Abstract
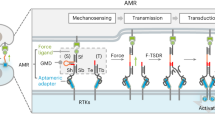
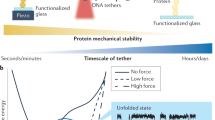
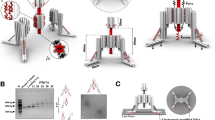
Cells interpret mechanical stimuli from their environments and neighbors, but the ability to engineer customized mechanosensing capabilities has remained a synthetic and mechanobiology challenge. Here we introduce tension-tuned synthetic Notch (SynNotch) receptors to convert extracellular and intercellular forces into specifiable gene expression changes. By elevating the tension requirements of SynNotch activation, in combination with structure-guided mutagenesis, we designed a set of receptors with mechanical sensitivities spanning the physiologically relevant picoNewton range. Cells expressing these receptors can distinguish between varying tensile forces and respond by enacting customizable transcriptional programs. We applied these tools to design a decision-making circuit, through which fibroblasts differentiate into myoblasts upon stimulation with distinct tension magnitudes. We also characterize cell-generated forces transmitted between cells during Notch signaling. Overall, this work provides insight into how mechanically induced changes in protein structure can be used to transduce physical forces into biochemical signals. The system should facilitate the further programming and dissection of force-related phenomena in biological systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Schoen, I., Pruitt, B. L. & Vogel, V. The Yin-Yang of rigidity sensing: how forces and mechanical properties regulate the cellular response to materials. Annu. Rev. Mater. Res. 43, 589–618 (2013).
Wells, R. G. Tissue mechanics and fibrosis. Biochim. Biophys. Acta 1832, 884–890 (2013).
Hahn, C. & Schwartz, M. A. Mechanotransduction in vascular physiology and atherogenesis. Nat. Rev. Mol. Cell Biol. 10, 53–62 (2009).
Mammoto, T., Mammoto, A. & Ingber, D. E. Mechanobiology and developmental control. Annu. Rev. Cell Dev. Biol. 29, 27–61 (2013).
Trylinski, M. & Schweisguth, F. Activation of Arp2/3 by WASp is essential for the endocytosis of δ only during cytokinesis in Drosophila. Cell Rep. 28, 1–10 (2019).
Bertolio, R. et al. Sterol regulatory element binding protein one couples mechanical cues and lipid metabolism. Nat. Commun. 10, 1326 (2019).
Van Helvert, S., Storm, C. & Friedl, P. Mechanoreciprocity in cell migration. Nat. Cell Biol. 20, 8–20 (2018).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Das, D. K. et al. Force-dependent transition in the T-cell receptor β-subunit allosterically regulates peptide discrimination and pMHC bond lifetime. Proc. Natl Acad. Sci. USA 112, 1517–1522 (2015).
Elosegui-Artola, A. et al. Force triggers YAP nuclear entry by regulating transport across nuclear pores. Cell 171, 1397–1410 (2017).
Sharma, D. et al. Single-molecule force spectroscopy reveals a mechanically stable protein fold and the rational tuning of its mechanical stability. Proc. Natl Acad. Sci. USA 104, 9278–9283 (2007).
Bornschlögl, T., Christof, J., Gebhardt, M. & Rief, M. Designing the folding mechanics of coiled coils. ChemPhysChem 10, 2800–2804 (2009).
Ng, S. P. et al. Designing an extracellular matrix protein with enhanced mechanical stability. Proc. Natl Acad. Sci. USA 104, 9633–9637 (2007).
Sadler, D. P. et al. Identification of a mechanical rheostat in the hydrophobic core of protein L. J. Mol. Biol. 393, 237–248 (2009).
Cao, Y., Yoo, T., Zhuang, S. & Li, H. Protein–protein interaction regulates proteins’ mechanical stability. J. Mol. Biol. 378, 1132–1141 (2008).
Cao, Y., Yoo, T. & Li, H. Single molecule force spectroscopy reveals engineered metal chelation is a general approach to enhance mechanical stability of proteins. Proc. Natl Acad. Sci. USA 105, 11152–11157 (2008).
Ringer, P. et al. Multiplexing molecular tension sensors reveals piconewton force gradient across talin-1. Nat. Methods 14, 1090–1096 (2017).
Hughes, J. H. & Kumar, S. Synthetic mechanobiology: engineering cellular force generation and signaling. Curr. Opin. Biotechnol. 40, 82–89 (2016).
Pan, Y. et al. Mechanogenetics for the remote and noninvasive control of cancer immunotherapy. Proc. Natl. Acad. Sci. USA 115, 992–997 (2018).
Liu, L. N. et al. Mechanoresponsive stem cells to target cancer metastases through biophysical cues. Sci. Transl. Med. 9, eaan2966 (2017).
Gordon, W. R. et al. Mechanical allostery: evidence for a force requirement in the proteolytic activation of Notch. Dev. Cell 33, 729–736 (2015).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and Notch signaling. Science 340, 991–994 (2013).
Chowdhury, F. et al. Defining single molecular forces required for Notch activation using Nano Yoyo. Nano Lett. 16, 1–20 (2016).
Seo, D. et al. A mechanogenetic toolkit for interrogating cell signaling in space and time. Cell 165, 1507–1518 (2016).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic Notch receptors. Cell 164, 780–791 (2016).
Verdorfer, T. & Gaub, H. E. Ligand binding stabilizes cellulosomal cohesins as revealed by AFM-based single-molecule force spectroscopy. Sci. Rep. 8, 9634 (2018).
Tiyanont, K. et al. Evidence for increased exposure of the Notch1 metalloprotease cleavage site upon conversion to an activated conformation. Structure 19, 546–554 (2011).
Wu, Y. et al. Therapeutic antibody targeting of individual Notch receptors. Nature 464, 1052–1057 (2010).
Aste-Amézaga, M. et al. Characterization of Notch1 antibodies that inhibit signaling of both normal and mutated Notch1 receptors. PLoS ONE 5, e9094 (2010).
Fortini, M. E. & Bilder, D. Endocytic regulation of Notch signaling. Curr. Opin. Genet. Dev. 19, 323–328 (2009).
Schwesinger, F. et al. Unbinding forces of single antibody-antigen complexes correlate with their thermal dissociation rates. Proc. Natl. Acad. Sci. USA 97, 9972–9977 (2000).
Varnum-Finney, B. et al. Immobilization of Notch ligand, δ-1, is required for induction of Notch signaling. J. Cell Sci. 113 Pt 23, 4313–4318 (2000).
De Odrowa̧z Piramowicz, M., Czuba, P., Targosz, M., Burda, K. & Szymoński, M. Dynamic force measurements of avidin-biotin and streptavdin-biotin interactions using AFM. Acta Biochim. Polym. 53, 93–100 (2006).
Weisel, J. W., Shuman, H. & Litvinov, R. I. Protein–protein unbinding induced by force: single-molecule studies. Curr. Opin. Struct. Biol. 13, 227–235 (2003).
Falk, R. et al. Generation of anti-Notch antibodies and their application in blocking Notch signalling in neural stem cells. Methods 58, 69–78 (2012).
Chopra, M. L. et al. Force generation via β-cardiac myosin, titin, and α-actinin drives cardiac sarcomere assembly from cell-matrix adhesions. Dev. Cell 44, 87–96 (2018).
Sakar, M. S. et al. Cellular forces and matrix assembly coordinate fibrous tissue repair. Nat. Commun. 7, 11036 (2016).
Kabadi, A. M. et al. Enhanced MyoD-induced transdifferentiation to a myogenic lineage by fusion to a potent transactivation domain. ACS Synth. Biol. 4, 689–699 (2015).
Balcioglu, H. E., van Hoorn, H., Donato, D. M., Schmidt, T. & Danen, E. H. J. The integrin expression profile modulates orientation and dynamics of force transmission at cell-matrix adhesions. J. Cell Sci. 128, 1316–1326 (2015).
Austen, K. et al. Extracellular rigidity sensing by talin isoform-specific mechanical linkages. Nat. Cell Biol. 17, 1597–1606 (2015).
Li, H., Carrion-Vazquez, M., Oberhauser, A. F., Marszalek, P. E. & Fernandez, J. M. Point mutations alter the mechanical stability of immunoglobulin modules. Nat. Struct. Biol. 7, 1117–1120 (2000).
Polacheck, W. J. & Chen, C. S. Measuring cell-generated forces: a guide to the available tools. Nat. Methods 13, 415–423 (2016).
Yang, Z., Yu, Z., Cai, Y., Du, R. & Cai, L. Engineering of an enhanced synthetic Notch receptor by reducing ligand-independent activation. Commun. Biol. 3, 116 (2020).
Gordon, W. R. et al. Structure of the Notch1-negative regulatory region: implications for normal activation and pathogenic signaling in T-ALL. Blood 113, 4381–4390 (2009).
Rand, M. D. et al. Calcium depletion dissociates and activates heterodimeric Notch receptors. Mol. Cell. Biol. 20, 1825–1835 (2000).
Frith, J. E. et al. Mechanically-sensitive miRNAs bias human mesenchymal stem cell fate via mTOR signalling. Nat. Commun. 9, 257 (2018).
Mangan, S. & Alon, U. Structure and function of the feed-forward loop network motif. Proc. Natl Acad. Sci. USA 100, 11980–11985 (2003).
Greber, D. & Fussenegger, M. An engineered mammalian band-pass network. Nucleic Acids Res. 38, e174 (2010).
Fridy, P. C. et al. A robust pipeline for rapid production of versatile nanobody repertoires. Nat. Methods 11, 1253–1260 (2014).
Langridge, P. D. & Struhl, G. Epsin-dependent ligand endocytosis activates Notch by force. Cell 171, 1383–1396 (2017).
Guo, B., McMillan, B. J. & Blacklow, S. C. Structure and function of the Mind bomb E3 ligase in the context of Notch signal transduction. Curr. Opin. Struct. Biol. 41, 38–45 (2016).
Messa, M. et al. Epsin deficiency impairs endocytosis by stalling the actin-dependent invagination of endocytic clathrin-coated pits. eLife 3, e03311 (2014).
Kaksonen, M. & Roux, A. Mechanisms of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 19, 313–326 (2018).
Serwas, D. et al. Mechanistic insights into actin force generation during vesicle formation from cryo-electron tomography. Dev. Cell 57, 1132–1145 (2022).
Seib, E. & Klein, T. The role of ligand endocytosis in Notch signalling. Biol. Cell 113, 401–418 (2021).
McMillan, B. J. et al. A tail of two sites: a bipartite mechanism for recognition of notch ligands by mind bomb E3 ligases. Mol. Cell 57, 912–924 (2015).
Dengl, S. et al. Hapten-directed spontaneous disulfide shuffling: a universal technology for site-directed covalent coupling of payloads to antibodies. FASEB J. 29, 1763–1779 (2015).
McMahan, J. B., Ngo, J. T. A Genetically encodable and chemically disruptable system for synthetic post-translational modification dependent signaling. Preprint at bioRxiv https://doi.org/10.1101/2022.05.29.493928 (2022).
Meloty-Kapella, L., Shergill, B., Kuon, J., Botvinick, E. & Weinmaster, G. Notch ligand endocytosis generates mechanical pulling force dependent on dynamin, epsins, and actin. Dev. Cell 12, 22 (2012).
Bloom, J. D., Labthavikul, S. T., Otey, C. R. & Arnold, F. H. Protein stability promotes evolvability. Proc. Natl Acad. Sci. USA 103, 5869–5874 (2006).
Nissim, L., Perli, S. D., Fridkin, A., Perez-Pinera, P. & Lu, T. K. Multiplexed and programmable regulation of gene networks with an integrated RNA and CRISPR/Cas toolkit in human cells. Mol. Cell 54, 698–710 (2014).
Sun, P., Enslen, H., Myung, P. S. & Maurer, R. A. Differential activation of CREB by Ca2+/calmodulin-dependent protein kinases type II and type IV involves phosphorylation of a site that negatively regulates activity. Genes Dev. 8, 2527–2539 (1994).
Wang, X. et al. Constructing modular and universal single molecule tension sensor using protein G to study mechano-sensitive receptors. Sci. Rep. 6, 21584 (2016).
Lee, C. K., Wang, Y. M., Huang, L. S. & Lin, S. Atomic force microscopy: determination of unbinding force, off rate and energy barrier for protein–ligand interaction. Micron 38, 446–461 (2007).
Johnson, K. C. & Thomas, W. E. How do we know when single-molecule force spectroscopy really tests single bonds? Biophys. J. 114, 2032–2039 (2018).
Acknowledgements
D.C.S. and A.M.M. were supported through Graduate Research Fellowship awards from the National Science Foundation. J.C.T was supported through a Cross-Disciplinary Fellowship awarded through BUnano (Boston University Nanotechnology Innovation Center). D.C.S and J.C.T. received support through the Boston University training program in Quantitative Biology and Physiology (QBP, NIH grant T32GM008764). D.C.S. was a recipient of a Kilachand Fellowship through the Multicellular Design Program (Boston University). Support for this work was provided through a seed grant from the Center for Multiscale & Translational Mechanobiology (Boston University) and through NIH research grants R35 GM128859 (to J.T.N.) and R01 HL147585 (to J.T.N.). Additional support was provided through the Reidy Family Career Development Professorship at Boston University (to J.T.N.). The schematics in Fig. 4 were created using biorender.com.
Author information
Authors and Affiliations
Contributions
D.C.S, J.C.T., A.M.M. and J.T.N. designed and performed experiments and analyzed the data. D.C.S. and J.T.N. wrote the manuscript. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.T.N. and D.C.S. are inventors on an issued patent (U.S. Patent, 10,858,443) held by the Trustees of Boston University. J.C.T. and A.M.M. declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Sanjay Kumar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods and Supplementary Figs. 1–15.
Supplementary Video 1
Time-lapse imaging of biotin-FITC induced trans-cellular activation. HEK293-FT reporter cells (UAS-H2B-mCherry) expressing anti-FITC receptors with the indicated mechanosensitive domains were cocultured with HEK293-FT antibiotin-TMD-DLL1 sender cells. trans-cellular coupling was induced by treatment with 2 nM biotin-FITC. Cells were imaged live for 24 h. Fluorescence emissions from T2A-BFP (blue, to identify receiver cells) and H2B-mCherry (red, to monitor reporter activity) were captured every 1.5 h. Sender cells were not fluorescently marked (that is, lacked BFP). Top row: merged images from detection of T2A-BFP (blue), H2B-mCherry (red) and transmitted light. Bottom row: emission from H2B-mCherry detection shown in grayscale.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sloas, D.C., Tran, J.C., Marzilli, A.M. et al. Tension-tuned receptors for synthetic mechanotransduction and intercellular force detection. Nat Biotechnol 41, 1287–1295 (2023). https://doi.org/10.1038/s41587-022-01638-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01638-y
This article is cited by
-
DNA-functionalized artificial mechanoreceptor for de novo force-responsive signaling
Nature Chemical Biology (2024)
-
Evidence and therapeutic implications of biomechanically regulated immunosurveillance in cancer and other diseases
Nature Nanotechnology (2024)
-
The meaning of ubiquitylation of the DSL ligand Delta for the development of Drosophila
BMC Biology (2023)