Abstract
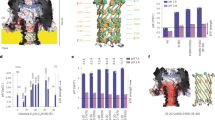
The electrical current blockade of a peptide or protein threading through a nanopore can be used as a fingerprint of the molecule in biosensor applications. However, threading of full-length proteins has only been achieved using enzymatic unfolding and translocation. Here we describe an enzyme-free approach for unidirectional, slow transport of full-length proteins through nanopores. We show that the combination of a chemically resistant biological nanopore, α-hemolysin (narrowest part is ~1.4 nm in diameter), and a high concentration guanidinium chloride buffer enables unidirectional, single-file protein transport propelled by an electroosmotic effect. We show that the mean protein translocation velocity depends linearly on the applied voltage and translocation times depend linearly on length, resembling the translocation dynamics of ssDNA. Using a supervised machine-learning classifier, we demonstrate that single-translocation events contain sufficient information to distinguish their threading orientation and identity with accuracies larger than 90%. Capture rates of protein are increased substantially when either a genetically encoded charged peptide tail or a DNA tag is added to a protein.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data used in this manuscript are available for download at https://figshare.com/s/5cd39ee415c62a316a6f.
Code availability
All data parsing (excluding DTW and GBC) were performed using the Pyth-ion package (https://github.com/wanunulab/Pyth-Ion) and figures were generated using Igor. For analysis, the raw 100 kHz nanopore current data was further low-pass filtered to 10 kHz using the low-pass filter function in Pyth-ion. DTW and GBC analyses were conducted via python scripts written and documented in Jupyter Notebooks, tslearn (v0.5.2)77, SciKit-Learn (v1.0.2)78, and a modified version of the PyPore79 nanopore data analysis library. The Jupyter notebook and associated files are available on GitHub (https://github.com/wanunulab/protein-gd). A detailed description of the DTW and GBC analyses is provided in Supplementary Figs. 26–33, Supplementary Tables 4–7 and Supplementary Notes 3 and 4.
Change history
21 September 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41587-023-01995-2
References
Shendure, J. et al. DNA sequencing at 40: past, present and future. Nature 550, 345–353 (2017).
Ameur, A., Kloosterman, W. P. & Hestand, M. S. Single-molecule sequencing: towards clinical applications. Trends Biotechnol. 37, 72–85 (2019).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133 (2009).
Venkatesan, B. M. & Bashir, R. Nanopore sensors for nucleic acid analysis. Nat. Nanotechnol. 6, 615–624 (2011).
Deamer, D., Akeson, M. & Branton, D. Three decades of nanopore sequencing. Nat. Biotechnol. 34, 518–524 (2016).
Smith, L. M. et al. Proteoform: a single term describing protein complexity. Nat. Methods 10, 186–187 (2013).
Bogaert, A., Fernandez, E. & Gevaert, K. N-terminal proteoforms in human disease. Trends Biochem. Sci. 45, 308–320 (2020).
Tolsma, T. O. & Hansen, J. C. Post-translational modifications and chromatin dynamics. Essays Biochem. 63, 89–96 (2019).
Conibear, A. C. Deciphering protein post-translational modifications using chemical biology tools. Nat. Rev. Chem. 4, 674–695 (2020).
MacCoss, M. J., Alfaro, J., Wanunu, M., Faivre, D. A. & Slavov, N. Sampling the proteome by emerging single-molecule and mass-spectrometry methods. Preprint at arXiv https://doi.org/10.48550/arXiv.2208.00530 (2022).
Slavov, N. Single-cell protein analysis by mass spectrometry. Curr. Opin. Chem. Biol. 60, 1–9 (2021).
Specht, H. & Slavov, N. Transformative opportunities for single-cell proteomics. J. Proteome Res. 17, 2565–2571 (2018).
Specht, H. et al. Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2. Genome Biol. 22, 50 (2021).
Alfaro, J. A. et al. The emerging landscape of single-molecule protein sequencing technologies. Nat. Methods 18, 604–617 (2021).
Zhao, Y. et al. Single-molecule spectroscopy of amino acids and peptides by recognition tunnelling. Nat. Nanotechnol. 9, 466–473 (2014).
Kennedy, E., Dong, Z., Tennant, C. & Timp, G. Reading the primary structure of a protein with 0.07 nm3 resolution using a subnanometre-diameter pore. Nat. Nanotechnol. 11, 968–976 (2016).
Swaminathan, J. et al. Highly parallel single-molecule identification of proteins in zeptomole-scale mixtures. Nat. Biotechnol. 36, 1076–1082 (2018).
van Ginkel, J. et al. Single-molecule peptide fingerprinting. Proc. Natl Acad. Sci. 115, 3338–3343 (2018).
Restrepo-Pérez, L., Joo, C. & Dekker, C. Paving the way to single-molecule protein sequencing. Nat. Nanotechnol. 13, 786–796 (2018).
Stefureac, R., Long, Y.-t, Kraatz, H.-B., Howard, P. & Lee, J. S. Transport of α-helical peptides through α-hemolysin and aerolysin pores. Biochemistry 45, 9172–9179 (2006).
Movileanu, L. Squeezing a single polypeptide through a nanopore. Soft Matter 4, 925–931 (2008).
Rodriguez-Larrea, D. & Bayley, H. Multistep protein unfolding during nanopore translocation. Nat. Nanotechnol. 8, 288–295 (2013).
Rosen, C. B., Bayley, H. & Rodriguez-Larrea, D. Free-energy landscapes of membrane co-translocational protein unfolding. Commun. Biol. 3, 160 (2020).
Payet, L. et al. Thermal unfolding of proteins probed at the single molecule level using nanopores. Anal. Chem. 84, 4071–4076 (2012).
Soni, N., Freundlich, N., Ohayon, S., Huttner, D. & Meller, A. Single-file translocation dynamics of SDS-denatured, whole proteins through sub-5 nm solid-state nanopores. ACS Nano 16, 11405–11414 (2022).
Oukhaled, G. et al. Unfolding of proteins and long transient conformations detected by single nanopore recording. Phys. Rev. Lett. 98, 158101 (2007).
Pastoriza-Gallego, M. et al. Dynamics of unfolded protein transport through an aerolysin pore. J. Am. Chem. Soc. 133, 2923–2931 (2011).
Merstorf, C. et al. Wild type, mutant protein unfolding and phase transition detected by single-nanopore recording. ACS Chem. Biol. 7, 652–658 (2012).
Pastoriza-Gallego, M. et al. Evidence of unfolded protein translocation through a protein nanopore. ACS Nano 8, 11350–11360 (2014).
Cressiot, B. et al. Protein transport through a narrow solid-state nanopore at high voltage: experiments and theory. ACS Nano 6, 6236–6243 (2012).
Keyser, U. F. et al. Direct force measurements on DNA in a solid-state nanopore. Nat. Phys. 2, 473–477 (2006).
Nivala, J., Marks, D. B. & Akeson, M. Unfoldase-mediated protein translocation through an alpha-hemolysin nanopore. Nat. Biotechnol. 31, 247–250 (2013).
Nivala, J., Mulroney, L., Li, G., Schreiber, J. & Akeson, M. Discrimination among protein variants using an unfoldase-coupled nanopore. ACS Nano 8, 12365–12375 (2014).
Yan, S. et al. Single molecule ratcheting motion of peptides in a Mycobacterium smegmatis porin A (MspA) nanopore. Nano Lett. 21, 6703–6710 (2021).
Chen, Z. et al. Controlled movement of ssDNA conjugated peptide through Mycobacterium smegmatis porin A (MspA) nanopore by a helicase motor for peptide sequencing application. Chem. Sci. 12, 15750–15756 (2021).
Brinkerhoff, H., Kang Albert, S. W., Liu, J., Aksimentiev, A. & Dekker, C. Multiple rereads of single proteins at single-amino acid resolution using nanopores. Science 374, 1509–1513 (2021).
Kang, X., Alibakhshi, M. A. & Wanunu, M. One-pot species release and nanopore detection in a voltage-stable lipid bilayer platform. Nano Lett. 19, 9145–9153 (2019).
Yu, L. et al. Stable polymer bilayers for protein channel recordings at high guanidinium chloride concentrations. Biophys. J. 120, 1537–1541 (2021).
Haynes, W. M. CRC Handbook of Chemistry and Physics (CRC Press, 2016).
Perkins, S. J. Protein volumes and hydration effects. Eur. J. Biochem. 157, 169–180 (1986).
Liu, G. P., Topping, T. B., Cover, W. H. & Randall, L. L. Retardation of folding as a possible means of suppression of a mutation in the leader sequence of an exported protein. J. Biol. Chem. 263, 14790–14793 (1988).
Sheshadri, S., Lingaraju, G. M. & Varadarajan, R. Denaturant mediated unfolding of both native and molten globule states of maltose binding protein are accompanied by large deltaCp’s. Protein Sci. 8, 1689–1695 (1999).
Nakane, J., Akeson, M. & Marziali, A. Evaluation of nanopores as candidates for electronic analyte detection. Electrophoresis 23, 2592–2601 (2002).
Meller, A., Nivon, L. & Branton, D. Voltage-driven DNA translocations through a nanopore. Phys. Rev. Lett. 86, 3435–3438 (2001).
Meller, A. & Branton, D. Single molecule measurements of DNA transport through a nanopore. Electrophoresis 23, 2583–2591 (2002).
Hornblower, B. et al. Single-molecule analysis of DNA-protein complexes using nanopores. Nat. Methods 4, 315–317 (2007).
Henrickson, S. E., Misakian, M., Robertson, B. & Kasianowicz, J. J. Driven DNA transport into an asymmetric nanometer-scale pore. Phys. Rev. Lett. 85, 3057–3060 (2000).
Mathé, J., Aksimentiev, A., Nelson, D. R., Schulten, K. & Meller, A. Orientation discrimination of single-stranded DNA inside the α-hemolysin membrane channel. Proc. Natl Acad. Sci. USA 102, 12377–12382 (2005).
Yang, G. et al. Solid-state synthesis and mechanical unfolding of polymers of T4 lysozyme. Proc. Natl Acad. Sci. USA 97, 139 (2000).
Aksimentiev, A. & Schulten, K. Imaging α-hemolysin with molecular dynamics: ionic conductance, osmotic permeability, and the electrostatic potential map. Biophys. J. 88, 3745–3761 (2005).
Doina, P. & Whye, T. Y. (eds.). Soft-DTW: a differentiable loss function for time-series. Proceedings of the 34th International Conference on Machine Learning Vol. 70, pp. 894–903 (PMLR, 2017).
Larkin, J. et al. High-bandwidth protein analysis using solid-state nanopores. Biophys. J. 106, 696–704 (2014).
Ling, D. Y. & Ling, X. S. On the distribution of DNA translocation times in solid-state nanopores: an analysis using Schrödinger’s first-passage-time theory. J. Phys. Condens. Matter 25, 375102 (2013).
Li, J. & Talaga, D. S. The distribution of DNA translocation times in solid-state nanopores. J. Phys. Condens. Matter 22, 454129 (2010).
Talaga, D. S. & Li, J. Single-molecule protein unfolding in solid state nanopores. J. Am. Chem. Soc. 131, 9287–9297 (2009).
Pavlenok, M., Yu, L., Herrmann, D., Wanunu, M. & Niederweis, M. Control of subunit stoichiometry in single-chain MspA nanopores. Biophys. J. https://doi.org/10.1016/j.bpj.2022.01.022 (2022).
Ouldali, H. et al. Electrical recognition of the twenty proteinogenic amino acids using an aerolysin nanopore. Nat. Biotechnol. 38, 176–181 (2020).
Versloot, R. C. A., Straathof, S. A. P., Stouwie, G., Tadema, M. J. & Maglia, G. β-Barrel nanopores with an acidic–aromatic sensing region identify proteinogenic peptides at low pH. ACS Nano https://doi.org/10.1021/acsnano.1c11455 (2022).
Versloot, R. C. A. et al. Quantification of protein glycosylation using nanopores. Nano Lett. 22, 5357–5364 (2022).
Huang, G. et al. PlyAB nanopores detect single amino acid differences in folded haemoglobin from blood. Angew. Chem. Int. Ed. 61, e202206227 (2022).
Noakes, M. T. et al. Increasing the accuracy of nanopore DNA sequencing using a time-varying cross membrane voltage. Nat. Biotechnol. 37, 651–656 (2019).
Muthukumar, M. Polymer translocation through a hole. J. Chem. Phys. 111, 10371–10374 (1999).
Ammenti, A., Cecconi, F., Marini Bettolo Marconi, U. & Vulpiani, A. A statistical model for translocation of structured polypeptide chains through nanopores. J. Phys. Chem. B 113, 10348–10356 (2009).
Phillips, J. C. et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 153, 044130 (2020).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Yoo, J. & Aksimentiev, A. New tricks for old dogs: improving the accuracy of biomolecular force fields by pair-specific corrections to non-bonded interactions. Phys. Chem. Chem. Phys. 20, 8432–8449 (2018).
Miyamoto, S. & Kollman, P. A. Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
Andersen, H. C. Rattle: a ‘velocity’ version of the shake algorithm for molecular dynamics calculations. J. Comput. Phys. 52, 24–34 (1983).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Song, L. et al. Structure of staphylococcal α-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1865 (1996).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Duan, X. & Quiocho, F. A. Structural evidence for a dominant role of nonpolar interactions in the binding of a transport/chemosensory receptor to its highly polar ligands. Biochemistry 41, 706–712 (2002).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Li, H., Robertson, A. D. & Jensen, J. H. Very fast empirical prediction and rationalization of protein pKa values. Proteins Struct. Funct. Bioinf. 61, 704–721 (2005).
Tavenard, R. et al. Tslearn, a machine learning toolkit for time series data. J. Mach. Learn. Res. 21, 1–6 (2020).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Schreiber, J. & Karplus, K. Analysis of nanopore data using hidden Markov models. Bioinformatics 31, 1897–1903 (2015).
Acknowledgements
We thank C. McCormick for assistance with editing the manuscript for clarity, and N. Slavov for helpful discussions regarding protein sequencing. We acknowledge funding from the National Institutes of Health under grants HG011087 (to M.W.) and GM115442 (to M.C.), and the National Science Foundation under grant PHY-1430124 (to A.A.). The supercomputer time was provided through the XSEDE allocation grant (MCA05S028) and the Leadership Resource Allocation MCB20012 on Frontera of the Texas Advanced Computing Center.
Author information
Authors and Affiliations
Contributions
L.Y. and M.W. conceived the project and designed the experiments. L.Y. developed the experimental protocol, carried out the experiments and analyzed the protein translocation data. X.K. and L.Y. fabricated the SU-8 wedge-on-pillar chips used in the experiments. F.L. and M.C. prepared and purified the protein samples. B.M. and A.A. designed and conducted the MD simulations. A.M. and A.F. performed the data analysis described in ‘protein-specific current signals’ (SDTW and GBC) and wrote the respective sections of the manuscript. J.C.F. performed the DNA–protein conjugation experiments. L.Y., B.M., A.A. and M.W. wrote the first draft of the manuscript, and all authors commented and edited it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Jeff Nivala and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–33, Supplementary notes 1–4 and Supplementary Tables 1–7.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yu, L., Kang, X., Li, F. et al. Unidirectional single-file transport of full-length proteins through a nanopore. Nat Biotechnol 41, 1130–1139 (2023). https://doi.org/10.1038/s41587-022-01598-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01598-3
This article is cited by
-
Engineered nanopores for exopeptidase protein sequencing
Nature Methods (2024)
-
Real-time detection of 20 amino acids and discrimination of pathologically relevant peptides with functionalized nanopore
Nature Methods (2024)
-
Full-length single-molecule protein fingerprinting
Nature Nanotechnology (2024)
-
Nanopore DNA sequencing technologies and their applications towards single-molecule proteomics
Nature Chemistry (2024)
-
Enzyme-less nanopore detection of post-translational modifications within long polypeptides
Nature Nanotechnology (2023)