Abstract
Children have reduced severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection rates and a substantially lower risk for developing severe coronavirus disease 2019 compared with adults. However, the molecular mechanisms underlying protection in younger age groups remain unknown. Here we characterize the single-cell transcriptional landscape in the upper airways of SARS-CoV-2-negative (n = 18) and age-matched SARS-CoV-2-positive (n = 24) children and corresponding samples from adults (n = 44), covering an age range of 4 weeks to 77 years. Children displayed higher basal expression of relevant pattern recognition receptors such as MDA5 (IFIH1) and RIG-I (DDX58) in upper airway epithelial cells, macrophages and dendritic cells, resulting in stronger innate antiviral responses upon SARS-CoV-2 infection than in adults. We further detected distinct immune cell subpopulations including KLRC1 (NKG2A)+ cytotoxic T cells and a CD8+ T cell population with a memory phenotype occurring predominantly in children. Our study provides evidence that the airway immune cells of children are primed for virus sensing, resulting in a stronger early innate antiviral response to SARS-CoV-2 infection than in adults.
Similar content being viewed by others
Main
It has repeatedly been reported that younger individuals have a substantially lower risk for developing coronavirus disease 2019 (COVID-19), despite a similar risk of infection, as reflected in dramatically increased mortality with increasing age1,2,3. These observations suggest that children may have a higher capability of controlling SARS-CoV-2 infection. It has been shown that an early cell-intrinsic innate immune response, mediated by pattern recognition receptors (PRRs) and the type I and III interferon (IFN) system, is crucial for the successful control of SARS-CoV-2 infection4. In line with these observations, recent studies compared adults and children with severe COVID-19 or those presenting to an emergency department and described an impaired IFN response in pediatric COVID-19 (refs. 5,6). However, the molecular mechanisms protecting against COVID-19 in younger age groups, particularly in those with no or only mild/moderate symptoms, remain unknown.
To understand the higher capacity of children for controlling SARS-CoV-2 infection at an early stage, we systematically characterized the transcriptional landscape of upper airways, an airway region with high susceptibility for SARS-CoV-2 infection7, in SARS-CoV-2-negative and SARS-CoV-2-positive children and adults.
Results
Different cellular composition in the upper airways of children and adults
We included study participants of three different COVID-19 cohorts: the RECAST study focusing on COVID-19 in children and their families, the Pa-COVID-19 study and the SC2 study8,9, including SARS-CoV-2-negative and SARS-CoV-2-positive children (n = 42) and adults (n = 44). The derived dataset comprised 268,745 cells in total (Fig. 1a). Samples from the upper airways (nose) were collected from individuals aged 4 weeks to 77 years with a positive SARS-CoV-2 PCR result along with age-matched SARS-CoV-2-negative controls (Supplementary Tables 1 and 2). Focusing on early infection, only mild/moderate COVID-19 cases were considered for this study (Fig. 1a). On the basis of the single-cell RNA sequencing (scRNA-seq) data, we identified 33 different cell types or states in the upper respiratory tract of these individuals, including 21 immune and 12 epithelial cell subtypes (Fig. 1b and Extended Data Fig. 1a,b). We observed striking differences between the pediatric and adult study participants regarding the composition of the immune cell and epithelial cell compartment in the nasal mucosa. While immune cells were rarely detected in nasal samples from healthy adults, samples from SARS-CoV-2-negative children contained high amounts of almost each immune cell subset with an overall dominance of neutrophils (Fig. 1b,c and Extended Data Fig. 2a). In adults, SARS-CoV-2 infection was associated with immune cell influx, while the proportion of immune and epithelial cells remained nearly stable in children (Fig. 1c and Extended Data Fig. 2a). Upon infection, children’s neutrophils showed an activated phenotype that was more pronounced than in infected adults, characterized by the enhanced expression of, for example, CCL3 and CXCR1/2 (Supplementary Table 4).
a, Nasal samples of children (n = 42) and adults (n = 44) were collected from individuals negative and positive (asymptomatic/mild/moderate COVID-19) for SARS-CoV-2 and subjected to scRNA-seq. b, Uniform manifold approximation and projection (UMAP) showing all identified individual immune and epithelial cell types and states. MC/basophil, mast cells or basophils; moMa, monocyte-derived macrophages; nrMa, non-resident macrophages; rMa, resident macrophages; mDC, myeloid dendritic cells; CD11c_mDC, CD11c-expressing mDC; pDCs, plasmacytoid dendritic cells; DNT, double-negative T cells; NK, natural killer cells; NKT, natural killer T cells; p_NKT, proliferating NKT cells; ag_T, aging T cells; CD8_Tm, memory CD8+ T cells; CTL1 and CTL2, type 1 and 2 cytotoxic T cells; IL-17A_CD8, IL-17A-expressing CD8 T cells; IL-17a_CD4, IL-17A expressing CD4+ T cells; p_T, proliferating T cells; Cil-diff, differentiating ciliated cells; Epi_low RNA, epithelial cells with low RNA; FOXN4, FOXN4+ cells. c, Scaled UMAPs displaying 45,000 cells per group reveal pronounced differences in the nasal cell composition of SARS-CoV-2-negative and -positive children and adults. d, Table showing all cell types/states significantly different between children and adults in non-infected individuals. Given are mean differences in percent; positive values indicate a higher number of cells of the respective cell population in children compared to adults. Comparisons were performed by Kruskal–Wallis test followed by Dunn’s two-sided post hoc comparison with Benjamini–Hochberg correction (***P < 0.001; Neutrophil, 5.54 × 10−7; CD8_Tm, 1.97 × 10−4; CTL2, 1.83 × 10−5; Basal, 6.96 × 10−5; Ciliated, 9.18 × 10−7; also see Extended Data Fig. 2a). e, Scatter plots representing changes of specific immune and epithelial cells over time. Depicted are the percentages of the respective cell type/state with respect to all cells for immune cells (left) or epithelial cells (right) of each individual. Individuals negative for SARS-CoV-2 are indicated in gray, individuals positive for SARS-CoV-2 in red. Lines represent curve fitting results by local polynomial regression (LOESS). The P values are from linear regression analysis (Benjamini–Hochberg adjusted two-tailed).
Interestingly, many of the epithelial cell populations showed a clear age dependency with, for example, goblet cells decreasing and ciliated cells increasing with age (Fig. 1d,e). A recent complementary study analyzed the cell composition of the nasal mucosa in healthy and SARS-CoV-2-infected children based on bulk RNA-seq and cell deconvolution methods. The authors were unable to identify children-specific goblet cells, but rather described that samples from healthy children were dominated by a ciliated cell signature, highlighting the limitations of bulk RNA approaches10.
SARS-CoV-2 entry receptors are not differentially expressed in children compared to adults
The expression level of ACE2, encoding the SARS-CoV-2 entry receptor, and TMPRSS2, FURIN, CTSB, CTSL and CTSV, encoding entry-associated proteases, was similar between children and adults and not upregulated by mild/moderate COVID-19 compared to the uninfected status (Extended Data Fig. 2b). Hence, these viral entry factors cannot explain the differences in SARS-CoV-2 pathophysiology between children and adults.
Children show enhanced viral sensing in airway epithelial cells
SARS-CoV-2 is a positive-strand RNA virus with a very high rate of replication11,12. Hence, the control of SARS-CoV-2 infection requires an optimal early and coordinated innate antiviral immunity. This response is activated by various PRRs. Recently, mounting evidence has been generated in support of MDA5 (IFIH1) as the major PRR for SARS-CoV-2 in epithelial cells with RIG-I (DDX58) possibly playing an additional, but minor, role13,14 (own unpublished data). An important enhancer of viral RNA sensing by MDA5 is LGP2 (DHX58)15. Importantly, PRRs, in particular MDA5 and LGP2, are only weakly expressed in many epithelial cell types but are profoundly upregulated by positive feedback regulation upon viral infection of the cell or by paracrine exposure to type I or III IFN. The dynamics of this feedback regulation are crucial for the successful control of an infecting virus (Fig. 2a). The importance of the PRR/IFN axis for the successful resolution of SARS-CoV-2 infection was recently demonstrated by clinical studies finding a strong association between inborn errors at various loci of the PRR/IFN system with an increased risk of severe COVID-19 (ref. 16). Similarly, and affecting even a much broader fraction of patients, autoantibodies directed against type I IFNs have been shown to occur at a remarkably high frequency in patients with severe COVID-19 (ref. 17).
a, Schematic representation of epithelial response to SARS-CoV-2 infection including genes involved in virus sensing and subsequent IFN response; created with BioRender.com. b, Dot plots depicting the expression of virus-sensing genes in epithelial cells of children and adults. Each significant increase comparing SARS-CoV-2-negative children (n = 18) with SARS-CoV-2-negative adults (n = 23), and SARS-CoV-2-positive children during the early (days post-onset of symptoms (dps) ≤ 4, n = 11) or late infection phase (dps 5–12, n = 11) with SARS-CoV-2-positive adults amid the early (n = 13) or late (n = 8) infection phase, respectively, is marked by a red circle (Benjamini–Hochberg-adjusted two-tailed, Wilcoxon P < 0.05). Ave. exp., average gene expression; Pct. exp., percentage of cells expressing the gene. c, Violin plots showing expression of prototypical virus-sensing genes in epithelial cells of SARS-CoV-2-positive patients (n = 45) in relation to days after first symptoms. d, Heat map showing scaled expression of 171 representative ISGs in all epithelial cells during the early SARS-CoV-2 infection phase (dps ≤ 4). Only selected genes are annotated; for a completely annotated heat map, see Extended Data Fig. 3. e, Differential expression of selected ISGs in an in vitro model comparing the response to SARS-CoV-2 infection in MDA5 (encoded by IFIH1)-overexpressing A549ACE2 cells and empty vector controls. n = 3 biologically independent replicates; error bars show mean ± s.e.m.; *P < 0.05, ***P<0.001 (IFNB1, 5.47 × 10−4; MX1, 2.63 × 10−2; BST2, 8.74 × 10−4; RSAD2, 4.24 × 10−2; IFIT1, 1.30 × 10−2) from one-tailed Student t-test.
Notably, we found a significantly higher basal expression level of the genes coding for RIG-I, MDA5 and LGP2 in epithelial cells in the upper respiratory tract of healthy children as compared to adults (Fig. 2b). This result suggests an increased ability of the respiratory mucosa of children to respond to viral infections, which is further supported by the highly increased amounts of innate immune cells in their upper airways (Fig. 1 and Extended Data Fig. 2a). In epithelial cells of SARS-CoV-2-positive children and adults, we observed a high expression level of those genes (Fig. 2c) in particular at the onset of COVID-19 symptoms that tended to decline until day 4 (day 0–4, referred to as early phase) and remained at lower levels in the later disease phase (days 5 to 12 post symptom onset, referred to as late phase). It can be assumed that higher basal expression of these PRRs would permit immediate sensing of SARS-CoV-2 by MDA5/LGP2 in infected epithelial cells (Fig. 2a). Strikingly, children’s airway epithelial cells displayed increased expression of these PRR genes compared to the expression level of these genes in epithelial cells in SARS-CoV-2-positive adults (Fig. 2b), in particular in the early disease phase after symptom onset. From day 5 onwards, virus sensing is largely comparable between children and adults (Fig. 2b, lower plot).
Following virus sensing, signaling through IRF3/NFκB leads to the expression of primary antiviral effectors, as well as antiviral cytokines such as IFNβ and IFNλ (Fig. 2a). IFNs act on epithelial cells in an autocrine and paracrine manner, further increasing MDA5/LGP2 responsiveness in the tissue and inducing a broad range of IFN-stimulated genes (ISGs). While we were not able to detect the expression of type I and type III IFNs themselves, ISGs showed an impressive activation pattern in epithelial cells of SARS-CoV-2-positive children, including many genes previously shown to exhibit strong antiviral activity against SARS-CoV-2, such as LY6E (ref. 18), IFITM2 and BST2 (ref. 19). In all epithelial cells, and in particular in ciliated cells, the magnitude of ISG expression considerably surpassed that of infected adults in both the early and late infection phases (Fig. 2d and Extended Data Fig. 3) with a generally decreasing trend in the late phase (Extended Data Fig. 3).
To demonstrate a direct association between MDA5 expression levels and activation of ISGs upon SARS-CoV-2 infection, we established an in vitro model using the human lung epithelial cell line A549, which exhibits very low basal expression levels of MDA5 similar to expression levels found in nasal epithelial cells of healthy adults. As expected, based on inefficient MDA5 sensing in concert with rapid replication and expression of virus-encoded antagonists20, only minute amounts of IFNB1 and ISG transcripts were induced upon SARS-CoV-2 infection in these cells. However, in cells with moderately increased basal expression of MDA5 by lentiviral transduction, permitting efficient virus sensing before the expression of antagonists, we observed a significant induction of the expression of IFNB1 and key ISGs including MX1, BST2 (tetherin), RSAD2 (viperin) and IFIT1 (Fig. 2e). These findings corroborate the central role of MDA5 expression before infection for sensing SARS-CoV-2 and inducing a swift and robust ISG response.
Enhanced innate immune cell activation in children
Studying immune–epithelial cell interactions revealed a stronger immune–epithelial cell cross talk in children versus adults particularly before infection. Among immune cells, non-resident macrophages (nrMa) and monocyte-derived macrophages (moMa) as well as CD11c+ dendritic cells (CD11c_mDC) were most interactive (Fig. 3a). These immune cell subsets showed a higher activation status in children as demonstrated by an increased expression level of several cytokine- and chemokine-coding genes such as IL1B, IL8, TNF, CCL3 and CCL4 (Fig. 3b). We furthermore observed an enhanced expression level of IFIH1 in moMa, nrMa and CD11c_mDC in children infected with SARS-CoV-2 and a significant increase of TLR2 in moMa and nrMa in the early phase of infection, suggesting that these cells might play an additional role in virus sensing and IFN production. This is further underlined by the fact that moMa, nrMa and CD11c_mDC of SARS-CoV-2-negative children expressed IFIH1 and TLR2 at higher levels than those of adults.
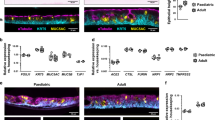
a, Heat maps depicting cell–cell communications between all identified cell types derived from linear-scaled ligand–receptor interaction counts in children (left, n = 18 SARS-CoV-2 negative, n = 11 SARS-CoV-2 positive early infection phase (dps ≤ 4), n = 11 SARS-CoV-2 positive late infection phase (dps 5–12)) and adults (right, n = 23 SARS-CoV-2 negative, n = 13 SARS-CoV-2 positive early infection phase, n = 8 SARS-CoV-2 positive late infection phase). b, Dot plots depicting expression of genes involved in antiviral and cytotoxic response of myeloid dendritic cells and different macrophage populations. Each significant increase comparing SARS-CoV-2-negative children (n = 18) with SARS-CoV-2-negative adults (n = 23) and SARS-CoV-2-positive children during the early (n = 11) or late (n = 11) infection phase with SARS-CoV-2-positive adults amid early (n = 13), or late (n = 8) infection phase, respectively, is marked by a red circle (Benjamini–Hochberg-adjusted two-tailed Wilcoxon, P < 0.05). Ave. exp., average gene expression; Pct. exp., percentage of cells expressing the gene. c, UMAP displaying the different T/NK cell subpopulations in the nose of children and adults. In comparison, scaled density plots are shown indicating the proportion of the different T/NK cells in children and adults separated by their SARS-CoV-2 infection status. Yellow represents high density; blue represents low density. d, Violin plots showing expression of representative immune mediators of the CTL2 subpopulation comparing SARS-CoV-2-negative children (n = 18) with SARS-CoV-2-negative adults (n = 23) and SARS-CoV-2-positive children during the early (n = 11) or late (n = 11) infection phase with SARS-CoV-2-positive adults amid the early (n = 13) or late (n = 8) infection phase, respectively. Each dot represents one cell. Plots show median, ***P < 0.001 (IFNG negative: 1.50 × 10−17, IFNG early phase: 1.14 × 10−47, IFNG late phase: 1.69 × 10−27, CCL5 negative: 5.97 × 10−30, CCL5 late phase: 6.32 × 10−73), derived from two-tailed Wilcoxon comparison.
Distinct immune cell subpopulations in children
Apart from the upregulated cell-intrinsic antiviral capacity of airway epithelial cells, macrophages and dendritic cells, we found specific patterns of immune cell subpopulations in children versus adults. We identified, among others, a subpopulation of KLRC1 (NKG2A)+ cytotoxic T cells (CTL2) occurring predominantly in children (Fig. 3c). NKG2A is a lectin-like inhibitory receptor on cytotoxic T cells playing a role in limiting excessive activation, preventing apoptosis and sustaining the virus-specific CD8+ T cell response21. Already without viral infection, this CD8 cytotoxic T cell subset was characterized by a strong expression level of cytotoxic mediators (Fig. 3d, Extended Data Fig. 4 and Supplementary Table 5). Furthermore, IFNG was highly expressed in these cells when comparing SARS-CoV-2-negative children to adults. Upon infection, children were characterized by a significantly higher expression level of IFNG compared to adults both in the early phase and in the later phase of infection. Similarly, the potent chemoattractant gene CCL5 was increased in children compared to adults with or without infection (Fig. 3d). The cytotoxic potential and the predominance of this cytotoxic T cell subset necessary for efficient killing of virus-infected cells provides further evidence for a better antivirus response in children compared to adults. In addition, SARS-CoV-2-infected children showed a distinct CD8+ T cell population (CD8_Tm) with a memory phenotype that was almost absent in adults (Fig. 3c, EDF4). It remains unclear whether these cells are beneficial for protection of the children against future reinfection.
Discussion
Our data provide clear evidence that the epithelial and immune cells of the upper airways (nose) of children are pre-activated and primed for virus sensing. This is likely a general feature of the children’s mucosal immune response, but of particular relevance for SARS-CoV-2. Very recently, scRNA-seq of fibroblasts infected with Chikungunya virus showed an extremely narrow window of opportunity for the cells to express IFNs before viral protein production shuts off the antiviral system22. This likely also explains the differences between SARS-CoV-2 and other respiratory viruses including respiratory syncytial virus, influenza A virus or SARS-CoV in terms of the induced host response. SARS-CoV-2 is characterized by extensive intracellular replication and a remarkable absence of IFN production and secretion. On the other hand, SARS-CoV-2 is highly sensitive to treatment with IFNs before or after infection, as shown in lung epithelial cells, even more so than SARS-CoV20,23. Primed virus sensing and a pre-activated innate immune response in children leads to efficient early production of IFNs in the infected airways, likely mediating substantial antiviral effects mirroring those observed in vitro in IFN-(pre)treated cells. Ultimately, this may lead to reduced virus replication and faster clearance in children. In fact, several studies already showed that children are much quicker in eliminating SARS-CoV-2 compared to adults, consistent with the concept that they shut down viral replication earlier24,25,26,27. For other respiratory viruses, such as respiratory syncytial virus and influenza A virus, that more efficiently induce an IFN response by themselves, a pre-activated innate immune response may be less relevant. The enhanced innate antiviral capacity in children together with the high IFN sensitivity of SARS-CoV-2 may explain why children are better able to control early-stage infection as compared to adults and therefore have a lower risk of developing severe COVID-19.
Methods
Patient recruitment and ethics approval
Individuals of three different cohorts were included in this study. Patients of the prospective observational cohort study Pa-COVID-19 (ref. 28) and its study arm RECAST (Understanding the increased resilience of children compared to adults in SARS-CoV-2 infection) were enrolled between August 2020 and June 2021 at Charité – Universitätsmedizin Berlin. Further patients were recruited in the prospective SC2-Study8,9 at University Hospital Leipzig between March 2020 and May 2021. Written informed consent was given by all patients and/or their parents before inclusion. All three studies were conducted in accordance with the Declaration of Helsinki and approved by the respective Institutional Review Boards (Pa-COVID-19/RECAST: EA2/066/20, SC2: 123/20-ek).
Patient cohort
From the three cohorts, patients classified as asymptomatic, mild or moderate based on WHO guidelines29 of COVID-19 severity were enrolled. In total, we analyzed nasal swabs of 45 confirmed SARS-CoV-2-positive patients comprising 24 children with ages ranging from 4 weeks to 17 years (median age 9.0 ± 5.6 years, 10 females, 14 males) and from 21 adults between 21 and 76 years (median age 39.0 ± 10.4 years, 12 females, 9 males). None of the children was hospitalized, but all were in domestic quarantine. Additionally, nasal swabs of 42 healthy, SARS-CoV-2-negative controls from 18 children between 4 and 16 years (median age 9.0 ± 3.8 years, 8 females, 10 males) and from 23 adults between 24 and 77 years were included (median age 46.0 ± 16.3 years, 13 females, 10 male; Supplementary Tables 3 and 4). All negative controls were examined for possible exposure to SARS-CoV-2 by detailed anamnesis. In the cases of known SARS-CoV-2 exposures, additional serological testing was conducted at the time of sampling and after 14 days or follow-up interviews were conducted to ensure that the participant as well as their household members showed no signs of infection for at least two consecutive weeks and that any routine testing yielded negative results.
Real-time PCR for SARS-CoV-2
RNA was extracted by using the MagNA Pure 96 DNA and Viral NA Small Volume Kit (Roche) on a MagNA Pure 96 System as recommended by the manufacturer. Real-time PCR with reverse transcription was performed targeting the envelope (E) gene and nucleocapsid (N) gene on the Roche Light Cycler 480 system (Tib-Molbiol).
Obtaining a single-cell suspension from human nasal swabs, preparation for scRNA-seq and subsequent preprocessing of the raw data
Sample processing, single-cell and library preparation, and data analysis were performed as documented previously8,9. Briefly, fresh nasopharyngeal swabs were transferred into cold DMEM/F12 medium (Gibco, 11039) and within 1 h processed further. Under biosafety S2, an equal volume of 13 mM dithiothreitol (AppliChem, A2948) was added to each sample. To achieve higher cell numbers, the solution was slowly pipetted up and down, and the swab was dipped roughly 20 times into the medium. Following incubation at 37 °C and 500 r.p.m. for 10 min on a thermomixer, samples were centrifuged at 350g at 4 °C for 5 min and the supernatant slowly removed. If the pellet showed any sign of red blood cells, it was resuspended in 1× PBS (Sigma-Aldrich, D8537), treated with Red Blood Cell Lysis Buffer (Roche, 11814389001) at 25 °C for 10 min and centrifuged at 350g at 4 °C for 5 min. If samples were not processed immediately, the cell pellet was resuspended in DMEM/F12 supplemented with 20% FBS (Gibco, 10500) and 10% dimethylsulfoxide (Sigma-Aldrich, D8418) and frozen at −80 °C. For the library preparations, cells were thawed at 37 °C, centrifuged at 350g at 4 °C for 5 min and further processed according to the protocol. To obtain a single-cell suspension, Accutase (Thermo Fisher, 00-4555-56) was added to the pellet, and the solution was incubated at room temperature for 10 min with careful pipetting of the cells after 5 min. The incubation was stopped by adding DMEM/F12 supplemented with 10% FBS and centrifugation at 350g at 4 °C for 5 min. Subsequently, the supernatant was removed and the cell pellet was resuspended in 1× PBS (volume was adjusted to the size of the cell pellet). The suspension was cleared of any cell debris using a 35-μm cell strainer (Falcon, 352235), and subsequently, cells were counted with a disposable Neubauer chamber (NanoEnTek, DHC-N01). Cell suspension was diluted to allow loading of 17,500 cells per sample. A single-cell and unique barcode emulsion was achieved by mixing the diluted cells with the master mix and loading them on the chip together with the Gel Beads and Partitioning Oil using the 10x Genomics Single Cell 3ʹ GEM, Library and Gel Bead Kit v3.1 (10x Genomics; PN 1000120; PN 1000121; PN 1000213) and loading the chip into the 10x Chromium Controller. The following reverse transcription, clean-up and cDNA amplification, as well as the library preparation, were performed according to the manufacturer’s manual. Notably, to make sure that the virus was inactivated, we prolonged the incubation at 85 °C during the reverse transcription to 10 min. Final 3ʹ RNA libraries were pooled for sequencing either on an S2 or S4 flow cell (S2: up to 13 samples, S4: up to 24 samples) and sequenced on the NovaSeq 6000 Sequencing System (Illumina, paired-end, single-indexing).
Single-cell datasets were aligned and preprocessed using Cellranger 3.0.1. A custom human hg19 reference genome (10x Genomics, version 3.1.0) with the SARS-CoV-2 genome (Refseq-ID: NC_045512) added as additional chromosome was used. For downstream analysis Seurat 3.2.2 was used. Cells with fewer than 3 genes and cells with more than or equal to 15% mitochondrial reads or fewer than 200 genes expressed were discarded. To remove doublets, a cutoff for the number of unique molecular identifiers and genes was determined manually per sample.
Samples were merged and exported as csv files containing counts and metadata and imported into scanpy 1.6.030. Following normalization to 10,000 reads per cell, expression values were log transformed and highly variable genes were calculated, which were used as a basis for the following preprocessing steps. The data were scaled, PCA transformed and aligned using harmony 0.0.5 (ref. 31) based on 100 principal components. Further alignment was performed using bbknn 1.4.0 (ref. 32) based on 50 prealigned principal components, 10 trees, 3 neighbors within batch and a trim setting of 85. Based on the integrated data, UMAP embedding and Leiden33 clustering were performed. This clustering was used as a basis for subclustering of immune and epithelial cell populations with the same algorithm. Clusters of epithelial34,35,36 and immune37,38,39 cell populations were assigned to cell types/stages according to the expression level of different marker genes (Extended Data Figs. 1 and 4). The T cell and macrophage/dendritic cell clusters were subclustered and then further refined manually.
The object was stored as h5ad, converted to h5seurat using SeuratDisk version 0.0.0.9014 and imported back into R version 4.1.0.
In total, 268,745 cells were included in the dataset. Cell numbers between the groups were equally distributed (negative children 51,595; negative adults 62,701; positive adults 51,500) with the exception of the group of positive children containing higher cell numbers (102,949). Different samples contributed varying numbers of cells. The percentages of contribution of each sample to its study group were compared using a Kruskal–Wallis test, which did not indicate significant differences between groups (P = 0.2).
To enable visual comparisons between UMAPs of different groups, equal numbers of cells (45,000) per group were randomly sampled using the SubsetData function in Seurat.
Putative cell–cell interactions were quantified using CellPhoneDB version 2.1.2 using default settings40. To reduce the influence of individual samples contributing a larger number of cells and to speed up computation, we capped the number of cells per sample at 2,000 randomly sampled cells. This was done using the SubsetData function in Seurat.
Identification of ISG gene set
For the analysis of PRR/IFN responses, a gene set of the most prominent ISGs expressed by lung epithelial cells was assembled. As described previously9, we treated A549 epithelial cells with a mix of IFNβ and IFNλ for 2, 8 or 24 h, and analyzed transcript levels by microarray analyses using the Illumina Human HT-12 v3 Expression Beadchip platform at the genomic and proteomics core facility at DKFZ. We identified ISGs as exhibiting a log2[fold change] > 0.8 at any time point, yielding 183 genes. We further included ISGs described to exhibit strong anti-SARS-CoV-2 activity19 (65 top-scoring genes) if not already included in our list. This eventually yielded a gene set of 217 genes also expressed in our scRNA-seq.
SARS-CoV-2 infection of MDA5-expressing A549 cells
A549 cells stably transduced with a lentiviral vector expressing human IFIH1 under the control of the murine ROSA26 promoter (termed A549 MDA5high in Fig. 2e) were provided by Nadine Gillich and Ralf Bartenschlager. We transduced A549 (termed MDA5low) and A549 MDA5high cells using lentiviral vectors encoding human ACE2 and TMPRSS2 to make them permissive for SARS-CoV-2 infection; to ensure consistent ACE2/TMPRSS2 expression across experiments, transduction was freshly performed 24 h before infection of cells. Infection with SARS-CoV-2 (strain BetaCoV/Germany/BavPat1/2020) was performed in our BSL3 facility at a multiplicity of infection of 0.1, and cells were collected at 24 h post infection. RNA was extracted using the Monarch Total RNA Miniprep Kit (New England Biolabs) and reverse transcribed by the High Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific). IFNB1 and ISG transcript levels were then assessed by real-time PCR using the iTaq Universal SYBR Green Supermix (BioRad) on a BioRad CFX96 Real-Time PCR Detection System. Data are presented as mean ± s.e.m. of three biologically independent replicates.
Statistics
Differential gene expression was calculated using rank_genes_groups() in scanpy version 1.6.0 and corrected for false discovery rates with statsmodels version 0.9.0 (ref. 41). Differences in cell type/stage compositions were assessed using the Kruskal–Wallis rank sum test followed by two-tailed Dunn’s post hoc test. Age dependencies were calculate fitting a linear regression model corrected for the COVID-19 status and adjusting the F-test P values using the Benjamini–Hochberg method42.
P values for dot plots and violin plots were calculated using the Seurat function FindMarkers() based on the Wilcoxon test and corrected with the Benjamini–Hochberg method42.
To test whether elevated MDA5 and RIG-I levels in A549 cells significantly increased by IFNB1 and ISG induction upon infection, an unpaired one-tailed Student t-test of three biologically independent repetitions was performed (GraphPad Prism v9.1).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Processed data in the form of a Seurat object are available without access restrictions on FigShare (https://doi.org/10.6084/m9.figshare.14938755). This dataset can be used to replicate and extend the analyses presented in this paper. Due to potential risk of de-identification of pseudonymized RNA-seq data, the raw sequencing data can be obtained through EGA (EGAS00001005461) for non-commercial research purposes alone, subject to controlled access as mandated by EU data protection laws. For access, contact the corresponding author, R.E. In addition, these data can be further visualized and analyzed in the Magellan COVID-19 data explorer at https://digital.bihealth.org.
References
Castagnoli, R. et al. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in children and adolescents: a systematic review. JAMA Pediatr. 174, 882–889 (2020).
Goldstein, E., Lipsitch, M. & Cevik, M. On the effect of age on the transmission of SARS-CoV-2 in households, schools, and the community. J. Infect. Dis. 223, 362–369 (2021).
O’Driscoll, M. et al. Age-specific mortality and immunity patterns of SARS-CoV-2. Nature 590, 140–145 (2021).
Park, A. & Iwasaki, A. Type I and type III interferons - induction, signaling, evasion, and application to combat COVID-19. Cell Host Microbe 27, 870–878 (2020).
Pierce, C. A. et al. Natural mucosal barriers and COVID-19 in children. JCI Insight 6, e148694 (2021).
Yoshida, M. et al. The local and systemic response to SARS-CoV-2 infection in children and adults. Preprint at medRxiv https://doi.org/10.1101/2021.03.09.21253012 (2021).
Hou, Y. J. et al. SARS-CoV-2 reverse genetics reveals a variable infection gradient in the respiratory tract. Cell 182, 429–446 (2020).
Chua, R. L. et al. COVID-19 severity correlates with airway epithelium–immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 38, 970–979 (2020).
Trump, S. et al. Hypertension delays viral clearance and exacerbates airway hyperinflammation in patients with COVID-19. Nat. Biotechnol. (2020).
Koch, C. M. et al. Immune response to SARS-CoV-2 in the nasal mucosa in children and adults. Preprint at medRxiv https://doi.org/10.1101/2021.01.26.21250269 (2021).
V’Kovski, P., Kratzel, A., Steiner, S., Stalder, H. & Thiel, V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. 19, 155–170 (2021).
Wolfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature 581, 465–469 (2020).
Rebendenne, A. et al. SARS-CoV-2 triggers an MDA-5-dependent interferon response which is unable to control replication in lung epithelial cells. J. Virol. 95, e02415–e02420 (2021).
Yin, X. et al. MDA5 governs the innate immune response to SARS-CoV-2 in lung epithelial cells. Cell Rep. 34, 108628 (2021).
Bruns, A. M., Leser, G. P., Lamb, R. A. & Horvath, C. M. The innate immune sensor LGP2 activates antiviral signaling by regulating MDA5-RNA interaction and filament assembly. Mol. Cell 55, 771–781 (2014).
Zhang, Q. et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 370, eabd4570 (2020).
Bastard, P. et al. Auto-antibodies to type I IFNs can underlie adverse reactions to yellow fever live attenuated vaccine. J. Exp. Med. 218, e20202486 (2021).
Pfaender, S. et al. LY6E impairs coronavirus fusion and confers immune control of viral disease. Nat. Microbiol. 5, 1330–1339 (2020).
Martin-Sancho, L. et al. Functional landscape of SARS-CoV-2 cellular restriction. Mol. Cell 81, 2656–2668.e8 (2021).
Hayn, M. et al. Systematic functional analysis of SARS-CoV-2 proteins uncovers viral innate immune antagonists and remaining vulnerabilities. Cell Rep. 35, 109126 (2021).
Rapaport, A. S. et al. The inhibitory receptor NKG2A sustains virus-specific CD8+ T cells in response to a lethal poxvirus infection. Immunity 43, 1112–1124 (2015).
Pott, F. et al. Single-cell transcriptomic analysis of antiviral responses and viral antagonism in Chikungunya virus-infected synovial fibroblasts. Preprint at bioRxiv https://doi.org/10.1101/2020.06.07.138610 (2021).
Lokugamage, K. G. et al. Type I interferon susceptibility distinguishes SARS-CoV-2 from SARS-CoV. J. Virol. 94, e01410-20 (2020).
Dufort, E. M. et al. Multisystem inflammatory syndrome in children in New York state. N. Engl. J. Med. 383, 347–358 (2020).
Feldstein, L. R. et al. Multisystem inflammatory syndrome in U.S. children and adolescents. N. Engl. J. Med. 383, 334–346 (2020).
Weisberg, S. P. et al. Distinct antibody responses to SARS-CoV-2 in children and adults across the COVID-19 clinical spectrum. Nat. Immunol. 22, 25–31 (2021).
Whittaker, E. et al. Clinical characteristics of 58 children with a pediatric inflammatory multisystem syndrome temporally associated with SARS-CoV-2. JAMA 324, 259–269 (2020).
Kurth, F. et al. Studying the pathophysiology of coronavirus disease 2019: a protocol for the Berlin prospective COVID-19 patient cohort (Pa-COVID-19). Infection 48, 619–626 (2020).
Aylward, B., Liang, W. & WHO-China-Joint-Mission. Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19) (WHO, 2020).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Polański, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Lukassen, S. et al. SARS-CoV-2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells. EMBO J. 39, e105114 (2020).
Plasschaert, L. W. et al. A single-cell atlas of the airway epithelium reveals the CFTR-rich pulmonary ionocyte. Nature 560, 377–381 (2018).
Vieira Braga, F. A. et al. A cellular census of human lungs identifies novel cell states in health and in asthma. Nat. Med. 25, 1153–1163 (2019).
Mogilenko, D. A. et al. Comprehensive profiling of an aging immune system reveals clonal GZMK+ CD8+ T cells as conserved hallmark of inflammaging. Immunity 54, 99–115 (2021).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Zhao, S. T. & Wang, C. Z. Regulatory T cells and asthma. J. Zhejiang Univ. Sci. B 19, 663–673 (2018).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Seabold, S. & Perktold, J. Statsmodels: econometric and statistical modeling with Python. Proc. 9th Python in Science Conference 57–61 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Acknowledgements
We thank all patients of the RECAST, Pa-COVID-19 and SC2 studies for donating nasal samples and clinical data. This study was supported by the BIH COVID-19 research program and the fightCOVID@DKFZ initiative, the European commission (ESPACE, 874719, Horizon 2020), the German Federal Ministry for Education and Research (BMBF)-funded de.NBI Cloud within the German Network for Bioinformatics Infrastructure (de.NBI; 031A537B, 031A533A, 031A538A, 031A533B, 031A535A, 031A537C, 031A534A and 031A532B), the BMBF-funded Medical Informatics Initiative (HiGHmed, 01ZZ1802A - 01ZZ1802Z), the BMBF-funded projects 01IK20337 and 82DZL0098B1, the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG)-funded CRC-TR 84 B08, CRC-1449 Z01 and project ID 272983813-TRR179 and the DFG COVID-19 focus funding project BI1693/2-1. This publication is part of the Human Cell Atlas—www.humancellatlas.org/publications.
Author information
Authors and Affiliations
Contributions
S.T., S. Lukassen, I.L., M.A.M. and R.E. conceived, designed and supervised the project. B.S., M.R., J.R., L.S., V.M.C. and M.A.M. designed the RECAST cohort. J.R., S.S., L.S. and S. Laudi recruited the patients and provided the human specimens, clinical data and annotation of the patients. J. Loske, A.S., J. Liebig, M.M. and R.L.C. performed scRNA experiments. V.G.M. and M.B. performed and analyzed the in vitro experiments. J. Loske, L.T., S. Lukassen and R.L.C. analyzed data. B.S., L.S., S. Laudi, V.M.C., M.R., C.C., M.B. and M.A.M. contributed with discussion of the results. B.T. and S.K. provided technical and experimental support. J. Loske, S. Lukassen, S.T., M.B., R.E. and I.L. wrote the manuscript; all authors read, revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Biotechnology thanks Melanie Neeland and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cell type marker genes.
(a-b) Dot plots depicting average and percent expression of genes used to classify (a) immune and (b) epithelial cells. Pct. Exp = percentage of cells expressing the gene, Ave. Exp. = average gene expression.
Extended Data Fig. 2 Cell composition in children and adults and ACE2 expression.
(a) Distribution of all cell types/ states in children and adults separated by SARS-CoV-2 infection status. Given are percentages of the total number of cells. Comparisons by Kruskal-Wallis test followed by Dunn’s two-sided post hoc comparison (*significance of comparison between children and adults, P < 0.05; Neutrophil: 5.54E-07, mDC: 1.47E-03, rMa: 3.71E-03, NK: 8.73E-03, p_NKT: 4.96E-02, ag_T: 6.09E-04, Treg: 8.50E-03, DNT: 1.62E-04, CD8_Tm: 1.97E-04, IL17A_CD8: 3.05E-04, IL17A_CD4: 2.68E-03, CTL2: 1.83E-05, CTL1: 3.60E-03, Plasma_cell: 1.83E-05, B_cell: 2.45E-03, immune_total: 1.61E-06, Basal: 6.96E-05, Cil_diff: 2.35E-03, Cil_acute_phase: 8.92E-06, Ciliated: 9.18E-07, Sec_diff: 5.09E-03). (b) Violin plots show distribution of the SARS-CoV-2 entry receptor ACE2 expression and the gene expression of associated proteases in epithelial cells of children and adults either with (n=24 children, n=21 adults) or without SARS-CoV-2 infection (n=18 children, n=23 adults). Plots show median. No significant differences were observed.
Extended Data Fig. 3 Epithelial IFN-response.
Heat maps depicting scaled expression (zero mean, unit variance) of known IFN-response genes in all epithelial cells and separately in ciliated cells across conditions during early and late SARS-CoV-2 infection phase (early: dps ≤ 4: n=11 children, n=13 adults; late dps >4: n=11 children, n=8 adults). Genes were filtered according to a minimum fold change of 1.5 between the lowest and highest expressing group, with the gene being selected according to their differential expression in all epithelial cells. Genes not meeting the criterion were greyed out to avoid inflating minor differences as a consequence of the scaling performed. Scaled Exp. = scaled expression.
Extended Data Fig. 4 Characteristics of NK and T-cells.
Expression profile of cytotoxic and aging-related genes in the NK and T-cell subtypes derived from n=86 nasal swap samples distinguishing the children’s and adults’ immune profile. Ave. Exp. = average gene expression, Pct. Exp. = percentage of cells expressing the gene.
Supplementary information
Supplementary Table
Supplementary Table 1. Characteristics of the individuals testing positive (n = 45; a) and negative (n = 42; b) included in the present study.
Supplementary Table
Supplementary Table 2. Summary of cohort characteristics.
Supplementary Tables
Supplementary Tables 3–5. (TS 3): Characteristics of sample preparation of all individuals (n = 86) included in the present study. (TS 4): Differentially expressed genes in Neutrophil. (TS 5): Differentially expressed genes in the different T/NK-cell clusters.
Rights and permissions
About this article
Cite this article
Loske, J., Röhmel, J., Lukassen, S. et al. Pre-activated antiviral innate immunity in the upper airways controls early SARS-CoV-2 infection in children. Nat Biotechnol 40, 319–324 (2022). https://doi.org/10.1038/s41587-021-01037-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01037-9
This article is cited by
-
The mechanisms of milder clinical symptoms of COVID-19 in children compared to adults
Italian Journal of Pediatrics (2024)
-
Evolution of enhanced innate immune suppression by SARS-CoV-2 Omicron subvariants
Nature Microbiology (2024)
-
Natural and hybrid immunity after SARS-CoV-2 infection in children and adolescents
Infection (2024)
-
Severe pediatric COVID-19: a review from the clinical and immunopathophysiological perspectives
World Journal of Pediatrics (2024)
-
SARS-CoV-2-Infektion und Autoimmunität
Zeitschrift für Rheumatologie (2024)