with increased risk of type 2 diabetes. Proc Natl Acad Sci USA 111, 13127–13132 (2014).
Majithia, A. R. et al. Prospective functional classification of all possible missense variants in PPARG. Nat. Genet. 48, 1570–1575 (2016).
Claussnitzer, M. et al. Leveraging cross-species transcription factor binding site patterns: from diabetes risk loci to disease mechanisms. Cell 156, 343–358 (2014).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Dimas, A. S. et al. Impact of type 2 diabetes susceptibility variants on quantitative glycemic traits reveals mechanistic heterogeneity. Diabetes. 63, 2158–2171 (2014).
Lotta, L. A. et al. Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nat. Genet. 49, 17–26 (2017).
Altshuler, D. & Daly, M. Guilt beyond a reasonable doubt. Nat. Genet. 39, 813–815 (2007).
Grove, M. L. et al. Best practices and joint calling of the HumanExome BeadChip: the CHARGE Consortium. PLoS. ONE. 8, e68095 (2013).
Price, A. L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 132–135 (2008). author reply 135–139.
Weale, M. E. Quality control for genome-wide association studies. Methods. Mol. Biol. 628, 341–372 (2010).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics. 55, 997–1004 (1999).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Eastwood, S. V. et al. Algorithms for the capture and adjudication of prevalent and incident diabetes in UK Biobank. PLoS. ONE. 11, e0162388 (2016).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Hoffmann, T. J. et al. Next generation genome-wide association tool: design and coverage of a high-throughput European-optimized SNP array. Genomics. 98, 79–89 (2011).
Hoffmann, T. J. et al. Design and coverage of high throughput genotyping arrays optimized for individuals of East Asian, African American, and Latino race/ethnicity using imputation and a novel hybrid SNP selection algorithm. Genomics. 98, 422–430 (2011).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS. Genet. 5, e1000529 (2009).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G. R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Winkler, T. W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Cook, J. P., Mahajan, A. & Morris, A. P. Guidance for the utility of linear models in meta-analysis of genetic association studies of binary phenotypes. Eur. J. Hum. Genet. 25, 240–245 (2017).
Ioannidis, J. P., Patsopoulos, N. A. & Evangelou, E. Heterogeneity in meta-analyses of genome-wide association investigations. PLoS. ONE. 2, e841 (2007).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012). S1–S3.
Wakefield, J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
Acknowledgements
A full list of acknowledgments appears in the Supplementary Note. Part of this work was conducted using the UK Biobank Resource under application number 9161.
Author information
Authors and Affiliations
Consortia
Contributions
Project coordination: A. Mahajan, A.P.M., J.I.R., M.I.M. Core analyses and writing: A. Mahajan, J.W., S.M.W., W. Zhao, N.R.R., A.Y.C., W.G., H.K., R.A.S., I. Barroso, T.M.F., M.O.G., J.B.M., M. Boehnke, D.S., A.P.M., J.I.R., M.I.M. Statistical analysis in individual studies: A. Mahajan, J.W., S.M.W., W. Zhao, N.R.R., A.Y.C., W.G., H.K., D.T., N.W.R., X.G., Y. Lu, M. Li, R.A.J., Y. Hu, S. Huo, K.K.L., W. Zhang, J.P.C., B.P.P., J. Flannick, N.G., V.V.T., J. Kravic, Y.J.K., D.V.R., H.Y., M.M.-N., K.M., R.L.-G., T.V.V., J. Marten, J. Li, A.V.S., P. An, S.L., S.G., G.M., A. Demirkan, J.F.T., V. Steinthorsdottir, M.W., C. Lecoeur, M. Preuss, L.F.B., P. Almgren, J.B.-J., J.A.B., M.C., K.-U.E., K.F.,H.G.d.H., Y. Hai, S. Han, S.J., F. Kronenberg, K.L., L.A.L., J.-J.L., H.L., C.-T.L., J. Liu, R.M., K.R., S.S., P.S., T.M.T., G.T., A. Tin, A.R.W., P.Y., J.Y., L.Y., R.Y., J.C.C., D.I.C., C.v.D., J. Dupuis, P.W.F., A. Köttgen, D.M.-K., N. Soranzo, R.A.S., A.P.M. Genotyping: A. Mahajan, N.R.R., A.Y.C., Y. Lu, Y. Hu, S. Huo, B.P.P., N.G., R.L.-G., P. An, G.M., E.A., N.A., C.B., N.P.B., Y.-D.I.C., Y.S.C., M.L.G., H.G.d.H., S. Hackinger, S.J., B.-J.K., P.K., J. Kriebel, F. Kronenberg, H.L., S.S.R., K.D.T., E.B., E.P.B., P.D., J.C.F., S.R.H., C. Langenberg, M.A.P., F.R., A.G.U., J.C.C., D.I.C., P.W.F., B.-G.H., C.H., E.I., S.L.R.K., J.S.K., Y. Liu, R.J.F.L., N. Soranzo, N.J.W., R.A.S., T.M.F., A.P.M., J.I.R., M.I.M. Cross-trait lookups in unpublished data: S.M.W., A.Y.C., Y. Lu, M. Li, M.G., H.M.H., A.E.J., D.J.L., E.M., G.M.P., H.R.W., S.K., C.J.W. Phenotyping: Y. Lu, Y. Hu, S. Huo, P. An, S.L., A. Demirkan, S. Afaq, S. Afzal, L.B.B., A.G.B., I. Brandslund, C.C., S.V.E., G.G., V. Giedraitis, A.T.-H., M.-F.H., B.I., M.E.J., T.J., A. Käräjämäki, S.S.K., H.A.K., P.K., F. Kronenberg, B.L., H.L., K.-H.L., A.L., J. Liu, M. Loh, V.M., R.M.-C., G.N., M.N., S.F.N., I.N., P.A.P., W.R., L.R., O.R., S.S., E.S., K.S.S., A.S., B.T., A. Tönjes, A.V., D.R.W., H.B., E.P.B., A. Dehghan, J.C.F., S.R.H., C. Langenberg, A.D. Morris, R.d.M., M.A.P., A.R., P.M.R., F.R.R., V. Salomaa, W.H.-H.S., R.V., J.C.C., J. Dupuis, O.H.F., H.G., B.-G.H., T.H., A.T.H., C.H., S.L.R.K., J.S.K., A. Köttgen, L.L., Y. Liu, R.J.F.L., C.N.A.P., J.S.P., O.P., B.M.P., M.B.S., N.J.W., T.M.F., M.O.G. Individual study design and principal investigators: N.G., P. An, B.-J.K., P. Amouyel, H.B., E.B., E.P.B., R.C., F.S.C., G.D., A. Dehghan, P.D., M.M.F., J. Ferrières, J.C.F., P. Frossard, V. Gudnason, T.B.H., S.R.H., J.M.M.H., M.I., F. Kee, J. Kuusisto, C. Langenberg, L.J.L., C.M.L., S.M., T.M., O.M., K.L.M., M.M., A.D. Morris, A.D. Murray, R.d.M., M.O.-M., K.R.O., M. Perola, A.P., M.A.P., P.M.R., F.R., F.R.R., A.H.R., V. Salomaa, W.H.-H.S., R.S., B.H.S., K. Strauch, A.G.U., R.V., M. Blüher, A.S.B., J.C.C., D.I.C., J. Danesh, C.v.D., O.H.F., P.W.F., P. Froguel, H.G., L.G., T.H., A.T.H., C.H., E.I., S.L.R.K., F. Karpe, J.S.K., A. Köttgen, K.K., M. Laakso, X.L., L.L., Y. Liu, R.J.F.L., J. Marchini, A. Metspalu, D.M.-K., B.G.N., C.N.A.P., J.S.P., O.P., B.M.P., R.R., N. Sattar, M.B.S., N. Soranzo, T.D.S., K. Stefansson, M.S., U.T., T.T., J.T., N.J.W., J.G.W., E.Z., I. Barroso, T.M.F., J.B.M., M. Boehnke, D.S., A.P.M., J.I.R., M.I.M.
Corresponding authors
Ethics declarations
Competing interests
J.C.F. has received consulting honoraria from Merck and from Boehringer-Ingelheim. D.I.C. received funding for exome chip genotyping in the WGHS from Amgen. O.H.F. works in ErasmusAGE, a center for aging research across the life course funded by Nestlé Nutrition (Nestec, Ltd.), Metagenics, Inc., and AXA. Nestlé Nutrition (Nestec, Ltd.), Metagenics, Inc., and AXA had no role in the design or conduct of the study; collection, management, analysis, or interpretation of the data; or preparation, review, or approval of the manuscript. E.I. is an advisor and consultant for Precision Wellness, Inc., and an advisor for Cellink for work unrelated to the present project. B.M.P. serves on the DSMB for a clinical trial funded by the manufacturer (Zoll LifeCor) and on the Steering Committee of the Yale Open Data Access Project funded by Johnson & Johnson. I. Barroso and spouse own stock in GlaxoSmithKline and Incyte Corporation. T.F. has consulted for Boeringer–Ingelheim and Sanofi on the genetics of diabetes. D.S. has received support from Pfizer, Regeneron, Genentech, and Eli Lilly. M.I.M. has served on advisory panels for Novo Nordisk and Pfizer and received honoraria from Novo Nordisk, Pfizer, Sanofi-Aventis, and Eli Lilly.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 3–8 and Supplementary Note
Supplementary Tables
Supplementary Tables 1, 2 and 9–11
Rights and permissions
About this article
Cite this article
Mahajan, A., Wessel, J., Willems, S.M. et al. Refining the accuracy of validated target identification through coding variant fine-mapping in type 2 diabetes. Nat Genet 50, 559–571 (2018). https://doi.org/10.1038/s41588-018-0084-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0084-1
This article is cited by
-
Association of birth weight with type 2 diabetes mellitus and the mediating role of fatty acids traits: a two-step mendelian randomization study
Lipids in Health and Disease (2024)
-
Multi-ancestry polygenic mechanisms of type 2 diabetes
Nature Medicine (2024)
-
Genetic determinants of plasma protein levels in the Estonian population
Scientific Reports (2024)
-
Association between fatty liver index and controlled attenuation parameters as markers of metabolic dysfunction-associated fatty liver disease and bone mineral density: observational and two-sample Mendelian randomization studies
Osteoporosis International (2024)
-
Lessons and Applications of Omics Research in Diabetes Epidemiology
Current Diabetes Reports (2024)