Abstract
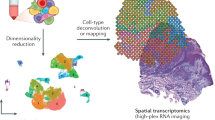
Measurement of the location of molecules in tissues is essential for understanding tissue formation and function. Previously, we developed Slide-seq, a technology that enables transcriptome-wide detection of RNAs with a spatial resolution of 10 μm. Here we report Slide-seqV2, which combines improvements in library generation, bead synthesis and array indexing to reach an RNA capture efficiency ~50% that of single-cell RNA-seq data (~10-fold greater than Slide-seq), approaching the detection efficiency of droplet-based single-cell RNA-seq techniques. First, we leverage the detection efficiency of Slide-seqV2 to identify dendritically localized mRNAs in neurons of the mouse hippocampus. Second, we integrate the spatial information of Slide-seqV2 data with single-cell trajectory analysis tools to characterize the spatiotemporal development of the mouse neocortex, identifying underlying genetic programs that were poorly sampled with Slide-seq. The combination of near-cellular resolution and high transcript detection efficiency makes Slide-seqV2 useful across many experimental contexts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Code availability
Code related to this manuscript can be found at https://github.com/MacoskoLab/slideseq-tools and https://github.com/rstickels/Slide_seqv2. The following package version numbers were used for data processing and associated analyses: https://github.com/broadinstitute/Drop-seq (Drop-seq-tools-2.3.0), https://broadinstitute.github.io/picard/ (picard-2.18.14), https://github.com/alexdobin/STAR (STAR-2.5.2a), https://github.com/theislab/scvelo (0.1.25), https://github.com/cole-trapnell-lab/monocle3 (beta) and https://github.com/satijalab/seurat (2.3.4). MATLAB 2017a, R3.5.3 and Python 3.7 were used for processing data.
References
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Codeluppi, S. et al. Spatial organization of the somatosensory cortex revealed by osmFISH. Nat. Methods 15, 932–935 (2018).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Hughes, T. K. et al. Second-strand synthesis-based massively parallel scRNA-seq reveals cellular states and molecular features of human inflammatory skin pathologies. Immunity 53, 878–894 (2020).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030 (2018).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Raj, A., van den Bogaard, P., Rifkin, S. A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145, dev165753 (2018).
Govindarajan, A., Israely, I., Huang, S.-Y. & Tonegawa, S. The dendritic branch is the preferred integrative unit for protein synthesis-dependent LTP. Neuron 69, 132–146 (2011).
Richter, J. D. & Klann, E. Making synaptic plasticity and memory last: mechanisms of translational regulation. Genes Dev. 23, 1–11 (2009).
Huber, K. M., Kayser, M. S. & Bear, M. F. Role for rapid dendritic protein synthesis in hippocampal mGluR-dependent long-term depression. Science 288, 1254–1257 (2000).
Kosik, K. S. Life at low copy number: how dendrites manage with so few mRNAs. Neuron 92, 1168–1180 (2016).
Ainsley, J. A., Drane, L., Jacobs, J., Kittelberger, K. A. & Reijmers, L. G. Functionally diverse dendritic mRNAs rapidly associate with ribosomes following a novel experience. Nat. Commun. 5, 4510 (2014).
Tushev, G. et al. Alternative 3′ UTRs modify the localization, regulatory potential, stability, and plasticity of mRNAs in neuronal compartments. Neuron 98, 495–511 (2018).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Welch, J. D., Hartemink, A. J. & Prins, J. F. SLICER: inferring branched, nonlinear cellular trajectories from single cell RNA-seq data. Genome Biol. 17, 106 (2016).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Lodato, S. & Arlotta, P. Generating neuronal diversity in the mammalian cerebral cortex. Annu. Rev. Cell Dev. Biol. 31, 699–720 (2015).
Telley, L. et al. Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364, eaav2522 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0591-3 (2020).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Ruediger, T. et al. Integration of opposing semaphorin guidance cues in cortical axons. Cereb. Cortex 23, 604–614 (2013).
Polleux, F., Giger, R. J., Ginty, D. D., Kolodkin, A. L. & Ghosh, A. Patterning of cortical efferent projections by semaphorin–neuropilin interactions. Science 282, 1904–1906 (1998).
Heavner, W. & Pevny, L. Eye development and retinogenesis. Cold Spring Harb. Perspect. Biol. 4, a008391 (2012).
Ashery-Padan, R., Marquardt, T., Zhou, X. & Gruss, P. Pax6 activity in the lens primordium is required for lens formation and for correct placement of a single retina in the eye. Genes Dev. 14, 2701–2711 (2000).
Barbieri, A. M. et al. A homeobox gene, vax2, controls the patterning of the eye dorsoventral axis. Proc. Natl Acad. Sci. USA 96, 10729–10734 (1999).
Andley, U. P. Crystallins in the eye: function and pathology. Prog. Retin. Eye Res. 26, 78–98 (2007).
Niederreither, K., Subbarayan, V., Dollé, P. & Chambon, P. Embryonic retinoic acid synthesis is essential for early mouse post-implantation development. Nat. Genet. 21, 444–448 (1999).
Fan, X. et al. Targeted disruption of Aldh1a1 (Raldh1) provides evidence for a complex mechanism of retinoic acid synthesis in the developing retina. Mol. Cell. Biol. 23, 4637–4648 (2003).
Snead, M. P. et al. Stickler syndrome, ocular-only variants and a key diagnostic role for the ophthalmologist. Eye 25, 1389–1400 (2011).
Fares-Taie, L. et al. ALDH1A3 mutations cause recessive anophthalmia and microphthalmia. Am. J. Hum. Genet. 92, 265–270 (2013).
Martin, H. C. et al. Quantifying the contribution of recessive coding variation to developmental disorders. Science 362, 1161–1164 (2018).
Kaplanis, J. et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586, 757–762 (2020).
McKernan, K. J. et al. Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Storey, J. D. A direct approach to false discovery rates. J. R. Stat. Soc. B Stat. Methodol. 64, 479–498 (2002).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Carlson M.org.Mm.eg.db genome wide annotation for mouse. R package version 3.8.2 http://bioconductor.org/packages/release/data/annotation/html/org.Mm.eg.db.html (2019).
Acknowledgements
We thank J. Dimidschstein and G. Fishell for their helpful advice on the developmental trajectory analysis. This work was supported by an NIH New Innovator Award (DP2 AG058488-01 to E.Z.M.), an NIH Early Independence Award (DP5, 1DP5OD024583 to F.C.), the NHGRI (R01, R01HG010647 to E.Z.M. and F.C.), the Burroughs Wellcome Fund CASI award (to F.C.) and the Schmidt Fellows Program at the Broad Institute and the Stanley Center for Psychiatric Research.
Author information
Authors and Affiliations
Contributions
F.C. and E.Z.M. supervised the work. R.R.S. and E.M. performed experiments. D.J.D. and P.A. contributed to experiments on the developing neocortex. R.R.S., F.C. and E.Z.M. analyzed the data. J.L. developed the Slide-seq tools software package. P.K. developed the bead synthesis protocol. J.L.M. performed optimization experiments. F.C., E.Z.M., R.R.S. and E.M. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
R.R.S., F.C. and E.Z.M. are listed as inventors on a pending patent application related to the development of Slide-seq.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9 and Tables 1, 2 and 6–8.
Supplementary Dataset 1
Plots of all genes dendritically enriched in Slide-seqV2.
Supplementary Dataset 2
Plots of all genes called as spatially significant in Slide-seqV2.
Supplementary Table 3
Dendritically enriched gene sets.
Supplementary Table 4
List of all genes called as spatially significant for Slide-seqV2 data in the developing cortex and eye.
Supplementary Table 5
List of genes unique to each method regarding the trajectory inference.
Rights and permissions
About this article
Cite this article
Stickels, R.R., Murray, E., Kumar, P. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat Biotechnol 39, 313–319 (2021). https://doi.org/10.1038/s41587-020-0739-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0739-1
This article is cited by
-
spVC for the detection and interpretation of spatial gene expression variation
Genome Biology (2024)
-
Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases
Genome Medicine (2024)
-
Multi-slice spatial transcriptome domain analysis with SpaDo
Genome Biology (2024)
-
Evaluating spatially variable gene detection methods for spatial transcriptomics data
Genome Biology (2024)
-
SRT-Server: powering the analysis of spatial transcriptomic data
Genome Medicine (2024)