Abstract
Tumor-associated epitopes presented on MHC-I that can activate the immune system against cancer cells are typically identified from annotated protein-coding regions of the genome, but whether peptides originating from novel or unannotated open reading frames (nuORFs) can contribute to antitumor immune responses remains unclear. Here we show that peptides originating from nuORFs detected by ribosome profiling of malignant and healthy samples can be displayed on MHC-I of cancer cells, acting as additional sources of cancer antigens. We constructed a high-confidence database of translated nuORFs across tissues (nuORFdb) and used it to detect 3,555 translated nuORFs from MHC-I immunopeptidome mass spectrometry analysis, including peptides that result from somatic mutations in nuORFs of cancer samples as well as tumor-specific nuORFs translated in melanoma, chronic lymphocytic leukemia and glioblastoma. NuORFs are an unexplored pool of MHC-I-presented, tumor-specific peptides with potential as immunotherapy targets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data: the raw Ribo-seq data (fastq.gz), offset-corrected BAM files used for translated ORF identification by RibORF and BigWig file generation, BigWig files for Ribo-seq data visualization in genome browsers and Ribo-seq translation levels (TPM) are deposited to NCBI GEO (GSE143263) for established cell lines (B721.221, A375 and HCT116) and for primary melanocytes (Thermo C0025C). GTEx, TCGA, CLL and healthy B cell samples RNA-seq transcription quantification of transcript isoforms is deposited to NCBI GEO GSE143263. Ribo-seq translation levels (TPM) of primary GBM and melanoma samples are deposited to NCBI GEO GSE143263. Raw data pertaining to primary patient samples is deposited to dbGaP: CLL1-5 Ribo-seq and CLL4, CLL5 RNA-seq data are available through dbGaP phs001998; Ribo-seq data for MEL2, MEL11 and GBM7 and matching RNA-seq data for MEL11 are available through dbGaP phs001451. B721.221 RNA-seq data for HLA-C (C*04:01, C*07:01) is deposited under GEO GSE131267. Melanoma RNA-seq data are deposited in dbGaP (https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs001451.v1.p1, ref. 4). Glioblastoma bulk RNA-seq data are available through dbGaP (https://www.ncbi.nlm.nih.gov/gap) with accession number phs001519.v1.p1 (ref. 3). Mass spectrometry data: the original mass spectra for immunopeptidomes of two melanoma patient-derived cell lines and the full proteome of a glioblastoma patient-derived cell line, tables of PSMs for all experiments, and the protein sequence databases used for searches have been deposited in the public proteomics repository MassIVE (https://massive.ucsd.edu) and are accessible at ftp://massive.ucsd.edu/MSV000084787. Original mass spectrometry data for the previously published monoallelic immunopeptidomes, B721.221 cell line full proteome and patient-derived cell line immunopeptidomes are accessible at ftp://massive.ucsd.edu/MSV000080527, ftp://massive.ucsd.edu/MSV000084172, and ftp://massive.ucsd.edu/MSV000084442. Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Code availability
Python scripts and Jupyter notebooks used in the analysis are available on GitHub at https://github.com/klarman-cell-observatory/Riboseq-nuORFs.
References
Hu, Z., Ott, P. A. & Wu, C. J. Towards personalized, tumour-specific, therapeutic vaccines for cancer. Nat. Rev. Immunol. 18, 168–182 (2018).
Hilf, N. et al. Actively personalized vaccination trial for newly diagnosed glioblastoma. Nature 565, 240–245 (2019).
Keskin, D. B. et al. Neoantigen vaccine generates intratumoral T cell responses in phase Ib glioblastoma trial. Nature 565, 234–239 (2019).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Robbins, P. F. et al. The intronic region of an incompletely spliced gp100 gene transcript encodes an epitope recognized by melanoma-reactive tumor-infiltrating lymphocytes. J. Immunol. 159, 303–308 (1997).
Van Den Eynde, B. J. et al. A new antigen recognized by cytolytic T lymphocytes on a human kidney tumor results from reverse strand transcription. J. Exp. Med. 190, 1793–1800 (1999).
Wang, R. F. et al. A breast and melanoma-shared tumor antigen: T cell responses to antigenic peptides translated from different open reading frames. J. Immunol. 161, 3596–3606 (1998).
Abelin, J. G. et al. Mass spectrometry profiling of HLA-associated peptidomes in mono-allelic cells enables more accurate epitope prediction. Immunity 46, 315–326 (2017).
Sarkizova, S. et al. A large peptidome dataset improves HLA class I epitope prediction across most of the human population. Nat. Biotechnol. 38, 199–209 (2019).
Laumont, C. M. et al. Global proteogenomic analysis of human MHC class I-associated peptides derived from non-canonical reading frames. Nat. Commun. 7, 10238 (2016).
Laumont, C. M. et al. Noncoding regions are the main source of targetable tumor-specific antigens. Sci. Transl. Med. 10, eaau5516 (2018).
Chen, J. et al. Pervasive functional translation of noncanonical human open reading frames. Science 367, 1140–1146 (2020).
Chong, C. et al. Integrated proteogenomic deep sequencing and analytics accurately identify non-canonical peptides in tumor immunopeptidomes. Nat. Commun. 11, 1293 (2020).
Nesvizhskii, A. I. Proteogenomics: concepts, applications and computational strategies. Nat. Methods 11, 1114–1125 (2014).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Fields, A. P. et al. A regression-based analysis of ribosome-profiling data reveals a conserved complexity to mammalian translation. Mol. Cell 60, 816–827 (2015).
Ji, Z., Song, R., Regev, A. & Struhl, K. Many lncRNAs, 5′ UTRs, and pseudogenes are translated and some are likely to express functional proteins. eLife 4, e08890 (2015).
Chew, G.-L. et al. Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140, 2828–2834 (2013).
Erhard, F. et al. Improved Ribo-seq enables identification of cryptic translation events. Nat. Methods 15, 363–366 (2018).
Martinez, T. F. et al. Accurate annotation of human protein-coding small open reading frames. Nat. Chem. Biol. 16, 458–468 (2019).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Iyer, M. K. et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 47, 199–208 (2015).
Mylonas, R. et al. Estimating the contribution of proteasomal spliced peptides to the HLA-I ligandome. Mol. Cell Proteom. 17, 2347–2357 (2018).
Rolfs, Z., Müller, M., Shortreed, M. R., Smith, L. M. & Bassani-Sternberg, M. Comment on ‘A subset of HLA-I peptides are not genomically templated: evidence for cis- and trans-spliced peptide ligands’. Sci. Immunol. 4, eaaw8457 (2019).
Yoshimura, A., Naka, T. & Kubo, M. SOCS proteins, cytokine signalling and immune regulation. Nat. Rev. Immunol. 7, 454–465 (2007).
Faridi, P. et al. A subset of HLA-I peptides are not genomically templated: Evidence for cis- and trans-spliced peptide ligands. Sci. Immunol. 3, eaar3947 (2018).
Liepe, J. et al. A large fraction of HLA class I ligands are proteasome-generated spliced peptides. Science 354, 354–358 (2016).
Raj, A. et al. Thousands of novel translated open reading frames in humans inferred by ribosome footprint profiling. eLife 5, e13328 (2016).
Hutter, C. & Zenklusen, J. C. The Cancer Genome Atlas: creating lasting value beyond its data. Cell 173, 283–285 (2018).
Blum, A., Wang, P. & Zenklusen, J. C. SnapShot: TCGA-analyzed tumors. Cell 173, 530 (2018).
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Consortium, G. TEx. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Georgiadis, P. et al. Evolving DNA methylation and gene expression markers of B-cell chronic lymphocytic leukemia are present in pre-diagnostic blood samples more than 10 years prior to diagnosis. BMC Genomics 18, 728 (2017).
Rodríguez, A. E. et al. Molecular characterization of chronic lymphocytic leukemia patients with a high number of losses in 13q14. PLoS ONE 7, e48485 (2012).
Rajasagi, M. et al. Systematic identification of personal tumor-specific neoantigens in chronic lymphocytic leukemia. Blood 124, 453–462 (2014).
Gonzalez, C. et al. Ribosome profiling reveals a cell-type-specific translational landscape in brain tumors. J. Neurosci. 34, 10924–10936 (2014).
Shraibman, B. et al. Identification of tumor antigens among the HLA peptidomes of glioblastoma tumors and plasma. Mol. Cell Proteom. 18, 1255–1268 (2019).
Blair, J. D., Hockemeyer, D., Doudna, J. A., Bateup, H. S. & Floor, S. N. Widespread translational remodeling during human neuronal differentiation. Cell Rep. 21, 2005–2016 (2017).
Freitas, M. et al. Expression of cancer/testis antigens is correlated with improved survival in glioblastoma. Oncotarget 4, 636–646 (2013).
Su, R. et al. Knockdown of SOX2OT inhibits the malignant biological behaviors of glioblastoma stem cells via up-regulating the expression of miR-194-5p and miR-122. Mol. Cancer 16, 171 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Malone, B. et al. Bayesian prediction of RNA translation from ribosome profiling. Nucleic Acids Res. 45, 2960–2972 (2017).
Bassani-Sternberg, M. et al. Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry. Nat. Commun. 7, 13404 (2016).
Harndahl, M. et al. Peptide binding to HLA class I molecules: homogenous, high-throughput screening, and affinity assays. J. Biomol. Screen. 14, 173–180 (2009).
Jurtz, V. et al. NetMHCpan-4.0: improved peptide-MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. J. Immunol. 199, 3360–3368 (2017).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature 526, 525–530 (2015).
Ferreira, P. G. et al. Transcriptome characterization by RNA sequencing identifies a major molecular and clinical subdivision in chronic lymphocytic leukemia. Genome Res. 24, 212–226 (2014).
Puente, X. S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526, 519–524 (2015).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinf. 12, 323 (2011).
Acknowledgements
We thank K. Gosik and R. Herbst for their help with the statistical analysis. We thank D. Fu for her help with the nonmetric multidimensional scaling analysis. We thank E. Hodis and J. Kwon for providing cultured primary melanocytes. We thank K.L. Ligon for providing the GBM cell line. We thank L. Gaffney for help with figure preparation. Work was supported by the Klarman Cell Observatory and HHMI (A.R.), NIH grant nos. NCI-1R01CA155010-02 (to C.J.W.), NHLBI-5R01HL103532-03 (to C.J.W.), NIH/NCI R21 CA216772-01A1 (to D.B.K.), NCI-SPORE-2P50CA101942-11A1 (to D.B.K), NHGRI T32HG002295 and NIH/NCI T32CA207021 (to S.S.), NCI R50CA211482 (to S.A.S.), NHGRI U41HG007234 and R01 HG004037 (to I.J.), NCI Clinical Proteomic Tumor Analysis Consortium grant nos. NIH/NCI U24-CA210986 and NIH/NCI U01 CA214125 (to S.A.C.) and NIH/NCI U24CA210979 (to D.R. Mani and G. Getz). This work was supported in part by The G. Harold and Leila Y. Mathers Foundation and the Bridge Project, a partnership between the Koch Institute for Integrative Cancer Research at MIT and the Dana-Farber/Harvard Cancer Center. C.J.W. is a scholar of the Leukemia and Lymphoma Society, and is supported in part by the Parker Institute for Cancer Immunotherapy. S.K. is a Cancer Research Institute/Hearst Foundation fellow. T.O. is a Leukemia and Lymphoma Society Fellow. B.A.K. is supported by a long-term EMBO fellowship (ALTF 14-2018). P.B. is supported by an Amy Strelzer Manasevit Grant and an American Society of Hematology Scholar Award. G.O. is supported by a postdoctoral fellowship sponsored by the American-Italian Cancer Foundation.
Author information
Authors and Affiliations
Contributions
T.O. and A.R. conceived the study. D.B.K., S.A.C., C.J.W., N.H. and A.R. directed the overall study design. T.O., E.C. and Y.T.C. generated Ribo-seq libraries. T.O., T.L. and S.C. performed Ribo-seq analysis. S.K., K.R.C., T.O., T.L., S.S., C.R.H., H.K. and A.A. generated the MS data and performed the associated data analysis. B.A.K. provided CLL RNA-seq data. F.A. performed GTEx, TCGA and CLL RNA-seq alignment and quantification under G.G.’s guidance. B.L. performed WGS analysis. D.B.K. and P.M.L. generated the single-HLA allele cell lines. D.B.K., G.O. and C.J.W. provided the patient-derived tumor cell lines. P.B. provided CLL samples. P.B., W.Z. and D.B.K. prepared peripheral blood mononuclear cells and B cells from patients with CLL and healthy donors. I.J. performed conservation analysis under M.K.’s guidance. S.J. performed MHC-I binding assays. Z.J. and S.A.S. provided computational support. T.O., T.L., K.R.C., S.K., S.S., D.B.K., S.A.C., C.J.W. and A.R. wrote the paper, with contributions from all coauthors.
Corresponding authors
Ethics declarations
Competing interests
A.R. is a founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas Therapeutics and until 31 August, 2020 was an SAB member of Syros Pharmaceuticals, Neogene Therapeutics, Asimov and ThermoFisher Scientific. From 1 August, 2020, A.R. is an employee of Genentech. C.J.W. and N.H. were cofounders, equity holders and SAB members of Neon Therapeutics, Inc. until May 2020, and now are equity holders of BionTech, Inc. D.B.K. has previously advised Neon Therapeutics, and has received consulting fees from Guidepoint, Neon Therapeutics, System Analytic Ltd and The Science Advisory Board. T.O. owns equity in BionTech, Moderna, Gilead, Novartis, Roche, 10X Genomics and Illumina. Since 3 August, 2020, T.O. is an employee of Flagship Labs 69. D.B.K. owns equity in Aduro Biotech, Agenus Inc., Armata Pharmaceuticals, Breakbio Corp., Biomarin Pharmaceutical Inc., Bristol-Myers Squibb Com., Celldex Therapeutics Inc., Editas Medicine Inc., Exelixis Inc., Gilead Sciences Inc., IMV Inc., Lexicon Pharmaceuticals Inc., and Stemline Therapeutics Inc. P.B. owns equity in Amgen Inc., Breakbio Corp., and Stemline Therapeutics Inc. S.A.S. has previously advised Neon Therapeutics and has received consulting fees from Neon Therapeutics. S.A.S. owns equity in Agenus Inc., Agios Pharmaceuticals, 152 Therapeutics, Breakbio Corp., Bristol-Myers Squibb and NewLink Genetics. S.A.C. is a SAB member of Kymera, PTM BioLabs and Seer and a scientific advisor to Pfizer and Biogen. T.O., T.L., K.R.C., S.K., N.H., D.B.K., S.A.C., C.J.W. and A.R. are coinventors on PCT/US2019/066104 directed to neoantigens and methods for identifying neoantigens as described in this paper.
Additional information
Peer review information Nature Biotechnology thanks Robert Bradley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
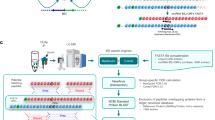
Extended Data Fig. 1 nuORFdb characteristics.
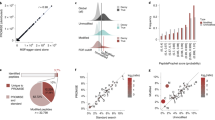
a. Hierarchical ORF prediction. Tree showing individual samples (leaves), combinations of samples (clades) and entire datasets of all reads (root) representing the nodes used to make ORF predictions (arrowheads). #: samples used in nuORFdb construction, but later discovered to be of poor quality and not used in any subsequent analyses; CHX: samples pre-treated with cycloheximide; Harr: samples pretreated with harringtonine, IFNy: samples pre-treated with interferon gamma. b. NuORFdb size relative to the annotated proteome, RNA-seq- and transcriptome-based databases. Number of ORFs (y axis) across four databases (x axis). c-d. Ribo-seq reveals mRNA reading frames. c. RNA-seq (blue) and Ribo-seq (green) reads aligned to the transcript of the MLEC gene. RNA-seq reads align to the entire length of the transcript, while Ribo-seq reads align exclusively to the translated portions. Ribo-seq supports translation of a 5’ uORF (red box, top). Histogram of +15nt-shifted 5’ ends of Ribo-seq reads supporting translation of the MLEC 5’ uORF (colorful) with corresponding full-length aligned reads below. 5‘ ends of full-length reads are outlined in colors matching their +15nt-shifted positions in the histogram (bottom). d. Histogram of 5’ ends of Ribo-seq reads supporting translation of annotated protein-coding ORFs at every third nucleotide (x axis) around the start codon (left) and the stop codon (right). The –12 position of the first peak indicates the placement of the ribosome at the start codon (position 0), which is computationally adjusted to +3 by adding +15nt to each 5’ end read location, as shown in (c).
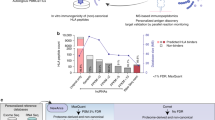
Extended Data Fig. 2 nuORFdb benchmarking.
a. Spectra search times (y axis) for the HLA-A*02:01 sample with different databases (x axis). b-c. nuORFdb minimizes the loss of sensitivity for annotated peptides, while enabling discovery of nuORF peptides. Number of annotated peptides (b) and nuORF peptides (c) discovered (y axis) across four databases (x axis). d. nuORFdb spectra mapping has the lowest % FDR among the three databases. %FDR for nuORF peptides (y axis) across databases (x axis). Global FDR for all peptides was set to 1%. e. nuORF peptides are discovered across multiple databases. Number of nuORF peptides unique to or shared across databases (y axis), as indicated by the black circles below (x axis). Bars on the bottom left indicate the total number of nuORF peptides discovered using each database. f. Ratios of nuORF types discovered vary depending on the database used for spectra mapping. Proportion of nuORFs of different types (y axis) in the set of nuORFs discovered by all three databases (Shared), using each database, or those specific to each database and not found by others (x axis). g. ORFs discovered using different databases vary in RNA-seq and Ribo-seq read coverage. Percent of annotated (UCSCdb) or nuORF (other databases) peptides with >0 reads (y axis) discovered using the four databases, or discovered uniquely by a database (x axis). h-k. MS spectrum mapping to the correct peptide sequence is more challenging using RNAdb and TransDb. h. Distribution of the number of considered matches for each spectrum across four databases. i. Difference between Spectrum Mill score for the top ranked (Rank1) and second best (Rank2) peptide sequences (y axis) across databases (x axis). n = 11007 (UCSC), 155 (Shared), 253 (nuORFdb), 68 (nuORFdb specific), 320 (RNAdb), 64 (RNAdb specific), 389 (TransDb), 149 (TransDb specific). Median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown. j. Distribution of the HLAthena-predicted binding score (MSi) (left) and percent of peptides with MSi score >= 0.8 (red line on the left) (x axis) across databases (y axis). k. Predicted hydrophobicity index (y axis) and retention time (x axis) of peptides discovered using different databases for the HLA-A*24:02 sample.
Extended Data Fig. 3 Additional filtering of MHC I IP, MS/MS-detected nuORF peptides.
a-d. Impact of filtering on nuORF number, types and false discovery rates. a,b. Total number of nuORF peptides (y axis) identified pre-filtering (solid bars) and retained post-filtering (hashed bars) overall (a) and for different nuORF types (x axis, b). c,d. False discovery rate (y axis) for annotated (gray) and nuORF (pink) peptides across 92 HLA alleles pre- and post-filtering (hashed) overall (c) and for different ORF types (x axis, d). e. Criteria used to filter peptides across ORF types. f. Filtering thresholds across nuORF categories. Filter cutoffs (vertical red lines) across different peptide spectral match scoring features (x axis) for different ORF types (y axis). n = 191897 (annotated), 2050 (5’ uORF), 1619 (Out-of-frame), 1542 (5’ overlap uORF), 855 (lincRNA), 514 (ncRNA Processed Transcript), 497 (3’ dORF), 376 (ncRNA Retained Intron), 341 (Pseudogene), 311 (3’ overlap dORF), 299 (Antisense), 163 (Other). Median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown. g. Filtering impact across categories. Percent of peptides (y axis) retained post-filtering across different ORF categories and overall (x axis).
Extended Data Fig. 4 nuORFs peptides in the MHC I immunopeptidome have comparable biochemical properties to annotated peptides.
a. MHC I immunopeptidome includes peptides from different nuORF categories. Number of unique proteins (x axis) detected by MHC I IP LC-MS/MS across expanded ORF types (y axis). b-g. Comparable biochemical features of nuORF and annotated peptides. b. Distribution of LC-MS/MS Spectrum Mill identification score (x axis) for annotated and nuORF peptides across ORF types (y axis). c. Peptide fragmentation score (x axis) for peptides identified across ORF types (y axis). d. Ribo-seq translation levels (x axis, log2(TPM+1)) of MHC I MS-detected ORFs across various ORF types (y axis). For all boxplots, n = 17426 (annotated), 806 (5’ uORF), 776 (lncRNA), 692 (5’ overlap uORF), 595 (Out-of-frame), 169 (3’ dORF), 120 (Pseudogene), 54 (3’ Overlap dORF), 48 (Other); median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown. e. Predicted hydrophobicity index (y axis) against the LC-MS/MS retention time (x axis) for annotated (grey) and nuORF (pink) peptide sequences for three representative HLA alleles. Dashed line: Lowess fit to the annotated peptides. Sample sizes, root mean square errors (rmse), and p-values (rank-sum test on residuals) are marked. f,g. Similar sequence motifs in nuORFs and annotated peptides. f. Non-metric multidimensional scaling (NMDS) plot of all MHC IP LC-MS/MS-detected annotated and nuORF 9 AA peptide sequences clustered by peptide sequence similarity for three representative HLA alleles. g. Consensus peptide sequence motif plots of all MHC IP LC-MS/MS-detected annotated and nuORF 9 AA peptide sequences.
Extended Data Fig. 5 Hierarchical ORF prediction based on Ribo-seq identifies short, overlapping, tissue-specific nuORFs.
a. nuORFs predictions are more sample and tissue specific than annotated ORFs. Proportion of annotated ORFs (grey) and nuORFs (pink) in the MHC I immunopeptidome (y axis, and pie chart). Hashed: proportion predicted only at the leaf and clade level, but not at the root. b. Two overlapping, MHC I MS-detected 5’ uORFs in LUZP1 as an example of tissue-specific, overlapping nuORFs identified by hierarchical ORF prediction. uORF2 (pink) was predicted in the CLL clade, and not at the root. uORF1 (cyan) was predicted at the root and not in the CLL clade. Detected peptides outlined in red with the HLA alleles where peptides were detected marked below. c. SOCS1 gene as an example of identification of short, overlapping nuORFs. SOCS1 gene encodes three translated proteins: the annotated ORF, an out-of-frame iORF, and a 5’ overlap ouORF. Two MHC I MS-detected peptides from 5’ ouORF outlined in yellow. Detected iORF peptide outlined in red and shown in higher magnification below. Bottom: Histogram of Ribo-seq reads supporting translation of the annotated ORF (blue) and the out-of-frame iORF (green).
Extended Data Fig. 6 nuORF peptides in the MHC I immunopeptidome and whole proteome of cancer cells.
a. nuORFdb helps map immunopeptidome even from samples and tumor types not used in constructing the reference. Total number of MHC I LC-MS/MS spectra mapped (y axis) across cancer samples (x axis). b-d. nuORFs of various types were detected in the MHC I immunopeptidome of cancer samples. Number (b) and proportion (c) of nuORFs (y axis) of different types identified in each cancer sample (x axis). d. Distribution of the fraction (y axis) of nuORF types (x axis) in B721.221 cells (dark grey) or across cancer samples (light grey). Asterisk: p < 0.05 (lncRNA p = 5 × 10−6, 5′ uORF p = 0.03; two-sided rank-sum test. n = 10 cancer samples, n = 100000 random samplings across alleles. Median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown. e-h. nuORFs are more abundant in the MHC I immunopeptidome than in the whole proteome. e. Percent of nuORF peptides (y axis) detected in the immunopeptidome (pink) and in the whole proteome (blue) of GBM11. f. Number of nuORFs (x axis) of different types (y axis) identified in the MHC I immunopeptidome (left) vs. whole proteome (hatched, right) in GBM11. g. Protein length (x axis, amino acids) of annotated (top) and nuORF (bottom) proteins detected in the MHC I immunopeptidome (pink) vs. in the whole proteome (blue). p-values: KS test. h. Proportion of all annotated ORFs (top) or nuORFs (bottom) detected in the whole proteome (blue), immunopeptidome (pink) or both (intersection) in GBM11.
Extended Data Fig. 7 nuORFs can be potential sources of neoantigens.
a. Approaches to identify potential nuORF-derived neoantigens. b. nuORFs have low sequence coverage by WES compared to WGS. Distribution of WES read coverage (x axis) across different ORF types (y axis). Bottom: WGS read coverage across all ORFs of all types. Vertical red line marks 30x coverage. n = 86421 (annotated), 61398 (lncRNA), 61248 (Out-of-frame), 33823 (5’ uORF), 31453 (3’ dORF), 20337 (5’ overlap uORF), 18316 (3’ overlap dORF), 7941 (Pseudogene), 2371 (Other), 323846 (WGS). Median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown. c. Somatic variants in the melanoma patient-derived cell line reflect the variants detected in the original tumor. Cancer-specific SNVs and InDels identified by WES from the primary tumor and by WGS from the tumor-derived cell line. d. Ribo-seq can be used to identify translated variants. Example of a translated SLC7A1 5’ uORF with a cancer-specific SNV. Top: histogram of Ribo-seq reads supporting the translation of the 5’ uORF. Middle: Ribo-seq reads supporting translation of the mutant (green) and wild-type alleles. Predicted neoantigen outlined in red.
Extended Data Fig. 8 SNVs in nuORFs expand the potential neoantigen repertoire.
a. PCAWG-TCGA analysis of SNVs in annotated ORFs and nuORFs. Number of all, transcribed (RNA-seq support), and transcribed nonsynonymous SNVs (y axis) in annotated ORFs and nuORFs (x axis) in CLL, GBM, and SKCM. In CLL, 2/73 samples had no transcribed SNVs, and 3/73 patients had no transcribed nonsynonymous SNVs. n = 73 (CLL,All), 71 (CLL, Expressed), 70 (CLL, Expressed nonsynonymous), 33 (GBM), 36 (SKCM) independent samples. Median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown. b. nuORFs with SNVs are translated in unrelated CLL samples. Number (left) and fraction (right) of transcribed nonsynonymous nuORF SNVs detected across 70 CLL samples (y axis) with Ribo-seq TPM > 0 in 0 or more unrelated CLL samples profiled by Ribo-seq (x axis). c. Transcription frequently indicates translation for annotated ORFs and nuORFs. Percent of annotated (grey) and nuORFs (pink) with RNA-seq and Ribo-seq support (y axis) in two CLL samples (x axis).
Extended Data Fig. 9 GBM and melanoma specific nuORFs.
a. RNA-seq expression (y axis, log2(TPM+1)) of GBM-specific nuORFs (x axis) in GTEx and tumor samples. b. Melanoma-specific nuORFs. RNA-seq expression (y axis, log2(TPM+1)) of melanoma-specific nuORFs (x axis) in GTEx and tumor samples. For all boxplots, n = 390 (CLL), 172 (GBM), 473 (SKCM), 10 donors/tissue across 31 tissues (GTEx). Median, with 25% and 75% (box range), and 1.5 IQR (whiskers) are shown.
Extended Data Fig. 10 GBM nuORFs.
a. Some nuORFs predicted to be GBM-specific are translated in non-cancerous samples. RNA-seq and Ribo-seq expression (log2(TPM+1)) of nuORFs predicted to be GBM-specific (y axis) in published primary GBM and non-cancer brain samples and differentiating hESCs (x axis). b. nuORFs are detected in published GBM and non-cancerous MHC I immunopeptidomes. Number of MS-detected nuORFs (x axis) of different types (y axis) in GBM (right) and non-cancerous brain (left) samples. c. LC-MS/MS spectrum of a peptide from SOX2-OT nuORF.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2, Methods and Figs. 1 and 2.
Supplementary Table 1
nuORFdb samples.
Supplementary Table 2
nuORFdb v.1.0 annotations.
Supplementary Table 3
B721 MHC-I peptides.
Supplementary Table 4
MS-detected ORFs.
Supplementary Table 5
Spliced peptides.
Supplementary Table 6
Whole proteome peptides B721.
Supplementary Table 7
Patient information.
Supplementary Table 8
Cancers MHC-I peptides.
Supplementary Table 9
Whole proteome peptides GBM11.
Supplementary Table 10
Results of peptide binding assays.
Supplementary Table 11
Cancer-specific ORFs.
Supplementary Table 12
Noncancer brain peptides, external data.
Supplementary Table 13
ORF type map.
Supplementary Table 14
CLL samples information.
Rights and permissions
About this article
Cite this article
Ouspenskaia, T., Law, T., Clauser, K.R. et al. Unannotated proteins expand the MHC-I-restricted immunopeptidome in cancer. Nat Biotechnol 40, 209–217 (2022). https://doi.org/10.1038/s41587-021-01021-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01021-3
This article is cited by
-
Non-canonical antigens are the largest fraction of peptides presented by MHC class I in mismatch repair deficient murine colorectal cancer
Genome Medicine (2024)
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
MARS an improved de novo peptide candidate selection method for non-canonical antigen target discovery in cancer
Nature Communications (2024)
-
Immunopeptidomics-based identification of naturally presented non-canonical circRNA-derived peptides
Nature Communications (2024)
-
Challenges in developing personalized neoantigen cancer vaccines
Nature Reviews Immunology (2024)