Abstract
The DNA sequencing technologies in use today produce either highly accurate short reads or less-accurate long reads. We report the optimization of circular consensus sequencing (CCS) to improve the accuracy of single-molecule real-time (SMRT) sequencing (PacBio) and generate highly accurate (99.8%) long high-fidelity (HiFi) reads with an average length of 13.5 kilobases (kb). We applied our approach to sequence the well-characterized human HG002/NA24385 genome and obtained precision and recall rates of at least 99.91% for single-nucleotide variants (SNVs), 95.98% for insertions and deletions <50 bp (indels) and 95.99% for structural variants. Our CCS method matches or exceeds the ability of short-read sequencing to detect small variants and structural variants. We estimate that 2,434 discordances are correctable mistakes in the ‘genome in a bottle’ (GIAB) benchmark set. Nearly all (99.64%) variants can be phased into haplotypes, further improving variant detection. De novo genome assembly using CCS reads alone produced a contiguous and accurate genome with a contig N50 of >15 megabases (Mb) and concordance of 99.997%, substantially outperforming assembly with less-accurate long reads.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data are available in NCBI BioProject PRJNA529679. CCS reads are available on NCBI SRA with accession code SRX5327410. Small variant calls are available on NCBI dbSNP with accession codes ss3783301452–ss3798736595. Structural variant calls are available on NCBI dbVar with accession nstd167. The trio binned Canu assemblies are available on NCBI Assembly with accession codes GCA_004796485.1 (maternal) and GCA_004796285.1 (paternal). Alignments to GRCh37 are available at ftp://ftp-trace.ncbi.nlm.nih.gov/giab/ftp/data/AshkenazimTrio/HG002_NA24385_son/PacBio_CCS_15kb/ or https://bit.ly/2RW1b3I. Additional data, including all assemblies and a track hub for the UCSC Genome Browser, are available at https://downloads.pacbcloud.com/public/publications/2019-HG002-CCS.
Code availability
Custom scripts are available at https://github.com/PacificBiosciences/hg002-ccs/. Google DeepVariant, a model trained on PacBio CCS reads, and instructions for use are available at https://github.com/google/deepvariant.
References
DNA Sequencing Costs: Data (National Human Genome Research Institute, accessed 7th December 2018); https://www.genome.gov/27541954/dna-sequencing-costs-data/
Sanger, F., Nicklen, S. & Coulson, A. R. DNA sequencing with chain-terminating inhibitors. Proc. Natl Acad. Sci. USA 74, 5463–5467 (1977).
Smith, L. M. et al. Fluorescence detection in automated DNA sequence analysis. Nature 321, 674–679 (1986).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Mouse Genome Sequencing Consortium. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Ronaghi, M., Karamohamed, S., Pettersson, B., Uhlén, M. & Nyrén, P. Real-time DNA sequencing using detection of pyrophosphate release. Anal. Biochem. 242, 84–89 (1996).
Bentley, D. R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
McKernan, K. J. et al. Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
Rothberg, J. M. et al. An integrated semiconductor device enabling non-optical genome sequencing. Nature 475, 348–352 (2011).
Sedlazeck, F. J., Lee, H., Darby, C. A. & Schatz, M. C. Piercing the dark matter: bioinformatics of long-range sequencing and mapping. Nat. Rev. Genet. 19, 329–346 (2018).
Chaisson, M. J. P. et al. Resolving the complexity of the human genome using single-molecule sequencing. Nature 517, 608–611 (2015).
Seo, J.-S. et al. De novo assembly and phasing of a Korean human genome. Nature 538, 243–247 (2016).
Cretu Stancu, M. et al. Mapping and phasing of structural variation in patient genomes using nanopore sequencing. Nat. Commun. 8, 1326 (2017).
Chaisson, M. J. P. et al. Multi-platform discovery of haplotype-resolved structural variation in human genomes. Nat. Commun. 10, 1784 (2019).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Mikheyev, A. S. & Tin, M. M. Y. A first look at the Oxford Nanopore MinION sequencer. Mol. Ecol. Resour. 14, 1097–1102 (2014).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Travers, K. J., Chin, C.-S., Rank, D. R., Eid, J. S. & Turner, S. W. A flexible and efficient template format for circular consensus sequencing and SNP detection. Nucleic Acids Res. 38, e159 (2010).
Loomis, E. W. et al. Sequencing the unsequenceable: expanded CGG-repeat alleles of the fragile X gene. Genome Res. 23, 121–128 (2013).
Hebert, P. D. N. et al. A sequel to Sanger: amplicon sequencing that scales. BMC Genomics 19, 219 (2018).
Zook, J. M. et al. Extensive sequencing of seven human genomes to characterize benchmark reference materials. Sci. Data 3, 160025 (2016).
Zook, J. M. et al. An open resource for accurately benchmarking small variant and reference calls. Nat. Biotechnol. 37, 561–566 (2019).
Myers, G. In Algorithms in Bioinformatics (eds Brown, D. & Morgenstern, B.) 52–67 (Springer, 2014).
Mandelker, D. et al. Navigating highly homologous genes in a molecular diagnostic setting: a resource for clinical next-generation sequencing. Genet. Med. 18, 1282–1289 (2016).
Ambardar, S., Gowda, M. & High-Resolution Full-Length, H. L. A. Typing method using third generation (Pac-Bio SMRT) sequencing technology. Methods Mol. Biol. 1802, 135–153 (2018).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Luo, R., Sedlazeck, F. J., Lam, T.-W. & Schatz, M. C. A multi-task convolutional deep neural network for variant calling in single molecule sequencing. Nat. Commun. 10, 998 (2019).
Poplin, R. et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 36, 983–987 (2018).
Patterson, M. et al. WhatsHap: Weighted haplotype assembly for future-generation sequencing reads. J. Comput. Biol. 22, 498–509 (2015).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics. 28, i333–i339 (2012).
Garcia, S. et al. Linked-Read sequencing resolves complex structural variants. Preprint at bioRxiv https://doi.org/10.1101/231662 (2017).
Chin, C.-S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Ruan, J. & Li, H. Fast and accurate long-read assembly with wtdbg2. Preprint at bioRxiv https://doi.org/10.1101/530972 (2019).
Koren, S. et al. De novo assembly of haplotype-resolved genomes with trio binning. Nat. Biotechnol. 36, 1174–1182 (2018).
Fungtammasan, A. & Hannigan, B. How well can we create phased, diploid, human genomes?: An assessment of FALCON-Unzip phasing using a human trio. Preprint at bioRxiv https://doi.org/10.1101/262196 (2018).
Chin, C.-S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Vollger, M. R. et al. Long-read sequence and assembly of segmental duplications. Nat. Methods 16, 88–94 (2019).
Schneider, V. A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Krusche, P. et al. Best practices for benchmarking germline small-variant calls in human genomes. Nat. Biotechnol. 37, 555–560 (2019).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Quinlan, A. R. BEDTools: The Swiss-Army Tool for Genome Feature Analysis. Curr. Protoc. Bioinforma. 47(11), 12.1–34 (2014).
Robinson, J., Soormally, A. R., Hayhurst, J. D. & Marsh, S. G. E. The IPD-IMGT/HLA Database—new developments in reporting HLA variation. Hum. Immunol. 77, 233–237 (2016).
Cleary, J. G. et al. Comparing variant call files for performance benchmarking of next-generation sequencing variant calling pipelines. Preprint at bioRxiv https://doi.org/10.1101/023754 (2015).
Li, H. et al. A synthetic-diploid benchmark for accurate variant-calling evaluation. Nat. Methods 15, 595–597 (2018).
Jeffares, D. C. et al. Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat. Commun. 8, 14061 (2017).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 31, 3210–3212 (2015).
Zerbino, D. R. et al. Ensembl 2018. Nucleic Acids Res. 46, D754–D761 (2018).
Jain, C., Koren, S., Dilthey, A., Phillippy, A. M. & Aluru, S. A fast adaptive algorithm for computing whole-genome homology maps. Bioinformatics. 34, i748–i756 (2018).
Acknowledgements
We would like to thank J. Harting for assistance with HLA typing, K. Robertshaw for figure generation, J. Wilson and J. Ziegle for providing PacBio CLR datasets and J. Puglisi for critical reading of the manuscript. S.K. and A.M.P. were supported by the Intramural Research Program of the National Human Genome Research Institute, National Institutes of Health. This work utilized the computational resources of the NIH HPC Biowulf cluster (https://hpc.nih.gov). This work was supported by NIH grant no. 1R01HG010040 to H.L. and NSFC grant nos. 31571353 and 31822029 to J.R. M.C.S. is funded by the National Science Foundation (grant no. DBI-1350041) and National Institutes of Health (grant no. R01-HG006677). F.J.S. and M.M. are funded by NIH grant no. UM1 HG008898. T.M. acknowledges funding from the German Research Foundation (DFG) (grant nos. 391137747 and 395192176). N.D.O. and J.M.Z. were supported by intramural funding from the National Institute of Standards and Technology and an interagency agreement with the U.S. Food and Drug Administration. This work utilized computational resources of DNAnexus and Google to apply DeepVariant to CCS reads. Certain commercial equipment, instruments or materials are identified to specify adequate experimental conditions or reported results. Such identification does not imply recommendation or endorsement by the National Institute of Standards, nor does it imply that the equipment, instruments or materials identified are necessarily the best available for the purpose.
Author information
Authors and Affiliations
Contributions
A.M.W., D.R.R., M.W.H. and P.P. designed the study. D.R.R. and P.P. developed the sample preparation protocol and performed sample preparation. D.R.R., P.P. and Y.Q. performed sequencing. A.C., A.K., C-S.C., M.A.D. and P.C. adapted the algorithms and implementation of DeepVariant. A.C., A.F., A.K., A.M.P., A.M.W., A.T., C-S.C., D.R.R., F.J.S., G.M., G.T.C., H.L., J.E., J.M.Z., J.R., M.A., M.A.D., M.C.S., M.M., N.D.O., P.C., P.P., R.J.H., S.K., T.M. and W.J.R. performed analysis. A.C., A.M.P., C-S.C., D.R.R., F.J.S., J.M.Z., M.A.D., M.C.S. and M.W.H. supervised analysis. A.C., A.M.W., D.R.R., G.M., J.M.Z., P.P., R.J.H., S.K. and W.J.R. wrote the manuscript. See Supplementary Note for more detailed author contributions. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.M.W., A.T., D.R.R., G.T.C., M.W.H., P.P., R.J.H., W.J.R. and Y.Q. are employees and shareholders of Pacific Biosciences. A.C., A.K., M.A.D. and P.C. are employees and shareholders of Google. A.F. and C-S.C. are employees and shareholders of DNAnexus. A.C. is a shareholder and was an employee of DNAnexus for a portion of this work.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 CCS protocol development.
(a) Distribution of polymerase read lengths for a 10 kb E.coli amplicon library and 30 kb E. coli whole genome library sequenced for 10 hours with identical conditions. (b) Distribution of polymerase read lengths for an 8 kb fragment from a BsaAI-digested lambda library sequenced for 4 hours with (5 hour) and without (0 hour) pre-extension to reduce “early-terminating” reads and select surviving polymerase-template complexes. (c) Sample preparation and sequencing workflow. (d) BioAnalyzer trace for the SMRTbell library, sheared to target 15–20 kb fragments. “FU” is fluorescence units. (e) BioAnalyzer trace for ELF fractions of the SMRTbell library. (f) The fraction centered around 15 kb was used for sequencing.
Supplementary Figure 2 CCS read accuracy and coverage uniformity.
(a) Distribution of accuracy predicted by the CCS algorithm for reads with fewer than 3 passes and at least 3 passes, which we consider a minimum pass count for CCS. Approximately half of reads have 3 or more passes; among those nearly all achieve Q20 predicted accuracy. (b) Distributions of HG002 concordance, measured against the GIAB benchmark, at levels of predicted read accuracy (R2 of median = 0.9980). Orange lines are medians; boxes extend from lower to upper quartiles; whiskers extend 1.5 interquartile distances; n=1,000 reads at each predicted accuracy. (c) Distribution of difference between concordance and predicted read accuracy shows that the prediction is well-calibrated to the empirical concordance. (d) Distribution of coverage in 500 bp windows at non-gap positions in GRCh37. (e) Coverage distributions at levels of [GC] content, measured in 500 bp windows. Orange lines are medians; boxes extend from lower to upper quartiles; whiskers extend 1.5 interquartile distances; n per distribution is listed above the plot.
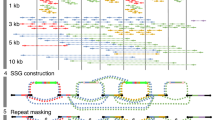
Supplementary Figure 3 CCS read pileups at HLA genes.
The 13.5 kb CCS reads provide phasing and full four-field resolution of HLA class I and II genes (Methods Mol. Biol. 1802, 135–153, 2018), including (a) HLA-A for which HG002 has alleles that differ in the first field, and (b) HLA-DPA1 for which HG002 has alleles that differ only in the fourth field from two intronic single nucleotide polymorphisms across 20 kb.
Supplementary Figure 4 Theoretical phase block N50 in HG002 at different read lengths.
To model the phase blocks achievable with a given read length, cuts were introduced between heterozygous variants in the GIAB trio-phased HG002 variant callset that are separated by more than the read length, which effectively assumes that adjacent heterozygous variants separated by less than the read length can be phased.
Supplementary Figure 5 Structural variant calling performance.
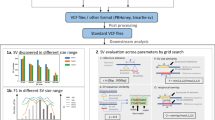
Precision, recall, and number of variant calls in the GIAB benchmark regions for the PacBio CCS mapping-based variant callers (a) pbsv and (b) Sniffles; the PacBio CCS assembly-based callers (c) paftools/Canu (polished) and (d) paftools/FALCON (unpolished); the PacBio CLR mapping-based callers (e) pbsv and (f) Sniffles; the Illumina short-read callers (g) Manta and (h) Delly; and the 10X Genomics callers (i) LongRanger and (j) paftools/Supernova. Negative length indicates a deletion; positive length indicates an insertion. The histogram bin size is 50 bp for variants shorter than 1 kb, and 500 bp for variants >1 kb. Precision and recall are measured with Truvari against the GIAB benchmark.
Supplementary Figure 6 Haplotype resolution in the Canu mixed assembly.
The Canu mixed assembly is larger than the haploid human genome size because it resolves some heterozygous loci into separate maternal and paternal haplotypes. (a, b) Loci where the long primary contig matches the paternal haplotype and a smaller contig matches the maternal haplotype. (c, d) Similar loci where the long primary contig matches the maternal haplotype and a smaller contig matches the paternal haplotype.
Supplementary Figure 7 Mis-phasing analysis of parental assemblies.
Parent-specific heterozygous SNVs were identified in the GIAB benchmark callset. The “Mis-phased SNVs fraction” is the fraction of parent-specific SNVs from the wrong parent (e.g. [SNVpat]/[SNVpat+SNVmat] in a maternal contig). No large contigs have a high mis-phased SNVs ratio, which suggests proper phasing of the (a) Canu paternal, (b) Canu maternal, (c) FALCON paternal, and (d) FALCON maternal assemblies.
Supplementary Figure 8 Coverage titration for variant calling, phasing, and assembly.
Precision and recall for (a) SNVs and (b) indels called with DeepVariant (CCS), subsampling in steps of 3%. (c) Precision and recall for structural variants called with pbsv, subsampling in steps of 10%. (d) Phase block N50 for phasing of the 28-fold DeepVariant (CCS) callset with WhatsHap, subsampling in steps of 10%. Phasing performance is similar with a callset produced at matched coverage (not shown). De novo assembly (e) completeness measured as total assembly size, (f) contiguity measured as contig N50, and (g) correctness measured as concordance to the HG002 GIAB benchmark for wtdbg2 assembly, subsampling reads in steps of 10%.
Supplementary Figure 9 Likely errors in the GIAB benchmark identified by CCS callsets.
Manual curation of discrepancies between the GIAB benchmark and CCS variant callsets identifies benchmark errors for all variant types that are correctable using the CCS variant callsets. Shown are four loci that the GIAB benchmark records as homozygous reference where CCS reads identify likely heterozygous variation: (a) Three SNVs supported by CCS reads and 6 kb matepair reads. (b) A 2 bp insertion supported by CCS reads, 10X Genomics reads, and 6 kb matepair reads. (c) A 328 bp insertion supported by CCS reads and assemblies. (d) An 83 bp deletion supported by CCS reads.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9, Supplementary Tables 1–12 and Supplementary Note.
Rights and permissions
About this article
Cite this article
Wenger, A.M., Peluso, P., Rowell, W.J. et al. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat Biotechnol 37, 1155–1162 (2019). https://doi.org/10.1038/s41587-019-0217-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0217-9
This article is cited by
-
Full-length isoform concatenation sequencing to resolve cancer transcriptome complexity
BMC Genomics (2024)
-
Analysis of five near-complete genome assemblies of the tomato pathogen Cladosporium fulvum uncovers additional accessory chromosomes and structural variations induced by transposable elements effecting the loss of avirulence genes
BMC Biology (2024)
-
Characterization and analysis of multi-organ full-length transcriptomes in Sphaeropteris brunoniana and Alsophila latebrosa highlight secondary metabolism and chloroplast RNA editing pattern of tree ferns
BMC Plant Biology (2024)
-
DNA satellite and chromatin organization at mouse centromeres and pericentromeres
Genome Biology (2024)
-
Genome assembly of Melilotus officinalis provides a new reference genome for functional genomics
BMC Genomic Data (2024)