Abstract
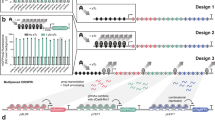
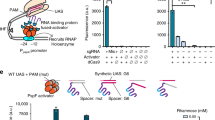
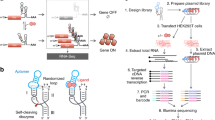
Engineering cellular phenotypes often requires the regulation of many genes. When using CRISPR interference, coexpressing many single-guide RNAs (sgRNAs) triggers genetic instability and phenotype loss, due to the presence of repetitive DNA sequences. We stably coexpressed 22 sgRNAs within nonrepetitive extra-long sgRNA arrays (ELSAs) to simultaneously repress up to 13 genes by up to 3,500-fold. We applied biophysical modeling, biochemical characterization and machine learning to develop toolboxes of nonrepetitive genetic parts, including 28 sgRNA handles that bind Cas9. We designed ELSAs by combining nonrepetitive genetic parts according to algorithmic rules quantifying DNA synthesis complexity, sgRNA expression, sgRNA targeting and genetic stability. Using ELSAs, we created three highly selective phenotypes in Escherichia coli, including redirecting metabolism to increase succinic acid production by 150-fold, knocking down amino acid biosynthesis to create a multi-auxotrophic strain and repressing stress responses to reduce persister cell formation by 21-fold. ELSAs enable simultaneous and stable regulation of many genes for metabolic engineering and synthetic biology applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
High-throughput sequencing data have been deposited in the NCBI Sequence Read Archive database (PRJNA504834). Sanger sequencing analysis is available as a Supplementary Note.
Code availability
A web interface to the ELSA Calculator is available at https://salislab.net/software. Python source code and a Dockerfile are available at https://github.com/hsalis/SalisLabCode.
References
Dominguez, A. A., Lim, W. A. & Qi, L. S. Beyond editing: repurposing CRISPR–Cas9 for precision genome regulation and interrogation. Nat. Rev. Mol. Cell Biol. 17, 5–15 (2016).
Barrangou, R. & Horvath, P. A decade of discovery: CRISPR functions and applications. Nat. Microbiol. 2, 17092 (2017).
Halperin, S. O. et al. CRISPR-guided DNA polymerases enable diversification of all nucleotides in a tunable window. Nature 560, 248–252 (2018).
Peters, J. M. et al. Enabling genetic analysis of diverse bacteria with Mobile-CRISPRi. Nat. Microbiol. 4, 244–250 (2019).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Hess, G. T., Tycko, J., Yao, D. & Bassik, M. C. Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell 68, 26–43 (2017).
Klann, T. S., Black, J. B. & Gersbach, C. A. CRISPR-based methods for high-throughput annotation of regulatory DNA. Curr. Opin. Biotechnol. 52, 32–41 (2018).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882.e1821 (2016).
Swiech, L. et al. In vivo interrogation of gene function in the mammalian brain using CRISPR–Cas9. Nat. Biotechnol. 33, 102–106 (2015).
Yao, L., Cengic, I., Anfelt, J. & Hudson, E. P. Multiple gene repression in cyanobacteria using CRISPRi. ACS Synth. Biol. 5, 207–212 (2015).
Zhao, Y. et al. CRISPR/dCas9‐mediated multiplex gene repression in Streptomyces. Biotechnol. J. 13, 1800121 (2018).
Kim, S. K., Seong, W., Han, G. H., Lee, D.-H. & Lee, S.-G. CRISPR interference-guided multiplex repression of endogenous competing pathway genes for redirecting metabolic flux in Escherichia coli. Microb. Cell Fact. 16, 188 (2017).
Ordon, J. et al. Generation of chromosomal deletions in dicotyledonous plants employing a user‐friendly genome editing toolkit. Plant J. 89, 155–168 (2017).
Hughes, R. A. & Ellington, A. D. Synthetic DNA synthesis and assembly: putting the synthetic in synthetic biology. Cold Spring Harb. Perspect. Biol. 9, a023812 (2017).
Stapley, J., Feulner, P. G., Johnston, S. E., Santure, A. W. & Smadja, C. M. Variation in recombination frequency and distribution across eukaryotes: patterns and processes. Phil. Trans. R. Soc. B 372, 20160455 (2017).
Vos, M. & Didelot, X. A comparison of homologous recombination rates in bacteria and archaea. ISME J. 3, 199 (2009).
Casini, A. et al. A pressure test to make 10 molecules in 90 days: external evaluation of methods to engineer biology. J. Am. Chem. Soc. 140, 4302–4316 (2018).
Najm, F. J. et al. Orthologous CRISPR–Cas9 enzymes for combinatorial genetic screens. Nat. Biotechnol. 36, 179–189 (2018).
Jack, B. R. et al. Predicting the genetic stability of engineered DNA sequences with the EFM calculator. ACS Synth. Biol. 4, 939–943 (2015).
Brophy, J. A. & Voigt, C. A. Principles of genetic circuit design. Nat. Methods 11, 508–520 (2014).
Lovett, S. T. Encoded errors: mutations and rearrangements mediated by misalignment at repetitive DNA sequences. Mol. Microbiol. 52, 1243–1253 (2004).
Shen, P. & Huang, H. V. Homologous recombination in Escherichia coli: dependence on substrate length and homology. Genetics 112, 441–457 (1986).
Tang, N. C. & Chilkoti, A. Combinatorial codon scrambling enables scalable gene synthesis and amplification of repetitive proteins. Nat. Mater. 15, 419–424 (2016).
Chen, Y.-J. et al. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 10, 659–664 (2013).
Casini, A. et al. R2oDNA designer: computational design of biologically neutral synthetic DNA sequences. ACS Synth. Biol. 3, 525–528 (2014).
Lorenz, R. et al. ViennaRNA package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Jiang, F. et al. Structures of a CRISPR–Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Briner, A. E. et al. Guide RNA functional modules direct Cas9 activity and orthogonality. Mol. Cell 56, 333–339 (2014).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Nielsen, A. A. & Voigt, C. A. Multi‐input CRISPR/Cas genetic circuits that interface host regulatory networks. Mol. Syst. Biol. 10, 763 (2014).
Dagdas, Y. S., Chen, J. S., Sternberg, S. H., Doudna, J. A. & Yildiz, A. A conformational checkpoint between DNA binding and cleavage by CRISPR–Cas9. Sci. Adv. 3, eaao0027 (2017).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014).
Farasat, I. & Salis, H. M. A biophysical model of CRISPR/Cas9 activity for rational design of genome editing and gene regulation. PLoS Comput. Biol. 12, e1004724 (2016).
Brophy, J. A. & Voigt, C. A. Antisense transcription as a tool to tune gene expression. Mol. Syst. Biol. 12, 854 (2016).
Nyerges, Á. et al. A highly precise and portable genome engineering method allows comparison of mutational effects across bacterial species. Proc. Natl Acad. Sci. USA 113, 2502–2507 (2016).
Lin, H., Bennett, G. N. & San, K. Y. Genetic reconstruction of the aerobic central metabolism in Escherichia coli for the absolute aerobic production of succinate. Biotechnol. Bioeng. 89, 148–156 (2005).
Li, X.-t et al. tCRISPRi: tunable and reversible, one-step control of gene expression. Sci. Rep. 6, 39076 (2016).
Chen, P.-Y., Qian, Y. & Del Vecchio, D. A model for resource competition in CRISPR-mediated gene repression. Preprint at bioRxiv https://doi.org/10.1101/266015 (2018).
Farasat, I. et al. Efficient search, mapping, and optimization of multi‐protein genetic systems in diverse bacteria. Mol. Syst. Biol. 10, 731 (2014).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
de Kok, S. et al. Rapid and reliable DNA assembly via ligase cycling reaction. ACS Synth. Biol. 3, 97–106 (2014).
Khlebnikov, A., Datsenko, K. A., Skaug, T., Wanner, B. L. & Keasling, J. D. Homogeneous expression of the PBAD promoter in Escherichia coli by constitutive expression of the low-affinity high-capacity AraE transporter. Microbiology 147, 3241–3247 (2001).
Ledoit, O. & Wolf, M. A well-conditioned estimator for large-dimensional covariance matrices. J. Multivar. Anal. 88, 365–411 (2004).
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Der, B. S. et al. DNAplotlib: programmable visualization of genetic designs and associated data. ACS Synth. Biol. 6, 1115–1119 (2016).
Keseler, I. M. et al. The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 45, D543–D550 (2016).
Gama-Castro, S. et al. RegulonDB version 9.0: high-level integration of gene regulation, coexpression, motif clustering and beyond. Nucleic Acids Res. 44, D133–D143 (2015).
Acknowledgements
We thank the Synthetic Biology Application Support Team at Integrated DNA Technologies (IDT) for providing insights into the gene synthesis process, the Penn State Proteomics and Mass Spectrometry Core Facility for the LC-MS analysis, the CSL Behring Fermentation Facility for use of their HPLC/RI and C. Praul and the Penn State Genomics Core Facility for technical support. This project was supported by funds from the Air Force Office of Scientific Research (grant no. FA9550-14-1-0089), an NSF Career Award to H.M.S. (grant no. CBET-1253641), the Defense Advanced Research Projects Agency (grant no. FA8750-17-C-0254) and the Department of Energy (grant no. DE-SC0019090).
Author information
Authors and Affiliations
Contributions
H.M.S., A.C.R. and S.M.H. conceived the study, designed the experiments and wrote the manuscript. A.C.R., S.M.H., P.R.C., A.H., D.P.C. and G.E.V. carried out experiments. A.C.R., S.M.H. and A.H. developed algorithms and performed data analysis.
Corresponding author
Ethics declarations
Competing interests
H.M.S. is the founder of De Novo DNA, which received funds from the Defense Advanced Research Projects Agency to commercialize this technology (grant no. D17PC00133).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14, Tables 1–4 and Note.
Supplementary Data 1
Genetic part sequences, measurements and calculations.
Supplementary Data 2
ELSA compositions, sequences, measurements and calculations.
Supplementary Data 3
ELSA-Stress: RNA-seq results and analysis.
Supplementary Data 4
ELSA-MultiAux: RNA-seq results and analysis.
Supplementary Data 5
MIQE: minimum information for RT–qPCR experiments.
Rights and permissions
About this article
Cite this article
Reis, A.C., Halper, S.M., Vezeau, G.E. et al. Simultaneous repression of multiple bacterial genes using nonrepetitive extra-long sgRNA arrays. Nat Biotechnol 37, 1294–1301 (2019). https://doi.org/10.1038/s41587-019-0286-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0286-9
This article is cited by
-
Improved prediction of bacterial CRISPRi guide efficiency from depletion screens through mixed-effect machine learning and data integration
Genome Biology (2024)
-
Site-specific transgene integration in chimeric antigen receptor (CAR) T cell therapies
Biomarker Research (2023)
-
Genome-wide promoter responses to CRISPR perturbations of regulators reveal regulatory networks in Escherichia coli
Nature Communications (2023)
-
Massively parallel characterization of engineered transcript isoforms using direct RNA sequencing
Nature Communications (2022)
-
Comparative optimization of combinatorial CRISPR screens
Nature Communications (2022)