Abstract
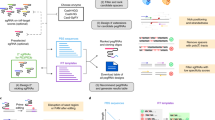
Prime editing enables the introduction of virtually any small-sized genetic change without requiring donor DNA or double-strand breaks. However, evaluation of prime editing efficiency requires time-consuming experiments, and the factors that affect efficiency have not been extensively investigated. In this study, we performed high-throughput evaluation of prime editor 2 (PE2) activities in human cells using 54,836 pairs of prime editing guide RNAs (pegRNAs) and their target sequences. The resulting data sets allowed us to identify factors affecting PE2 efficiency and to develop three computational models to predict pegRNA efficiency. For a given target sequence, the computational models predict efficiencies of pegRNAs with different lengths of primer binding sites and reverse transcriptase templates for edits of various types and positions. Testing the accuracy of the predictions using test data sets that were not used for training, we found Spearman’s correlations between 0.47 and 0.81. Our computational models and information about factors affecting PE2 efficiency will facilitate practical application of prime editing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The deep sequencing data from this study have been submitted to the National Center for Biotechnology Information Sequence Read Archive under accession number PRJNA624815. The data sets used in this study are provided as Supplementary Tables 3, 4 and 5.
Code availability
Source codes for DeepPE and the custom Python script used for the prime editing efficiency calculations are provided as Supplementary Codes 1 and 2 and are also available at https://github.com/hkimlab-PE/PE_SupplementaryCode.
Change history
08 February 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41587-024-02159-6
References
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Lin, Q. et al. Prime genome editing in rice and wheat. Nat. Biotechnol. 38, 582–585 (2020).
Chari, R., Mali, P., Moosburner, M. & Church, G. M. Unraveling CRISPR–Cas9 genome engineering parameters via a library-on-library approach. Nat. Methods 12, 823–826 (2015).
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR–Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Kim, H. K. et al. In vivo high-throughput profiling of CRISPR–Cpf1 activity. Nat. Methods 14, 153–159 (2017).
Allen, F. et al. Predicting the mutations generated by repair of Cas9-induced double-strand breaks. Nat. Biotechnol. 37, 64–72 (2018).
Kim, H. K. et al. Deep learning improves prediction of CRISPR–Cpf1 guide RNA activity. Nat. Biotechnol. 36, 239–241 (2018).
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018).
Kim, H. K. et al. SpCas9 activity prediction by DeepSpCas9, a deep learning-based model with high generalization performance. Sci. Adv. 5, eaax9249 (2019).
Kim, H. K. et al. High-throughput analysis of the activities of xCas9, SpCas9-NG and SpCas9 at matched and mismatched target sequences in human cells. Nat. Biomed. Eng. 4, 111–124 (2020).
Song, M. et al. Sequence-specific prediction of the efficiencies of adenine and cytosine base editors. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0453-z (2020).
Arbab, M. et al. Determinants of base editing outcomes from target library analysis and machine learning. Cell 182, 463–480 (2020).
Kim, N. et al. Prediction of the sequence-specific cleavage activity of Cas9 variants. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0537-9 (2020).
Wang, D. et al. Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning. Nat. Commun. 10, 4284 (2019).
Schlub, T. E., Smyth, R. P., Grimm, A. J., Mak, J. & Davenport, M. P. Accurately measuring recombination between closely related HIV-1 genomes. PLoS Comput. Biol. 6, e1000766 (2010).
Sack, L. M., Davoli, T., Xu, Q., Li, M. Z. & Elledge, S. J. Sources of error in mammalian genetic screens. G3 (Bethesda) 6, 2781–2790 (2016).
Feldman, D., Singh, A., Garrity, A. J. & Blainey, P. C. Lentiviral co-packaging mitigates the effects of intermolecular recombination and multiple integrations in pooled genetic screens. Preprint at https://doi.org/10.1101/262121 (2018).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
Dang, Y. et al. Optimizing sgRNA structure to improve CRISPR–Cas9 knockout efficiency. Genome Biol. 16, 280 (2015).
Nielsen, S., Yuzenkova, Y. & Zenkin, N. Mechanism of eukaryotic RNA polymerase III transcription termination. Science 340, 1577–1580 (2013).
Lin, Y. et al. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014).
Chen, H., Choi, J. & Bailey, S. Cut site selection by the two nuclease domains of the Cas9 RNA-guided endonuclease. J. Biol. Chem. 289, 13284–13294 (2014).
Zeng, Y. et al. The initiation, propagation and dynamics of CRISPR–SpyCas9 R-loop complex. Nucleic Acids Res. 46, 350–361 (2018).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Anders, C., Bargsten, K. & Jinek, M. Structural plasticity of PAM recognition by engineered variants of the RNA-guided endonuclease Cas9. Mol. Cell. 61, 895–902 (2016).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Nishimasu, H. et al. Engineered CRISPR–Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Miller, S. M. et al. Continuous evolution of SpCas9 variants compatible with non-G PAMs. Nat. Biotechnol. 38, 471–481 (2020).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR–Cas9 variants. Science 368, 290–296 (2020).
Du, D. et al. Genetic interaction mapping in mammalian cells using CRISPR interference. Nat. Methods 14, 577–580 (2017).
Shen, J. P. et al. Combinatorial CRISPR–Cas9 screens for de novo mapping of genetic interactions. Nat. Methods 14, 573–576 (2017).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. Preprint at https://arxiv.org/abs/1603.02754 (2016).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Mnih, V. et al. Human-level control through deep reinforcement learning. Nature 518, 529–533 (2015).
Abadi, M. et al. In Proc. of the 12th USENIX Conference on Operating Systems Design and Implementation 265–283 (USENIX Association, 2016).
Acknowledgements
We would like to thank D. Kim, S. Park and Y. Kim for assisting with the experiments. This work was supported, in part, by the National Research Foundation of Korea (grants 2017R1A2B3004198 (H.H.K.), 2017M3A9B4062403 (H.H.K.), 2020R1C1C1003284 (H.K.K) and 2018R1A5A2025079 (H.H.K)), the Brain Korea 21 Plus Project (Yonsei University College of Medicine) and the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (grants HI17C0676 (H.H.K.) and HI16C1012 (H.H.K.)).
Author information
Authors and Affiliations
Contributions
G.Y. and H.K.K. performed the wet experiments, including high-throughput evaluation of PE2 efficiencies. S.M., S.L., S.Y. and H.K.K. developed DeepPE and the related web tools. J.P. substantially contributed to bioinformatics analyses and DeepPE development. H.K.K. and H.H.K. conceived of and designed the study. H.K.K., G.Y. and H.H.K. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Yonsei University has filed a patent application based on this work, in which H.K.K., G.Y. and H.H.K. are listed as inventors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Texts 1–3, Supplementary Figs. 1–23 and Supplementary Tables 1 and 2.
Supplementary Table 3
Data sets of PE2 efficiencies at endogenous sites
Supplementary Table 4
Data sets HT-training, HT-test, Type-training, Type-test, Position-training and Position-test
Supplementary Table 5
Data sets of PE2 efficiencies generated using HCT116 and MDA-MB-231 cells
Supplementary Table 6
Oligonucleotides used in this study
Supplementary Table 7
Exact P values for Figs. 2 and 3 and Supplementary Fig. 15
Supplementary Software 1
Codes relevant to the PE efficiency analysis
Supplementary Software 2
Codes relevant to DeepPE
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, H.K., Yu, G., Park, J. et al. Predicting the efficiency of prime editing guide RNAs in human cells. Nat Biotechnol 39, 198–206 (2021). https://doi.org/10.1038/s41587-020-0677-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0677-y
This article is cited by
-
Efficient prime editing in two-cell mouse embryos using PEmbryo
Nature Biotechnology (2024)
-
High-throughput evaluation of genetic variants with prime editing sensor libraries
Nature Biotechnology (2024)
-
Recent advances in CRISPR-based functional genomics for the study of disease-associated genetic variants
Experimental & Molecular Medicine (2024)
-
BacPE: a versatile prime-editing platform in bacteria by inhibiting DNA exonucleases
Nature Communications (2024)
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)