Abstract
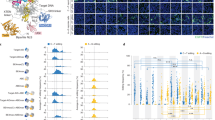
Although base editors are useful tools for precise genome editing, current base editors can only convert either adenines or cytosines. We developed a dual adenine and cytosine base editor (A&C-BEmax) by fusing both deaminases with a Cas9 nickase to achieve C-to-T and A-to-G conversions at the same target site. Compared to single base editors, A&C-BEmax’s activity on adenines is slightly reduced, whereas activity on cytosines is higher and RNA off-target activity is substantially decreased.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Targeted amplicon sequencing data have been deposited in the NCBI Sequence Read Archive database under accession codes PRJNA558944, PRJNA559260, PRJNA559237, PRJNA559051, PRJNA592341 and PRJNA592333. The RNA-sequencing data used in this study have been deposited in the NCBI Sequence Read Archive database under accession code PRJNA592597. Plasmids encoding A&C-BEmax, AID-BE4max, Lenti ABE7.10-N-AIDmax and Lenti AID-BEmax are available from Addgene.
References
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Rees, H. A. & Liu, D. R. J. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Bae, S., Park, J. & Kim, J.-S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Gehrke, J. M. et al. An APOBEC3A–Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. 36, 977–982 (2018).
Zuo, E. et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos. Science 364, 289–292 (2019).
Jin, S. et al. Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science 364, 292–295 (2019).
Zhou, C. et al. Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis. Nature 571, 275–278 (2019).
Grünewald, J. et al. CRISPR DNA base editors with reduced RNA off-target and self-editing activities. Nat. Biotechnol. 37, 1041–1048 (2019).
Wienert, B., Martyn, G. E., Funnell, A. P., Quinlan, K. G. & Crossley, M. J. Wake-up sleepy gene: reactivating fetal globin for β-hemoglobinopathies. Trends Genet. 34, 927–940 (2018).
Martyn, G. E. et al. Natural regulatory mutations elevate the fetal globin gene via disruption of BCL11A or ZBTB7A binding. Nat. Genet. 50, 498–503 (2018).
Martyn, G. E. et al. A natural regulatory mutation in the proximal promoter elevates fetal globin expression by creating a de novo GATA1 site. Blood 133, 852–856 (2019).
Kurita, R. et al. Establishment of immortalized human erythroid progenitor cell lines able to produce enucleated red blood cells. PloS ONE 8, e59890 (2013).
Wienert, B. et al. KLF1 drives the expression of fetal hemoglobin in British HPFH. Blood 130, 803–807 (2017).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Wang, Y. et al. Comparison of cytosine base editors and development of the BEable-GPS database for targeting pathogenic SNVs. Genome Biol. 20, 218 (2019).
Nishimasu, H. et al. Engineered CRISPR–Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Yang, L. et al. Increasing targeting scope of adenosine base editors in mouse and rat embryos through fusion of TadA deaminase with Cas9 variants. Protein Cell 9, 814–819 (2018).
Biyun, Z. et al. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-877/v1 (2020).
Hwang, G.-H. et al. Web-based design and analysis tools for CRISPR base editing. BMC Bioinformatics 19, 542 (2018).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Acknowledgements
We thank M. Crossley at the University of New South Wales, Australia, for providing the HUDEP-2(ΔGγ) cell line and H. Yang and C. Zhou at the Institute of Neuroscience, Chinese Academy of Sciences, for the suggestions of RNA off-targeting analysis. This work was partially supported by grants from the National Key R&D Program of China (2019YFA0110802 to D.L. and 2019YFA0802800 to M.L.), the National Natural Science Foundation of China (81670470 and 81873685 to D.L., and 31925011 and 91940306 to L.Y.), grants from the Shanghai Municipal Commission for Science and Technology (18411953500 to D.L.), the Innovation Program of the Shanghai Municipal Education Commission (2019-01-07-00-05-E00054 to D.L.), the support of the ECNU Public Platform for Innovation (011) and Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Contributions
X.Z., D.L. and M.L. designed the experiments and analyzed the data. X.Z., B.Z., L.C., L.X., L.L., W.H., S.Y., L.Y., Ha.H., Ho.H., Y.L., L.W., X.M., X.P. and Y.W. performed experiments. X.Z., W.Y., Z.Y. and G.C. analyzed the transcriptome data. R.K. and Y.N. generated HUDEP-2 cells. Y.W. and Li.Y. performed bioinformatic analysis to profile A&C-BEmax-editable variants and SNPs. D.L. and M.L. supervised the research. D.L., X.Z., S.S., H.G. and B.Z. analyzed the data and wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors have submitted patent applications (nos. 201810929391.3, 201810930979.0 and 201810930980.3) based on the results reported in this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17, Supplementary Note, Supplementary Tables 3–6 and Supplementary Sequences 1–3.
Supplementary Table 1
Number of pathogenic sites or SNPs correctable by A&C-BEmax.
Supplementary Table 2
Analysis of potential codon and amino acid conversion by A&C-BEmax.
Source data
Source Data Fig. 1
Statistical source data.
Rights and permissions
About this article
Cite this article
Zhang, X., Zhu, B., Chen, L. et al. Dual base editor catalyzes both cytosine and adenine base conversions in human cells. Nat Biotechnol 38, 856–860 (2020). https://doi.org/10.1038/s41587-020-0527-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0527-y
This article is cited by
-
CRISPR technologies for genome, epigenome and transcriptome editing
Nature Reviews Molecular Cell Biology (2024)
-
Adenine transversion editors enable precise, efficient A•T-to-C•G base editing in mammalian cells and embryos
Nature Biotechnology (2024)
-
CRISPR/Cas9-mediated base editors and their prospects for mitochondrial genome engineering
Gene Therapy (2024)
-
CRISPR applications in cancer diagnosis and treatment
Cellular & Molecular Biology Letters (2023)
-
Recent advances in therapeutic CRISPR-Cas9 genome editing: mechanisms and applications
Molecular Biomedicine (2023)