Abstract
Programmable genome integration of large, diverse DNA cargo without DNA repair of exposed DNA double-strand breaks remains an unsolved challenge in genome editing. We present programmable addition via site-specific targeting elements (PASTE), which uses a CRISPR–Cas9 nickase fused to both a reverse transcriptase and serine integrase for targeted genomic recruitment and integration of desired payloads. We demonstrate integration of sequences as large as ~36 kilobases at multiple genomic loci across three human cell lines, primary T cells and non-dividing primary human hepatocytes. To augment PASTE, we discovered 25,614 serine integrases and cognate attachment sites from metagenomes and engineered orthologs with higher activity and shorter recognition sequences for efficient programmable integration. PASTE has editing efficiencies similar to or exceeding those of homology-directed repair and non-homologous end joining-based methods, with activity in non-dividing cells and in vivo with fewer detectable off-target events. PASTE expands the capabilities of genome editing by allowing large, multiplexed gene insertion without reliance on DNA repair pathways.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw reads for RNA sequencing and the atgRNA efficiency screen are available at Sequence Read Archive under BioProject accession number PRJNA700575 (ref. 78). Expression plasmids are available from Addgene at https://www.addgene.org/browse/article/28223250/ under UBMTA. The human genome GRCh38 can be accessed at https://www.ncbi.nlm.nih.gov/assembly/GCF_000001405.26/. Source data are provided with this paper.
Code availability
Code to predict atgRNA efficiency and support information is available at https://github.com/abugoot-lab/atgRNA_rank79.
References
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Wright, A. V., Nuñez, J. K. & Doudna, J. A. Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 164, 29–44 (2016).
Nami, F. et al. Strategies for in vivo genome editing in nondividing cells. Trends Biotechnol. 36, 770–786 (2018).
Suzuki, K. et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Rouet, P., Smih, F. & Jasin, M. Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease. Mol. Cell. Biol. 14, 8096–8106 (1994).
Chapman, J. R., Taylor, M. R. G. & Boulton, S. J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Geisinger, J. M. & Stearns, T. CRISPR/Cas9 treatment causes extended TP53-dependent cell cycle arrest in human cells. Nucleic Acids Res. 48, 9067–9081 (2020).
Wang, H. et al. Development of a self-restricting CRISPR–Cas9 system to reduce off-target effects. Mol. Ther. Methods Clin. Dev. 18, 390–401 (2020).
Kanca, O. et al. An efficient CRISPR-based strategy to insert small and large fragments of DNA using short homology arms. eLife 8, e51539 (2019).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Anzalone, A. V. et al. Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing. Nat. Biotechnol. 40, 731–740 (2021).
Wang, J. et al. Efficient targeted insertion of large DNA fragments without DNA donors. Nat. Methods 19, 331–340 (2022).
Ivics, Z., Hackett, P. B., Plasterk, R. H. & Izsvák, Z. Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells. Cell 91, 501–510 (1997).
Brown, W. R. A., Lee, N. C. O., Xu, Z. & Smith, M. C. M. Serine recombinases as tools for genome engineering. Methods 53, 372–379 (2011).
Calos, M. P. The C31 integrase system for gene therapy. Curr. Gene Ther. 6, 633–645 (2006).
Mulholland, C. B. et al. A modular open platform for systematic functional studies under physiological conditions. Nucleic Acids Res. 43, e112 (2015).
Ehrhardt, A., Engler, J. A., Xu, H., Cherry, A. M. & Kay, M. A. Molecular analysis of chromosomal rearrangements in mammalian cells after øC31-mediated integration. Hum. Gene Ther. 17, 1077–1094 (2006).
Liu, J., Jeppesen, I., Nielsen, K. & Jensen, T. G. Phi c31 integrase induces chromosomal aberrations in primary human fibroblasts. Gene Ther. 13, 1188–1190 (2006).
Kovač, A. et al. RNA-guided retargeting of Sleeping Beauty transposition in human cells. eLife 9, e53868 (2020).
Ma, S. et al. Enhancing site-specific DNA integration by a Cas9 nuclease fused with a DNA donor-binding domain. Nucleic Acids Res. 48, 10590–10601 (2020).
Chen, S. P. & Wang, H. H. An engineered Cas–transposon system for programmable and site-directed DNA transpositions. CRISPR J. 2, 376–394 (2019).
Bhatt, S. & Chalmers, R. Targeted DNA transposition in vitro using a dCas9–transposase fusion protein. Nucleic Acids Res. 47, 8126–8135 (2019).
Hew, B. E., Sato, R., Mauro, D., Stoytchev, I. & Owens, J. B. RNA-guided piggyBac transposition in human cells. Synth. Biol. 4, ysz018 (2019).
Chaikind, B., Bessen, J. L., Thompson, D. B., Hu, J. H. & Liu, D. R. A programmable Cas9–serine recombinase fusion protein that operates on DNA sequences in mammalian cells. Nucleic Acids Res. 44, 9758–9770 (2016).
Akopian, A., He, J., Boocock, M. R. & Stark, W. M. Chimeric recombinases with designed DNA sequence recognition. Proc. Natl Acad. Sci. USA 100, 8688–8691 (2003).
Gordley, R. M., Smith, J. D., Gräslund, T. & Barbas, C. F. III Evolution of programmable zinc finger-recombinases with activity in human cells. J. Mol. Biol. 367, 802–813 (2007).
Mercer, A. C., Gaj, T., Fuller, R. P. & Barbas, C. F. III Chimeric TALE recombinases with programmable DNA sequence specificity. Nucleic Acids Res. 40, 11163–11172 (2012).
Gersbach, C. A., Gaj, T., Gordley, R. M., Mercer, A. C. & Barbas, C. F. III Targeted plasmid integration into the human genome by an engineered zinc-finger recombinase. Nucleic Acids Res. 39, 7868–7878 (2011).
Prorocic, M. M. et al. Zinc-finger recombinase activities in vitro. Nucleic Acids Res. 39, 9316–9328 (2011).
Gordley, R. M., Gersbach, C. A. & Barbas, C. F. III Synthesis of programmable integrases. Proc. Natl Acad. Sci. USA 106, 5053–5058 (2009).
Xu, Z. et al. Accuracy and efficiency define Bxb1 integrase as the best of fifteen candidate serine recombinases for the integration of DNA into the human genome. BMC Biotechnol. 13, 87 (2013).
Kay, M. A., He, C. -Y. & Chen, Z. -Y. A robust system for production of minicircle DNA vectors. Nat. Biotechnol. 28, 1287–1289 (2010).
Dang, Y. et al. Optimizing sgRNA structure to improve CRISPR–Cas9 knockout efficiency. Genome Biol. 16, 280 (2015).
Oscorbin, I. P., Wong, P. F. & Boyarskikh, U. A. The attachment of a DNA‐binding Sso7d‐like protein improves processivity and resistance to inhibitors of M‐MuLV reverse transcriptase. FEBS Lett. 594, 4338–4356 (2020).
Ghosh, P., Kim, A. I. & Hatfull, G. F. The orientation of mycobacteriophage Bxb1 integration is solely dependent on the central dinucleotide of attP and attB. Mol. Cell 12, 1101–1111 (2003).
Sun, D. et al. A functional genetic toolbox for human tissue-derived organoids. eLife 10, e67886 (2021).
Keravala, A. et al. A diversity of serine phage integrases mediate site-specific recombination in mammalian cells. Mol. Genet. Genomics 276, 135–146 (2006).
Singh, S., Ghosh, P. & Hatfull, G. F. Attachment site selection and identity in Bxb1 serine integrase-mediated site-specific recombination. PLoS Genet. 9, e1003490 (2013).
Zhang, Q., Azarin, S. M. & Sarkar, C. A. Model-guided engineering of DNA sequences with predictable site-specific recombination rates. Nat. Commun. 13, 4152 (2022).
Jiang, T., Zhang, X. -O., Weng, Z. & Xue, W. Deletion and replacement of long genomic sequences using prime editing. Nat. Biotechnol. 40, 227–234 (2021).
Choi, J. et al. Precise genomic deletions using paired prime editing. Nat. Biotechnol. 40, 218–226 (2022).
Jusiak, B. et al. Comparison of integrases identifies Bxb1-GA mutant as the most efficient site-specific integrase system in mammalian cells. ACS Synth. Biol. 8, 16–24 (2019).
Schwinn, M. K. et al. CRISPR-mediated tagging of endogenous proteins with a luminescent peptide. ACS Chem. Biol. 13, 467–474 (2018).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Schnepp, B. C., Jensen, R. L., Chen, C. -L., Johnson, P. R. & Clark, K. R. Characterization of adeno-associated virus genomes isolated from human tissues. J. Virol. 79, 14793–14803 (2005).
Wold, W. S. M. & Toth, K. Adenovirus vectors for gene therapy, vaccination and cancer gene therapy. Curr. Gene Ther. 13, 421–433 (2013).
Wesselhoeft, R. A., Kowalski, P. S. & Anderson, D. G. Engineering circular RNA for potent and stable translation in eukaryotic cells. Nat. Commun. 9, 2629 (2018).
Azuma, H. et al. Robust expansion of human hepatocytes in Fah–/–/Rag2–/–/Il2rg–/– mice. Nat. Biotechnol. 25, 903–910 (2007).
Bateman, A. et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2020).
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science 365, 48–53 (2019).
Klompe, S. E., Vo, P. L. H., Halpin-Healy, T. S. & Sternberg, S. H. Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration. Nature 571, 219–225 (2019).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
Maeder, M. L. et al. Development of a gene-editing approach to restore vision loss in Leber congenital amaurosis type 10. Nat. Med. 25, 229–233 (2019).
Mackay, D. S. et al. Screening of a large cohort of Leber congenital amaurosis and retinitis pigmentosa patients identifies novel LCA5 mutations and new genotype–phenotype correlations. Hum. Mutat. 34, 1537–1546 (2013).
Marson, F. A. L., Bertuzzo, C. S. & Ribeiro, J. D. Classification of CFTR mutation classes. Lancet Respir. Med. 4, e36 (2016).
Eyquem, J. et al. Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection. Nature 543, 113–117 (2017).
Tareen, A. & Kinney, J. B. Logomaker: beautiful sequence logos in Python. Bioinformatics 36, 2272–2274 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Picelli, S., Åsa K., Björklund, A.K., Reinius, B., Sagasser, S. Winberg, G. and Sandberg, R. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5–W9 (2008).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Sena-Esteves, M. & Gao, G. Introducing genes into mammalian cells: viral vectors. Cold Spring Harb. Protoc. 2020, 095513 (2020).
Su, Q., Sena-Esteves, M. & Gao, G. Release of the cloned recombinant adenovirus genome for rescue and expansion. Cold Spring Harb. Protoc. 2019, https://doi.org/10.1101/pdb.prot095539 (2019).
Su, Q., Sena-Esteves, M. & Gao, G. Purification of the recombinant adenovirus by cesium chloride gradient centrifugation. Cold Spring Harb. Protoc. 2019, https://doi.org/10.1101/pdb.prot095547 (2019).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Roux, S., Enault, F., Hurwitz, B. L. & Sullivan, M. B. VirSorter: mining viral signal from microbial genomic data. PeerJ 3, e985 (2015).
Durrant, M. G., Li, M. M., Siranosian, B. A., Montgomery, S. B. & Bhatt, A. S. A bioinformatic analysis of integrative mobile genetic elements highlights their role in bacterial adaptation. Cell Host Microbe 28, 140–153 (2020).
Yarnall, M. T. N. et al. Genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. SRA https://www.ncbi.nlm.nih.gov/bioproject/PRJNA700575/ (2022).
Yarnall, M. T. N. et al. Genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. GitHub https://github.com/abugoot-lab/atgRNA_rank (2022).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
McCarthy, D. J. & Smyth, G. K. Testing significance relative to a fold-change threshold is a TREAT. Bioinformatics 25, 765–771 (2009).
Acknowledgements
We would like to thank B. Desimone, F. Chen, J. Joung, A. Serj-Hansen, G. Feng, J. Wilde, M. Calos, T. Aida, Y. Cha and M. Mittens for helpful discussions, E.V. Koonin and K. Makarova for helpful discussions with integrase discovery and annotation, P. Reginato, D. Weston and E. Boyden for MiSeq instrumentation, S. Jacobs and A. Ainbinder for ddPCR instrumentations, S. Bhatia and S. March Riera for hepatocyte assistance, G. Paradis and M. Griffin for flow cytometry assistance and J. Crittenden for editing the manuscript. L.V. is supported by a Swiss National Science Foundation Postdoc Mobility Fellowship. O.O.A. and J.S.G. are supported by NIH grants 1R21-AI149694, R01-EB031957 and R56-HG011857, The McGovern Institute Neurotechnology program, the K. Lisa Yang and Hock E. Tan Center for Molecular Therapeutics in Neuroscience, G. Harold & Leila Y. Mathers Charitable Foundation, MIT John W. Jarve (1978) Seed Fund for Science Innovation, Impetus Grants, Cystic Fibrosis Foundation Pioneer Grant, Google Ventures, FastGrants, the Harvey Family Foundation and the McGovern Institute. S.K.G. was supported by the Intramural Research Program of the National Library of Medicine, NIH.
Author information
Authors and Affiliations
Contributions
O.O.A. and J.S.G. conceived the study. O.O.A. and J.S.G. designed and participated in all experiments. M.T.N.Y., E.I.I. and C.S.-U. led many of the experiments and assay readouts. R.N.K. helped with cell culture, cloning, plasmid sequencing, NGS and in vivo experiments. M.T.N.Y. and C.S.-U. helped with ddPCR, sequencing experiments and cloning. L.V. helped with various PASTE editing experiments and characterization of integrases. W.Z. synthesized mRNA and performed the electroporation experiments. J.L. and S.K.G. performed the computational mining to uncover integrases and annotated these new systems. K.J. performed the ML modeling of the pooled atgRNA screening and developed a guide design software package. N.R., L.Z., K.H., J.A.W, A.P.K., A.E.Z. and C.A.V. synthesized synthetic guides and advised on synthetic RNA experiments. J.M.H. and A.U. provided select mRNA constructs and advised on mRNA experiments. H.M., J.X. and G.G. produced AAV and AdV. S.K.D., Y.M. and D.R.R. provided primary human hepatocytes and advice for in vivo experiments with humanized mouse models. L.F. and G.B. provided humanized liver mice, managed in vivo injections and collections and advised on the in vivo aspects of the project. O.O.A. and J.S.G. wrote the manuscript with help from all authors.
Corresponding authors
Ethics declarations
Competing interests
O.O.A., J.S.G., J.L., L.V. and K.J. are co-inventors on patent applications filed by Massachusetts Institute of Technology relating to work in this manuscript. O.O.A. and J.S.G. are cofounders of Sherlock Biosciences, Proof Diagnostics, Moment Biosciences and Tome Biosciences. O.O.A. and J.S.G. were advisors for Beam Therapeutics during the course of this project. K.H., J.A.W., A.P.K. and A.E.Z. are employees and shareholders of Synthego. S.K.D., Y.M. and D.R.R. are employees of PhoenixBio. L.F. and G.B. are employees of Yecuris Corporation. N.R., L.Z. and C.A.V. are employees of Integrated DNA Technologies. The remaining authors declare no competing interest.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Evaluation of prime integration activity for diverse attB sequences and optimization of PASTE editing through dosage and mutagenesis.
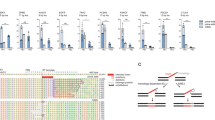
a) Prime editing efficiency for the insertion of different length BxbINT attB sites at ACTB. Data are mean (n = 2 or 3) ± s.e.m. b) Prime editing efficiency for this insertion of a BxbINT attB site at ACTB with targeting and non-targeting guides. Data are mean (n = 3) ± s.e.m. c) Prime editing efficiency for the insertion of different integrases’ attB sites at ACTB. Both orientations of landing sites are profiled (F, forward; R, reverse). Data are mean (n = 3) ± s.e.m. d) PASTE editing efficiency for the insertion of EGFP at ACTB with and without a nicking guide. Data are mean (n = 3) ± s.e.m. e) PASTE integration efficiency of EGFP at ACTB measured with different doses of a single-vector delivery of components. Data are mean (n = 2 or 3) ± s.e.m. f) PASTE integration efficiency of EGFP at ACTB measured with different ratios of a single-vector delivery of components to the EGFP template vector. Data are mean (n = 3) ± s.e.m. g) PASTE efficiency at the ACTB target compared between atgRNAs containing either the v1 or v2 scaffold designs. Data are mean (n = 3) ± s.e.m. h) PASTE integration efficiency of EGFP at ACTB with different RT domain fusions. Data are mean (n = 2 or 3) ± s.e.m. i) PASTE integration efficiency of EGFP at ACTB with different RT domain fusions and linkers. Data are mean (n = 2 or 3) ± s.e.m. j) PASTE integration efficiency of EGFP at ACTB with mutant RT domains. Data are mean (n = 3) ± s.e.m. k) Optimization of PASTE constructs with a panel of linkers and RT modifications for Gluc integration at the ACTB locus using atgRNAs with the v2 scaffold. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 2 Characterization of PASTE payload sizes and integration junctions.
a) PASTE integration efficiency at the ACTB locus of varying sized cargos transfected at a fixed DNA amount and variable molar ratio. b) PASTE integration efficiency at the ACTB locus of varying sized cargos transfected at a variable DNA amounts. c) Schematic of PASTE integration, including resulting attB and attL sites that are generated and PCR primers for assaying the integration junctions. d) PCR and gel electrophoresis readout of left integration junction from PASTE insertion of GFP at the ACTB locus. Insertion is analyzed for in-frame and out-of-frame GFP integration experiments as well as for a no prime control. Expected sizes of the PCR fragments are shown using the primers shown in the schematic in subpanel A. e) PCR and gel electrophoresis readout of right integration junction from PASTE insertion of GFP at the ACTB locus. Insertion is analyzed for in-frame and out-of-frame GFP integration experiments as well as for a no prime control. Expected sizes of the PCR fragments are shown using the primers shown in the schematic in subpanel A. f) Sanger sequencing shown for the right integration junction for an in-frame fusion of GFP via PASTE to the N-terminus of ACTB. g) Sanger sequencing shown for the left integration junction for an in-frame fusion of GFP via PASTE to the N-terminus of β-actin. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 3 Validation of design rules for efficient PASTE insertion at endogenous genomic loci.
a) Schematic of various parameters that affect PASTE integration of ~1 kb GFP insert. On the atgRNA, the PBS, RT, and attB lengths can alter the efficiency of AttB insertion. Nicking guide selection also affects overall gene integration efficiency. b) The impact of PBS and RT length on PASTE integration of GFP at the ACTB locus. c) The impact of PBS and RT length on PASTE integration of GFP at the LMNB1 locus. d) The impact of attB length on PASTE integration of GFP at the ACTB locus. e) The impact of attB length on PASTE integration of GFP at the LMNB1 locus. f) The impact of attB length on PASTE integration of GFP at the NOLC1 locus. g) The impact of minimal PBS, RT, and attB lengths on PASTE integration efficiency of GFP at the ACTB locus. h) The impact of minimal PBS, RT, and attB lengths on PASTE integration efficiency of GFP at the LMNB1 locus. i) PASTE integration efficiency of EGFP at varying endogenous loci. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 4 Heatmaps depicting the effect of PBS, RT, and attB lengths on atgRNA efficiency of attachment site insertion from high-throughput pooled screening of 10,580 guides targeting a variety of loci.
Bar charts indicating normalized summation across relevant PBS, RT, or attB parameter axes are shown on heatmap sides.
Extended Data Fig. 5 Effect of nicking guides on insertion of diverse cargos.
a) PASTE insertion efficiency at ACTB and LMNB1 loci with two different nicking guide designs. b) Attachment site insertion at the SERPINA1 locus with a panel of different nicking guides at varying distances. c) Effect of nicking guides on PASTE integration efficiency at the LMNB1 locus with two different atgRNA designs. d) PASTE integration efficiency at ACTB and LMNB1 with target and non-targeting spacers and matched atgRNAs with and without BxbINT expression. e) Integration of a panel of different gene cargo at LMNB1 locus via PASTE. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 6 Further characterization of PASTE specificity and effects on cellular transcriptome.
a) Comparison of indel rates generated by PASTE and HITI mediated insertion of EGFP at the ACTB and LMNB1 loci in HepG2 cells. b) Effect of attB site integration on protein production. Samples treated with either ACTB, LMNB1 non-targeting guides were harvest and analyzed for protein expression by Western blot. Quantified band intensities relative to GAPDH controls are shown below samples. c) GFP integration activity at predicted BxbINT and PASTE ACTB Cas9 guide off-target sites in the human genome. d) GFP integration activity at predicted HITI ACTB Cas9 guide off-target sites. e) Validation of ddPCR assays for detecting editing at predicted BxbINT off-target sites using synthetic amplicons. f) Validation of ddPCR assays for detecting editing at predicted PASTE ACTB Cas9 guide off-target sites using synthetic amplicons. g) Validation of ddPCR assays for detecting editing at predicted HITI ACTB Cas9 guide off-target sites using synthetic amplicons. h) Analysis of on-target and off-target integration events across 3 single-cell clones for PASTE and 3 single-cell clones for no prime condition. i) Volcano plots depicting the fold expression change of sequenced mRNAs versus significance (p-value). Each dot represents a unique mRNA transcript and significant transcripts are shaded according to either upregulation (red) or downregulation (blue). Fold expression change is measured against ACTB-targeting guide-only expression (including cargo). Significance is determined by moderated t-statistic80 adjusted for a log-fold cut off of 0.58581. j) Top significantly upregulated and downregulated genes for BxbINT-only conditions. Genes are shown with their corresponding Z-scores of counts per million (cpm) for BxbINT only expression, GFP-only expression, PASTE targeting ACTB for EGFP insertion, Prime targeting ACTB for EGFP expression without BxbINT, and guide/cargo only. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 7 Additional characterization of attP mutants for improved editing and multiplexing.
a) Integration efficiencies of wildtype and mutant attP sites with PASTE at the ACTB locus. b) attP single mutants are characterized for PASTE EGFP integration at the ACTB locus. c) Relative enrichment values (calculated as ratio of integrated reads to total reads) for the wildtype Bxb1 and top 5 mutants from the mutagenesis screen d) Comparison of integration efficiency between PASTEv3 and Twin-PE integration at the ACTB locus, with both single atgRNA (46 bp) or dual atgRNA with PASTE-Replace (38 bp). e) Comparison of integration efficiency and residual attB formation between PASTEv3 with PASTE-Replace and Twin-PE integration at the NOLC1 locus with dual atgRNAs containing either a 46 bp or 42 bp attB sequence. f) Comparison of integration efficiency and residual attB formation between PASTEv3 with PASTE-Replace and Twin-PE integration at the CCR5 locus with dual atgRNAs containing a 38 bp attB sequence. g) Comparison of residual attB formation between PASTEv3 with PASTE-Replace and Twin-PE integration at the ACTB locus. h) Characterization of integration of a 5 kb payload at the ACTB locus with all 16 possible dinucleotides for attB/attP pairs between the atgRNA and minicircle. i) Schematic of the pooled attB/attP dinucleotide orthogonality assay. Each attB dinucleotide sequence is co-transfected with a barcoded pool of all 16 attP dinucleotide sequences and BxbINT, and relative integration efficiencies are determined by next generation sequencing of barcodes. All 16 attB dinucleotides are profiled in an arrayed format with attP pools. j) Relative insertion preferences for all possible attB/attP dinucleotide pairs determined by the pooled orthogonality assay. k) Orthogonality of BxbINT dinucleotides as measured by a pooled reporter assay. Each web logo motif shows the relative integration of different attP sequences in a pool at a denoted attB sequence with the listed dinucleotide. l) Representative fluorescence images of multiplexed PASTE gene tagging of ACTB, LMNB1, and NOLC1. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 8 Therapeutic applications of PASTE and further characterization of integrases.
a) Schematic of protein production assay for PASTE-integrated transgene. SERPINA1 and CPS1 transgenes are tagged with HIBIT luciferase for readout with both ddPCR and luminescence. b) Integration efficiency of SERPINA1 and CPS1 transgenes in HEK293FT cells at the ACTB locus. c) Integration efficiency of SERPINA1 and CPS1 transgenes in HepG2 cells at the ACTB locus. d) Intracellular levels of SERPINA1-HIBIT and CPS1-HIBIT in HepG2 cells. e) Secreted levels of SERPINA1-HIBIT and CPS1-HIBIT in HepG2 cells. f) Integration of SERPINA1 and CPS1 genes that are HIBIT tagged as measured by a protein expression luciferase assay. g) Integration of SERPINA1 and CPS1 genes that are HIBIT tagged as measured by a protein expression luciferase assay normalized to a standardized HIBIT ladder, enabling accurate quantification of protein levels. h) PASTE integration activity with most active integrases compared to BxbINT. i) Characterization of integrase activity on truncated attachment sites using integrase reporters in HEK293FT cells. j) PASTE integration activity with computationally selected integrases with shorter attB sites. Data are mean (n = 3) ± s.e.m.
Extended Data Fig. 9 Evaluation of viral templates for PASTE and characterization of editing in non-dividing cells.
a) Schematic of PASTE performance in the presence of cell cycle inhibition. Cells are transfected with plasmids for insertion with PASTE or Cas9-induced HDR and treated with aphidicolin to arrest cell division. Efficiency of PASTE and HDR are read out with ddPCR or amplicon sequencing, respectively. b) Editing efficiency of single mutations by HDR at EMX1 locus with two Cas9 guides in the presence or absence of cell division read out with amplicon sequencing. Data are mean (n = 3) ± s.e.m. c) HDR mediated editing of the EMX1 locus is significantly diminished in non-dividing HEK293FT cells blocked by 5 µM aphidicolin treatment. Data are mean (n = 3) ± s.e.m. d) Integration efficiency of various sized GFP inserts up to 13.3 kb at the ACTB locus with PASTE in the presence or absence of cell division. Data are mean (n = 3) ± s.e.m. e) Effect of insert minicircle DNA amount on PASTE-mediated insertion at the ACTB locus in dividing and non-dividing HEK293FT cells blocked by 5 µM aphidicolin treatment. Data are mean (n = 3) ± s.e.m. f) PASTE efficiency of EGFP integration at the ACTB locus in K562 cells. Data are mean (n = 3) ± s.e.m. g) Insertion templates delivered via AAV transduction. Templates were co-delivered via AAV dosing at levels indicated. Data are mean (n = 3) ± s.e.m. h) PASTE integration of GFP at the ACTB locus with the GFP template delivered via AAV in HEK293FT cells. i) PASTE integration of GFP at the ACTB locus with the GFP template delivered via AAV at different doses in HEK293FT cells. Data are mean (n = 3) ± s.e.m. j) Integration efficiency of AdV delivery of integrase, guides, and cargo in HEK293FT and HepG2 cells. BxbINT and guide RNAs or cargo were delivered either via plasmid transfection (Pl), AdV transduction (AdV), or omitted (-). SpCas9-RT was only delivered as plasmid or omitted. Data are mean (n = 3) ± s.e.m. k) Delivery of PASTE system components with mRNA and synthetic guides, paired with either AdV or plasmid cargo. Data are mean (n = 3) ± s.e.m. l) Attachment site insertion efficiency at the LMNB1 locus using PASTE delivered as mRNA with synthetic atgRNA and nicking guides. Data are mean (n = 3) ± s.e.m. m) Integration efficiency at the LMNB1 locus using PASTE delivered as mRNA (Trilink versions), synthetic atgRNA and nicking guides, and adenoviral delivered EGFP cargo. All conditions contain full length PASTE mRNA and are optionally supplemented with additional Bxb1 mRNA as indicated. Data are mean (n = 2) ± s.e.m.
Extended Data Fig. 10 Additional characterization of in vivo liver editing with PASTE.
a) PASTE integration using delivery of circular mRNA with synthetic guides and either AdV or plasmid cargo. Data are mean (n = 3) ± s.e.m. b) PASTE integration of GFP at the ACTB locus with dose titration of PASTE components and GFP cargo delivered as AdV in HepG2 cells. Data are mean (n = 3) ± s.e.m. c) Evaluation of a 3-primer NGS assay for measuring integration efficiency, akin to junctional readouts by ddPCR. Using amplicon standards mixed at predefined ratios (x-axis), we can ascertain the accuracy of the measured editing (y-axis) by NGS. d) Analysis of primary human hepatocyte (PXB-cells®) EGFP integration at the ACTB locus using adenoviral delivery for PASTEv1 and guides and AAV for the EGFP template. Viral doses are as indicated. Shown is mean ± s.e.m with n = 2. e) Analysis of all liver editing outcomes for adenoviral EGFP template integration at the ACTB locus using PASTE in vivo. f) Analysis of attB site insertion efficiency at the ACTB locus using PASTE in vivo. Data are mean (n = 8). g) Analysis of adenoviral EGFP template integration efficiency into available attB sites at the ACTB locus using PASTE in vivo. Data are mean (n = 8). h) Analysis of indel frequency at the ACTB locus using PASTE in vivo. Data are mean (n = 8). i) Analysis of attB-site associated indels during in vivo integration with PASTE via alignment of representative reads to the ACTB locus containing the desired attB site.
Supplementary information
Supplementary Information
Supplementary Tables 1–11.
Supplementary Data 1
Pooled atgRNA screening read counts for different atgRNA sequences.
Supplementary Data 2
Editing results for different atgRNAs in the pooled screen.
Supplementary Data 3
List of computationally identified serine integrases and predicted attB/attP sequences.
Source data
Source Data Fig. 1
Unprocessed nucleic acid gels and western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yarnall, M.T.N., Ioannidi, E.I., Schmitt-Ulms, C. et al. Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. Nat Biotechnol 41, 500–512 (2023). https://doi.org/10.1038/s41587-022-01527-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01527-4
This article is cited by
-
HiHo-AID2: boosting homozygous knock-in efficiency enables robust generation of human auxin-inducible degron cells
Genome Biology (2024)
-
BacPE: a versatile prime-editing platform in bacteria by inhibiting DNA exonucleases
Nature Communications (2024)
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)
-
Targeted DNA integration in human cells without double-strand breaks using CRISPR-associated transposases
Nature Biotechnology (2024)
-
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Nature Communications (2024)