Abstract
Clustering of mutations has been observed in cancer genomes as well as for germline de novo mutations (DNMs). We identified 1,796 clustered DNMs (cDNMs) within whole-genome-sequencing data from 1,291 parent–offspring trios to investigate their patterns and infer a mutational mechanism. We found that the number of clusters on the maternal allele was positively correlated with maternal age and that these clusters consisted of more individual mutations with larger intermutational distances than those of paternal clusters. More than 50% of maternal clusters were located on chromosomes 8, 9 and 16, in previously identified regions with accelerated maternal mutation rates. Maternal clusters in these regions showed a distinct mutation signature characterized by C>G transversions. Finally, we found that maternal clusters were associated with processes involving double-strand-breaks (DSBs), such as meiotic gene conversions and de novo deletion events. This result suggested accumulation of DSB-induced mutations throughout oocyte aging as the mechanism underlying the formation of maternal mutation clusters.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
23 July 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41588-021-00905-z
References
Veltman, J. A. & Brunner, H. G. De novo mutations in human genetic disease. Nat. Rev. Genet. 13, 565–575 (2012).
Kong, A. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475 (2012).
Wong, W. S. W. et al. New observations on maternal age effect on germline de novo mutations. Nat. Commun. 7, 10486 (2016).
Goldmann, J. M. et al. Parent-of-origin-specific signatures of de novo mutations. Nat. Genet. 48, 935–939 (2016).
Crow, J. F. The origins, patterns and implications of human spontaneous mutation. Nat. Rev. Genet. 1, 40–47 (2000).
Ségurel, L., Wyman, M. J. & Przeworski, M. Determinants of mutation rate variation in the human germline. Annu. Rev. Genomics Hum. Genet. 15, 47–70 (2014).
Michaelson, J. J. et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell 151, 1431–1442 (2012).
Schrider, D. R., Hourmozdi, J. N. & Hahn, M. W. Pervasive multinucleotide mutational events in eukaryotes. Curr. Biol. 21, 1051–1054 (2011).
Yuen, R. K. et al. Genome-wide characteristics of de novo mutations in autism. NPJ Genom. Med. 1, 160271–1602710 (2016).
Besenbacher, S. et al. Multi-nucleotide de novo mutations in humans. PLoS Genet. 12, e1006315 (2016).
Terekhanova, N. V., Bazykin, G. A., Neverov, A., Kondrashov, A. S. & Seplyarskiy, V. B. Prevalence of multinucleotide replacements in evolution of primates and Drosophila. Mol. Biol. Evol. 30, 1315–1325 (2013).
Francioli, L. C. et al. Genome-wide patterns and properties of de novo mutations in humans. Nat. Genet. 47, 822–826 (2015).
Rahbari, R. et al. Timing, rates and spectra of human germline mutation. Nat. Genet. 48, 126–133 (2016).
Harris, K. & Nielsen, R. Error-prone polymerase activity causes multinucleotide mutations in humans. Genome Res. 24, 1445–1454 (2014).
Bodian, D. L. et al. Utility of whole-genome sequencing for detection of newborn screening disorders in a population cohort of 1,696 neonates. Genet. Med. 18, 221–230 (2015).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Titus, S. et al. Impairment of BRCA1-related DNA double-strand break repair leads to ovarian aging in mice and humans. Sci. Transl. Med. 5, 172ra21 (2013).
White, R. R. & Vijg, J. Do DNA double-strand breaks drive aging? Mol. Cell 63, 729–738 (2016).
Oktay, K., Turan, V., Titus, S., Stobezki, R. & Liu, L. BRCA mutations, DNA repair deficiency, and ovarian aging. Biol. Reprod. 93, 67 (2015).
Kong, A. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010).
Halldorsson, B. V. et al. The rate of meiotic gene conversion varies by sex and age. Nat. Genet. 48, 1377–1384 (2016).
Martin, H. C. et al. Multicohort analysis of the maternal age effect on recombination. Nat. Commun. 6, 7846 (2015).
Campbell, C. L., Furlotte, N. A., Eriksson, N., Hinds, D. & Auton, A. Escape from crossover interference increases with maternal age. Nat. Commun. 6, 6260 (2015).
Arbeithuber, B., Betancourt, A. J., Ebner, T. & Tiemann-Boege, I. Crossovers are associated with mutation and biased gene conversion at recombination hotspots. Proc. Natl. Acad. Sci. USA 112, 2109–2114 (2015).
Lercher, M. J. & Hurst, L. D. Human SNP variability and mutation rate are higher in regions of high recombination. Trends Genet. 18, 337–340 (2002).
Webster, M. T. & Hurst, L. D. Direct and indirect consequences of meiotic recombination: implications for genome evolution. Trends Genet. 28, 101–109 (2012).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Zámborszky, J. et al. Loss of BRCA1 or BRCA2 markedly increases the rate of base substitution mutagenesis and has distinct effects on genomic deletions. Oncogene 36, 746–755 (2017).
Moynahan, M. E., Chiu, J. W., Koller, B. H. & Jasin, M. Brca1 controls homology-directed DNA repair. Mol. Cell 4, 511–518 (1999).
Patel, K. J. et al. Involvement of Brca2 in DNA repair. Mol. Cell 1, 347–357 (1998).
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010).
Kong, A. et al. Recombination rate and reproductive success in humans. Nat. Genet. 36, 1203–1206 (2004).
Ottolini, C. S. et al. Genome-wide maps of recombination and chromosome segregation in human oocytes and embryos show selection for maternal recombination rates. Nat. Genet. 47, 727–735 (2015).
Middlebrooks, C. D. et al. Evidence for dysregulation of genome-wide recombination in oocytes with nondisjoined chromosomes 21. Hum. Mol. Genet. 23, 408–417 (2014).
Jónsson, H. et al. Parental influence on human germline de novo mutations in 1,548 trios from Iceland. Nature 549, 519–522 (2017).
Raczy, C. et al. Isaac: ultra-fast whole-genome secondary analysis on Illumina sequencing platforms. Bioinformatics 29, 2041–2043 (2013).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Derrien, T. et al. Fast computation and applications of genome mappability. PLoS One 7, e30377 (2012).
Gel, B. et al. regioneR: an R/Bioconductor package for the association analysis of genomic regions based on permutation tests. Bioinformatics 32, 289–291 (2016).
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Blokzijl, F., Janssen, R., Van Boxtel, R. & Cuppen, E. MutationalPatterns: an integrative R package for studying patterns in base substitution catalogues. Preprint at https://www.biorxiv.org/content/early/2016/08/30/071761 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Acknowledgements
This study was funded by the Inova Health System with support from Fairfax County and philanthropic support from the Odeen family. We thank the Inova Translational Medicine Institute staff for supporting the study. We also thank the families who participated in the genomic studies that made this research possible. This work was partly financially supported by grants from the Netherlands Organization for Scientific Research (916-14-043 to C.G. and 918-15-667 to J.A.V.) and the European Research Council (ERC Starting grant DENOVO 281964 to J.A.V.).
This study used data generated by the Genome of the Netherlands Project. A full list of the investigators is available from http://www.nlgenome.nl/. Funding for the project was provided by the Netherlands Organization for Scientific Research under award number 184021007, dated July 9, 2009 and made available as a Rainbow Project of the Biobanking and Biomolecular Research Infrastructure Netherlands (BBMRI-NL). The sequencing was carried out in collaboration with the Beijing Institute for Genomics (BGI).
Author information
Authors and Affiliations
Contributions
C.G. and J.E.N. designed the study. J.M.G., V.B.S. and W.S.W.W. performed the data analyses. W.S.W.W. carried out QC and de novo mutation calling. T.V. performed Sanger validation. B.D.S., J.F.D. and J.E.N. supervised the data collection, sequencing and writing of the manuscript. D.L.B. assisted in data analyses and interpretation. J.M.G., V.B.S., W.S.W.W., J.A.V. and C.G. drafted the manuscript. P.B.N. acquired part of the replication data. All authors contributed to the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Linear models of age effects.
(a) Linear models for the numbers of clustered and unclustered DNMs. (b) Linear models for the numbers of cluster events. Grey shades indicate standard errors.
Supplementary Figure 2 Parental ages by the number of clusters per individual.
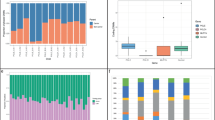
(a) Primary cohort and (b) replication cohort. Boxplot whiskers depict distance from quartile to a maximum of 1.58 times the interquartile range. Numbers indicate number of individuals per group. While the maternal age increases with the number of clusters, the paternal age does not.
Supplementary Figure 3 Differences between maternal and paternal cDNMs in the replication cohort.
(a) The fraction of probands with maternal and paternal clustered mutations (y-axis), grouped by parental age quantiles. Error bars indicate the binomial 95% confidence intervals. (b) The number of paternal and maternal cDNMs (y-axis) stratified by the distance to the nearest other cDNM (x-axis). (c) The size of paternal and maternal age effect of clusters with at least one phased cDNM (y-axis) by inter-mutational distance (x-axis). Whiskers indicate the 95% confidence interval. (d) Age of fathers at conception and (e) age of the mothers at conception (y-axis) by the number of mutations in the offspring’s largest mutation cluster originating from the respective parent (x-axis). We considered only clusters where at least one cDNM is on the allele from the respective parent (paternal allele for d and maternal allele for e). Numbers indicate the size of each group. Boxplot compartments: box: interquartile range; line: median; whiskers: extreme values <1.5 × interquartile ranges from box borders).
Supplementary Figure 4
Numbers of phased unclustered DNMs and cDNMs per chromosome in the primary cohort.
Supplementary Figure 5 Patterns of cDNMs across the chromosomes in the replication cohort.
(a) The fraction of phased cDNMs per chromosome. Error bars indicate the binomial 95% confidence intervals. (b) The nucleotide substitution spectrum of maternal and paternal clusters and unclustered DNMs. Error bars indicate the binomial 95% confidence intervals. (c) The nucleotide substitution spectrum of cDNMs by location. Error bars indicate the binomial 95% confidence intervals.
Supplementary Figure 6 cDNM-enriched regions on chromosomes 8 and 9.
Overview of regions enriched for maternal cluster mutations. X-axis and ideograms indicate chromosomal position. The red and blue histograms indicate the number of maternal cDNMs and paternal cDNMs identified in this study, respectively. The pale red and pale blue histograms indicate the number of maternal and paternal unclustered DNMs. The lowest track indicates normalized cSNP C>G score, which is predictive for maternal DNMs. (a) Full chromosome 8. (b) Region with increased maternal mutation rate on chromosome 9 (chr9: 0-10,000,000).
Supplementary Figure 7 cDNM-enriched regions in the replication cohort.
Overview of regions enriched for maternal cluster mutations. X-axis and ideograms indicate chromosomal position. The red and blue histograms indicate the number of maternal cDNMs and paternal cDNMs identified in this study, respectively. The pale red and pale blue histograms indicate the number of maternal and paternal unclustered DNMs. The lowest track indicates normalized cSNP C>G score, which is predictive for maternal DNMs. (a) Full chromosome 16. (b) Full chromosome 8. (c) Region with increased maternal mutation rate on chromosome 9 (chr9: 0-10,000,000). (d) Region with increased maternal mutation rate on chromosome 2 (chr2: 0-10,000,000).
Supplementary Figure 8
Relation between cSNP C>G score and the number of phased clusters in genomic bins of 1 Mb.
Supplementary Figure 9
DNMs within 100 kb of the two de novo deletion events in the replication cohort.
Supplementary Figure 10 Recombination scores of cDNM regions.
Recombination scores (as defined by Kong et al.20) of cDNM regions. (a) Recombination scores of genomic regions harboring unclustered DNM and cDNM in primary cohort. (b) Recombination scores of genomic regions harboring unclustered DNM and cDNM in replication cohort. (c) Recombination scores of genomic regions harbouring cSNPs. The numbers indicate one-sided p-values for a difference between the groups, based on Wilcoxon rank sum test.
Supplementary Figure 11 Fitting of cancer signatures.
(a) Fitting to unclustered DNMs and cDNMs. (b) Fitting to maternal cDNMs and paternal cDNMs. The solid error bars indicate the standard deviation of resampled mutations’ contributions; the dashed error bars indicate 95% confidence intervals of the resampled mutations’ contributions.
Supplementary Figure 12 Principal component analysis of sequencing-quality statistics.
The quality control variables are described in Supplementary Table 18. (a) First two principal components plotted against each other and colored by software version of data analysis pipeline. Spearman-correlation coefficient of PC1 and average coverage: −0.893. (b) Variance explained by principal components. (c) Principal components two and three plotted against each other and colored by estimated ancestry of sequenced individual.
Supplementary Figure 13
Number of callable bases by sequencing batch.
Supplementary Figure 14
Number of filtered DNMs versus average genome coverage in the proband.
Supplementary Figure 15 C>G mutations in cSNPs.
(a) cSNPs depleted by CpG>CpT mutations, but enriched by remaining C>G mutations, reproducing hallmarks of cDNM spectra. (b) Fraction of non-CpG C>G nucleotide substitutions in cSNP spectra decreases with inter-mutational distances, showing a lower fraction of real clusters at higher distances.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Note and Supplementary Tables 1–4, 7–12, 14–25
Supplementary Table 5: cDNMs
List of clustered DNMs
Supplementary Table 6: Clusters per trio
Number of clusters per trio
Supplementary Table 13: cSNPs
List of cSNPs
Rights and permissions
About this article
Cite this article
Goldmann, J.M., Seplyarskiy, V.B., Wong, W.S.W. et al. Germline de novo mutation clusters arise during oocyte aging in genomic regions with high double-strand-break incidence. Nat Genet 50, 487–492 (2018). https://doi.org/10.1038/s41588-018-0071-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0071-6
This article is cited by
-
Meta-analysis of 46,000 germline de novo mutations linked to human inherited disease
Human Genomics (2024)
-
A genomic mutational constraint map using variation in 76,156 human genomes
Nature (2024)
-
Identification and characterisation of de novo germline structural variants in two commercial pig lines using trio-based whole genome sequencing
BMC Genomics (2023)
-
Revealing parental mosaicism: the hidden answer to the recurrence of apparent de novo variants
Human Genomics (2023)
-
A mutation rate model at the basepair resolution identifies the mutagenic effect of polymerase III transcription
Nature Genetics (2023)