Abstract
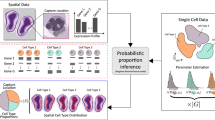
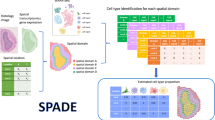
A limitation of spatial transcriptomics technologies is that individual measurements may contain contributions from multiple cells, hindering the discovery of cell-type-specific spatial patterns of localization and expression. Here, we develop robust cell type decomposition (RCTD), a computational method that leverages cell type profiles learned from single-cell RNA-seq to decompose cell type mixtures while correcting for differences across sequencing technologies. We demonstrate the ability of RCTD to detect mixtures and identify cell types on simulated datasets. Furthermore, RCTD accurately reproduces known cell type and subtype localization patterns in Slide-seq and Visium datasets of the mouse brain. Finally, we show how RCTD’s recovery of cell type localization enables the discovery of genes within a cell type whose expression depends on spatial environment. Spatial mapping of cell types with RCTD enables the spatial components of cellular identity to be defined, uncovering new principles of cellular organization in biological tissue. RCTD is publicly available as an open-source R package at https://github.com/dmcable/RCTD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Slide-seqV2 data generated for this study are available at the Broad Institute Single Cell Portal at https://singlecell.broadinstitute.org/single_cell/study/SCP948. Additional publicly available data from other studies that were used for analysis are also included in this repository.
Code availability
RCTD is implemented in the open-source R package RCTD, with source code freely available at https://github.com/dmcable/RCTD. Additional code used for analysis in this paper is available at https://github.com/dmcable/RCTD/tree/dev/AnalysisPaper.
References
Stickels, R. R. et al. Sensitive spatial genome wide expression profiling at cellular resolution. Nature Biotechnology (in the press).
10x Genomics. 10x Genomics: Visium spatial gene expression (2020).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Pelkey, K. A. et al. Hippocampal GABAergic inhibitory interneurons. Physiol. Rev. 97, 1619–1747 (2017).
Cembrowski, M. S. et al. The subiculum is a patchwork of discrete subregions. elife 7, e37701 (2018).
Edsgärd, D., Johnsson, P. & Sandberg, R. Identification of spatial expression trends in single-cell gene expression data. Nat. Methods 15, 339–342 (2018).
Sun, S., Zhu, J. & Zhou, X. Statistical analysis of spatial expression patterns for spatially resolved transcriptomic studies. Nat. Methods 17, 193–200 (2020).
Svensson, V., Teichmann, S. A. & Stegle, O. SpatialDE: identification of spatially variable genes. Nat. Methods 15, 343–346 (2018).
Wagner, A., Regev, A. & Yosef, N. Revealing the vectors of cellular identity with single-cell genomics. Nat. Biotechnol. 34, 1145–1160 (2016).
Regev, A. et al. Science forum: the Human Cell Atlas. eLife 6, e27041 (2017).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Moncada, R. et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat. Biotechnol. 38, 333–342 (2020).
Townes, F. W., Hicks, S. C., Aryee, M. J. & Irizarry, R. A. Feature selection and dimension reduction for single-cell RNA-seq based on a multinomial model. Genome Biol. 20, 295 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Pliner, H. A., Shendure, J. & Trapnell, C. Supervised classification enables rapid annotation of cell atlases. Nat. Methods 16, 983–986 (2019).
Leek, J. T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat. Rev. Genet. 11, 733–739 (2010).
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS ONE 13, e0209648 (2018).
Tsoucas, D. et al. Accurate estimation of cell-type composition from gene expression data. Nat. Commun. 10, 2975 (2019).
Kozareva, V. et al. A transcriptomic atlas of the mouse cerebellum reveals regional specializations and novel cell types. Preprint at bioRxiv https://doi.org/10.1101/2020.03.04.976407 (2020).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030 (2018).
Brown, A. M. et al. Molecular layer interneurons shape the spike activity of cerebellar Purkinje cells. Sci. Rep. 9, 1742 (2019).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Zhang, M. et al. Molecular, spatial and projection diversity of neurons in primary motor cortex revealed by in situ single-cell transcriptomics. Preprint at bioRxiv https://doi.org/10.1101/2020.06.04.105700 (2020).
Sunkin, S. M. et al. Allen Brain Atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res. 41, D996–D1008 (2012).
Capogna, M. Neurogliaform cells and other interneurons of stratum lacunosum-moleculare gate entorhinal–hippocampal dialogue. J. Physiol. 589, 1875–1883 (2011).
Leão, R. N. et al. OLM interneurons differentially modulate CA3 and entorhinal inputs to hippocampal CA1 neurons. Nat. Neurosci. 15, 1524–1530 (2012).
Gampe, K. et al. NTPDase2 and purinergic signaling control progenitor cell proliferation in neurogenic niches of the adult mouse brain. Stem Cells 33, 253–264 (2015).
Dikow, N. et al. 3p25.3 microdeletion of GABA transporters SLC6A1 and SLC6A11 results in intellectual disability, epilepsy and stereotypic behavior. Am. J. Med. Genet. A 164, 3061–3068 (2014).
Lee, T.-S. et al. GAT1 and GAT3 expression are differently localized in the human epileptogenic hippocampus. Acta Neuropathol. 111, 351–363 (2006).
Kulkarni, A., Anderson, A. G., Merullo, D. P. & Konopka, G. Beyond bulk: a review of single cell transcriptomics methodologies and applications. Curr. Opin. Biotechnol. 58, 129–136 (2019).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Sakamoto, Y., Ishiguro, M. & Kitagawa, G. Akaike Information Criterion Statistics 1st edn, Vol. 1 (Springer Netherlands, 1986).
Zhou, M., Li, L., Dunson, D. & Carin, L. Lognormal and gamma mixed negative binomial regression. Proc. Int. Conf. Mach. Learn. 2012, 1343–1350 (2012).
Swami, A. Non-Gaussian mixture models for detection and estimation in heavy-tailed noise. In Proceedings of the 2000 IEEE International Conference on Acoustics, Speech, and Signal Processing 3802–3805 (IEEE, 2000).
Turlach, B. A. & Weingessel, A. quadprog: functions to solve quadratic programming problems. R package version 1.5-5 (2013).
Duchi, J. Sequential convex programming, notes for EE364b: Convex Optimization II (Stanford University, 2018).
SatijaLab. Analysis, visualization, and integration of spatial datasets with Seurat. https://satijalab.org/seurat/articles/spatial_vignette.html (2020).
Acknowledgements
We thank R. Stickels for providing valuable input on the analysis. We thank members of the Chen lab, Irizarry lab and Macosko lab for helpful discussions. D.M.C. was supported by a Fannie and John Hertz Foundation Fellowship and an NSF Graduate Research Fellowship. This work was supported by an NIH Early Independence Award (1DP5OD024583 to F.C.), the Burroughs Wellcome Fund (F.C.) and the NHGRI (R01HG010647 to E.Z.M. and F.C.) as well as the Schmidt Fellows Program at the Broad Institute and the Stanley Center for Psychiatric Research. R.A.I. was supported by NIH grants R35GM131802 and R01HG005220.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
D.M.C., F.C., R.A.I. and E.Z.M. conceived the study. F.C., E.M. and E.Z.M. designed the Slide-seq experiment. E.M. generated the Slide-seq data. D.M.C., R.A.I. and F.C. developed the statistical methods. D.M.C., F.C., R.A.I. and E.Z.M. designed the analysis. D.M.C., R.A.I., F.C., A.G. and L.S.Z. analyzed the data. D.M.C., F.C., R.A.I., E.Z.M. and L.S.Z. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Table 1 and Figs. 1–27.
Rights and permissions
About this article
Cite this article
Cable, D.M., Murray, E., Zou, L.S. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat Biotechnol 40, 517–526 (2022). https://doi.org/10.1038/s41587-021-00830-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-00830-w
This article is cited by
-
vissE: a versatile tool to identify and visualise higher-order molecular phenotypes from functional enrichment analysis
BMC Bioinformatics (2024)
-
Multi-slice spatial transcriptome domain analysis with SpaDo
Genome Biology (2024)
-
Niche-DE: niche-differential gene expression analysis in spatial transcriptomics data identifies context-dependent cell-cell interactions
Genome Biology (2024)
-
Architecting the metabolic reprogramming survival risk framework in LUAD through single-cell landscape analysis: three-stage ensemble learning with genetic algorithm optimization
Journal of Translational Medicine (2024)
-
Gene panel selection for targeted spatial transcriptomics
Genome Biology (2024)