Abstract
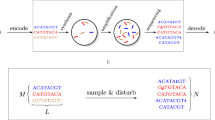
The density and long-term stability of DNA make it an appealing storage medium, particularly for long-term data archiving. Existing DNA storage technologies involve the synthesis and sequencing of multiple nominally identical molecules in parallel, resulting in information redundancy. We report the development of encoding and decoding methods that exploit this redundancy using composite DNA letters. A composite DNA letter is a representation of a position in a sequence that consists of a mixture of all four DNA nucleotides in a predetermined ratio. Our methods encode data using fewer synthesis cycles. We encode 6.4 MB into composite DNA, with distinguishable composition medians, using 20% fewer synthesis cycles per unit of data, as compared to previous reports. We also simulate encoding with larger composite alphabets, with distinguishable composition deciles, to show that 75% fewer synthesis cycles are potentially sufficient. We describe applicable error-correcting codes and inference methods, and investigate error patterns in the context of composite DNA letters.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All raw sequencing data are available from the European Nucleotide Archive (ENA) under accession PRJEB32427. This includes sequencing of the large-scale experiment described in Figs. 2–4, sequencing of the experiment with large alphabets described in Fig. 5 and sequencing of the error analysis experiment described in Fig. 5. All other data are available within the article or its supplementary information.
Code availability
All original software code included in this study is available online. Alteration of the previously published DNA fountain code to support composite DNA is available from https://github.com/leon-anavy/dna-fountain. Code used for Reed–Solomon error correction (altered from previously published code) is available from https://github.com/leon-anavy/Reed-Solomon. Custom code used for the analyses presented in this study is available from https://github.com/leon-anavy/composite-DNA.
Change history
16 September 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Cox, J. P. Long-term data storage in DNA. Trends Biotechnol. 19, 247–250 (2001).
Zhirnov, V., Zadegan, R. M., Sandhu, G. S., Church, G. M. & Hughes, W. L. Nucleic acid memory. Nat. Mater. 15, 366–370 (2016).
Church, G. M., Gao, Y. & Kosuri, S. Next-generation digital information storage in DNA. Science 337, 1628 (2012).
Goldman, N. et al. Towards practical, high-capacity, low-maintenance information storage in synthesized DNA. Nature 494, 77–80 (2013).
Bornholt, J. et al. Toward a DNA-based archival storage system. IEEE Micro 37, 98–104 (2017).
Tabatabaei Yazdi, S. M. H. et al. A rewritable, random-access DNA-based storage system. Sci. Rep. 5, 14138 (2015).
Erlich, Y. & Zielinski, D. DNA Fountain enables a robust and efficient storage architecture. Science 355, 950–954 (2017).
Organick, L. et al. Random access in large-scale DNA data storage. Nat. Biotechnol. 36, 242–248 (2018).
Gabrys, R., Kiah, H. M. & Milenkovic, O. Asymmetric lee distance codes for DNA-based storage. In Proc. 2015 IEEE International Symposium on Information Theory (ISIT) 909–913 (IEEE, 2015)..
Levy, M. & Yaakobi, E. Mutually uncorrelated codes for DNA storage. In Proc. 2017 IEEE International Symposium on Information Theory (ISIT) 3115–3119 (IEEE, 2017).
Lee, H. H., Kalhor, R., Goela, N., Bolot, J. & Church, G. M. Terminator-free template-independent enzymatic DNA synthesis for digital information storage. Nat. Commun. 10, 2383 (2019).
Palluk, S. et al. De novo DNA synthesis using polymerase–nucleotide conjugates. Nat. Biotechnol. 36, 645–650 (2018).
Roquet, N., Park, H. & Bhatia, S. P. Nucleic acid-based data storage. US patent 20180137418 (2017).
LeProust, E. M. et al. Synthesis of high-quality libraries of long (150mer) oligonucleotides by a novel depurination controlled process. Nucleic Acids Res. 38, 2522–2540 (2010).
Barrett, M. T. et al. Comparative genomic hybridization using oligonucleotide microarrays and total genomic DNA. Proc. Natl Acad. Sci. USA 101, 17765–17770 (2004).
Kosuri, S. & Church, G. M. Large-scale de novo DNA synthesis: technologies and applications. Nat. Methods 11, 499–507 (2014).
Choi, Y. et al. High information capacity DNA-based data storage with augmented encoding characters using degenerate bases. Sci. Rep. 9, 6582 (2019).
Grass, R. N., Heckel, R., Puddu, M., Paunescu, D. & Stark, W. J. Robust chemical preservation of digital information on DNA in silica with error-correcting codes. Angew. Chem. Int. Ed. Engl. 54, 2552–2555 (2015).
Reed, I. S. & Solomon, G. Polynomial codes over certain finite fields. J. Soc. Ind. Appl. Math. 8, 300–304 (1960).
MacKay, D. J. C. Fountain codes. IEE Proc. Comm. 152, 1062 (2005).
Jiménez-Sánchez, A. DNA computer code based on expanded genetic alphabet. Eur. J. Comput. Sci. Inf. Technol. 2, 8–20 (2014).
Tabatabaei Yazdi, S. M. H. et al. DNA-based storage: trends and methods. IEEE Trans. Mol. Biol. Multiscale Commun. 1, 230–248 (2015).
Raviv, N., Schwartz, M. & Yaakobi, E. Rank modulation codes for DNA storage. In Proc. 2017 IEEE International Symposium on Information Theory (ISIT) 3125–3129 (IEEE, 2017).
Yazdi, S. M. H. T., Kiah, H. M., Gabrys, R. & Milenkovic, O. Mutually uncorrelated primers for DNA-based data storage. Preprint at https://arxiv.org/abs/1709.05214 (2017).
Takahashi, C. N., Nguyen, B. H., Strauss, K. & Ceze, L. Demonstration of end-to-end automation of DNA data storage. Sci. Rep. 9, 4998 (2019).
Hoshika, S. et al. Hachimoji DNA and RNA: a genetic system with eight building blocks. Science 363, 884–887 (2019).
Bains, W. Hybridization methods for DNA sequencing. Genomics 11, 94–301 (1991).
Pevzner, P. A. Rearrangements of DNA sequences and SBH. Comput. Chem. 18, 221–223 (1994).
Preparata, F. P. & Oliver, J. S. DNA sequencing by hybridization using semi-degenerate bases. J. Comput. Biol. 11, 753–765 (2004).
Snir, S., Yeger-Lotem, E., Chor, B., and Yakhini, Z. Using restriction enzymes to improve sequencing by hybridization. Technical report CS-2002-14 (Technion, 2002).
Chen, Z. et al. Highly accurate fluorogenic DNA sequencing with information theory-based error correction. Nat. Biotechnol. 35, 1170–1178 (2017).
Davidson, E. H. The Regulatory Genome: Gene Regulatory Networks in Development and Evolution (Academic, 2006).
Sandelin, A., Alkema, W., Engström, P., Wasserman, W. W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32, D91–D94 (2004).
Levy, L. et al. A synthetic oligo library and sequencing approach reveals an insulation mechanism encoded within bacterial σ54 promoters. Cell Rep. 21, 845–858 (2017).
Sharon, E. et al. Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nat. Biotechnol. 30, 521–530 (2012).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Mikutis, G. et al. Silica-encapsulated DNA-based tracers for aquifer characterization. Environ. Sci. Technol. 52, 12142–12152 (2018).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina paired-end read merger. Bioinformatics 30, 614–620 (2014).
Shakespeare, W. The Complete Works of William Shakespeare http://www.gutenberg.org/ebooks/100 (1994)
Huffman, D. A. A method for the construction of minimum-redundancy codes. Proc. IRE 40, 1098–1101 (1952).
Acknowledgements
We thank T. Katz-Ezov and T. Hashimshony from the Technion Genome Center for advice and assistance with oligonucleotide design and sequencing experiments. We also thank P. Weiss from Twist Bioscience for technical support and assistance with DNA synthesis. Finally, we thank the Yakhini and Amit research groups for valuable comments and discussions. L. Anavy is supported by the Adams Fellowships Program of the Israel Academy of Sciences and Humanities. This project received funding from the European Union’s Horizon 2020 Research and Innovation Programme under grant agreement 664918 (MRG-Grammar).
Author information
Authors and Affiliations
Contributions
L.A. and Z.Y. initiated and designed the coding and algorithmic approach. L.A. developed the software and performed data analysis. I.V. and O.A. performed the experiments. L.A., R.A. and Z.Y. wrote the manuscript. R.A. and Z.Y. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
L.A, Z.Y and R.A are the inventors of a patent application for the method described in this article. The initial filing was assigned United States Provisional Patent Application No. 62/674,114. The remaining authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17 and Supplementary Note
Supplementary Table 1
Physical density calculations of composite DNA storage. This includes the large-scale experiment and the dilution experiment.
Supplementary Table 2
Logical density calculations of composite DNA storage. This includes all the experiments, theoretical encodings and simulation experiments.
Supplementary Table 3
Oligonucleotide design for large-alphabet experiments and error analysis.
Supplementary Table 4
Oligonucleotide design for the large-scale composite DNA storage.
Supplementary Table 5
Oligonucleotide design for the simulations of large composite alphabet DNA storage.
Supplementary Table 6
Simulation results of large composite alphabet DNA storage
Rights and permissions
About this article
Cite this article
Anavy, L., Vaknin, I., Atar, O. et al. Data storage in DNA with fewer synthesis cycles using composite DNA letters. Nat Biotechnol 37, 1229–1236 (2019). https://doi.org/10.1038/s41587-019-0240-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0240-x
This article is cited by
-
Efficient DNA-based data storage using shortmer combinatorial encoding
Scientific Reports (2024)
-
A universal system for boosting gene expression in eukaryotic cell-lines
Nature Communications (2024)
-
Reconstruction algorithms for DNA-storage systems
Scientific Reports (2024)
-
In-vitro validated methods for encoding digital data in deoxyribonucleic acid (DNA)
BMC Bioinformatics (2023)
-
A biological camera that captures and stores images directly into DNA
Nature Communications (2023)