Abstract
Understanding complex tissues requires single-cell deconstruction of gene regulation with precision and scale. Here, we assess the performance of a massively parallel droplet-based method for mapping transposase-accessible chromatin in single cells using sequencing (scATAC-seq). We apply scATAC-seq to obtain chromatin profiles of more than 200,000 single cells in human blood and basal cell carcinoma. In blood, application of scATAC-seq enables marker-free identification of cell type-specific cis- and trans-regulatory elements, mapping of disease-associated enhancer activity and reconstruction of trajectories of cellular differentiation. In basal cell carcinoma, application of scATAC-seq reveals regulatory networks in malignant, stromal and immune cells in the tumor microenvironment. Analysis of scATAC-seq profiles from serial tumor biopsies before and after programmed cell death protein 1 blockade identifies chromatin regulators of therapy-responsive T cell subsets and reveals a shared regulatory program that governs intratumoral CD8+ T cell exhaustion and CD4+ T follicular helper cell development. We anticipate that scATAC-seq will enable the unbiased discovery of gene regulatory factors across diverse biological systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All single-cell sequencing data are available through the Gene Expression Omnibus under accession GSE129785. There are no restrictions on data availability or use. Species-mixing and PBMC datasets are available in pre- and post-processed formats here: https://support.10xgenomics.com/single-cell-atac/datasets. WashU browser sessions of aggregated scATAC-seq data (by cluster, as shown in each Figure) are available here: Fig. 2 single-cell clusters: http://epigenomegateway.wustl.edu/legacy/?genome=hg19&session=HcbHMSgBCc&statusId=28207718. Fig. 4 single-cell clusters: http://epigenomegateway.wustl.edu/legacy/?genome=hg19&session=tYJVrV7zzk&statusId=834543265. Fig. 5 single-cell clusters: http://epigenomegateway.wustl.edu/legacy/?genome=hg19&session=7UZG0iF90b&statusId=807471043. Whole exome sequencing data from patients SU006 and SU008 were previously described48 and obtained from the Sequence Read Archive under accession PRJNA533341.
Code availability
Custom code for main analyses used in this work has been deposited on GitHub: https://github.com/GreenleafLab/10x-scATAC-2019.
References
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Schep, A. N.et al. Structured nucleosome fingerprints enable high-resolution mapping of chromatin architecture within regulatory regions. Genome Res. 25, 1757–1770 (2015).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Cusanovich, D. A. et al. Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Cusanovich, D. A. et al. The cis-regulatory dynamics of embryonic development at single-cell resolution. Nature 555, 538–542 (2018).
Satpathy, A. T. et al. Transcript-indexed ATAC-seq for precision immune profiling. Nat. Med. 24, 580–590 (2018).
Buenrostro, J. D. et al. Integrated single-cell analysis maps the continuous regulatory landscape of human hematopoietic differentiation. Cell 173, 1535–1548.e16 (2018).
Cusanovich, D. A. et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell 174, 1309–1324.e18 (2018).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Neal, J. T. et al. Organoid modeling of the tumor immune microenvironment. Cell 175, 1972–1988.e16 (2018).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361–376.e17 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Pliner, H. A. et al. Cicero predicts cis-regulatory dna interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871.e8 (2018).
Grajales-Reyes, G. E. et al. Batf3 maintains autoactivation of Irf8 for commitment of a CD8α(+) conventional DC clonogenic progenitor. Nat. Immunol. 16, 708–717 (2015).
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Mumbach, M. R. et al. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 49, 1602–1612 (2017).
GTEx Consortium et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Crispino, J. D. & Weiss, M. J. Erythro-megakaryocytic transcription factors associated with hereditary anemia. Blood 123, 3080–3088 (2014).
Nutt, S. L. & Kee, B. L. The transcriptional regulation of B cell lineage commitment. Immunity 26, 715–725 (2007).
Johnson, J. L. et al. Lineage-determining transcription factor TCF-1 initiates the epigenetic identity of T cells. Immunity 48, 243–257.e10 (2018).
Gerber, E. E. et al. Integrin-modulating therapy prevents fibrosis and autoimmunity in mouse models of scleroderma. Nature 503, 126–130 (2013).
Castellanos, J. G. et al. Microbiota-Induced TNF-like ligand 1A drives group 3 innate lymphoid cell-mediated barrier protection and intestinal T cell activation during colitis. Immunity 49, 1077–1089.e5 (2018).
Niebuhr, B. et al. Runx1 is essential at two stages of early murine B-cell development. Blood 122, 413–423 (2013).
Stehling-Sun, S., Dade, J., Nutt, S. L., DeKoter, R. P. & Camargo, F. D. Regulation of lymphoid versus myeloid fate ‘choice’ by the transcription factor Mef2c. Nat. Immunol. 10, 289–296 (2009).
Kerenyi, M. A. & Orkin, S. H. Networking erythropoiesis. J. Exp. Med. 207, 2537–2541 (2010).
Fulkerson, P. C. Transcription factors in eosinophil development and as therapeutic targets. Front. Med. 4, 115 (2017).
Satpathy, A. T., Wu, X., Albring, J. C. & Murphy, K. M. Re(de)fining the dendritic cell lineage. Nat. Immunol. 13, 1145–1154 (2012).
Manz, M. G., Traver, D., Miyamoto, T., Weissman, I. L. & Akashi, K. Dendritic cell potentials of early lymphoid and myeloid progenitors. Blood 97, 3333–3341 (2001).
Becker, A. M. et al. IRF-8 extinguishes neutrophil production and promotes dendritic cell lineage commitment in both myeloid and lymphoid mouse progenitors. Blood 119, 2003–2012 (2012).
Rodrigues, P. F. et al. Distinct progenitor lineages contribute to the heterogeneity of plasmacytoid dendritic cells. Nat. Immunol. 19, 711–722 (2018).
Chang, A. L. S. et al. Pembrolizumab for advanced basal cell carcinoma: an investigator-initiated, proof-of-concept study. J. Am. Acad. Dermatol. 80, 564–566 (2019).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624.e24 (2017).
Atwood, S. X., Li, M., Lee, A., Tang, J. Y. & Oro, A. E. GLI activation by atypical protein kinase C ι/λ regulates the growth of basal cell carcinomas. Nature 494, 484–488 (2013).
Jaiswal, S. et al. CD47 is upregulated on circulating hematopoietic stem cells and leukemia cells to avoid phagocytosis. Cell 138, 271–285 (2009).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Okazaki, T. & Honjo, T. The PD-1-PD-L pathway in immunological tolerance. Trends Immunol. 27, 195–201 (2006).
Betancur, P. A. et al. A CD47-associated super-enhancer links pro-inflammatory signalling to CD47 upregulation in breast cancer. Nat. Commun. 8, 14802 (2017).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, pii: eaav1898 (2018).
Li, H. et al. Dysfunctional CD8 T cells form a proliferative, dynamically regulated compartment within human melanoma. Cell 176, 775–789.e18 (2018).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Preprint at bioRxiv https://doi.org/10.1101/648899 (2019).
Pauken, K. E. et al. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science 354, 1160–1165 (2016).
Sen, D. R. et al. The epigenetic landscape of T cell exhaustion. Science 354, 1165–1169 (2016).
Scott-Browne, J. P. et al. Dynamic changes in chromatin accessibility occur in CD8+ T cells responding to viral infection. Immunity 45, 1327–1340 (2016).
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Pearce, E. L. et al. Control of effector CD8+ T cell function by the transcription factor eomesodermin. Science 302, 1041–1043 (2003).
Sullivan, B. M., Juedes, A., Szabo, S. J., von Herrath, M. & Glimcher, L. H. Antigen-driven effector CD8 T cell function regulated by T-bet. Proc. Natl Acad. Sci. USA 100, 15818–15823 (2003).
Intlekofer, A. M. et al. Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin. Nat. Immunol. 6, 1236–1244 (2005).
Chou, C. et al. c-Myc-induced transcription factor AP4 is required for host protection mediated by CD8+ T cells. Nat. Immunol. 15, 884–893 (2014).
Yu, B. et al. Epigenetic landscapes reveal transcription factors that regulate CD8+ T cell differentiation. Nat. Immunol. 18, 573–582 (2017).
Phan, A. T. et al. Constitutive glycolytic metabolism supports CD8+ T cell effector memory differentiation during viral infection. Immunity 45, 1024–1037 (2016).
Mackay, L. K. et al. The developmental pathway for CD103(+)CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Au-Yeung, B. B. et al. A sharp T-cell antigen receptor signaling threshold for T-cell proliferation. Proc. Natl Acad. Sci. USA 111, E3679–E3688 (2014).
Chen, J. et al. NR4A transcription factors limit CAR T cell function in solid tumours. Nature 567, 530–534 (2019).
Milner, J. J. et al. Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours. Nature 552, 253–257 (2017).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Quigley, M. et al. Transcriptional analysis of HIV-specific CD8+ T cells shows that PD-1 inhibits T cell function by upregulating BATF. Nat. Med. 16, 1147–1151 (2010).
Martinez, G. J. et al. The transcription factor NFAT promotes exhaustion of activated CD8+ T cells. Immunity 42, 265–278 (2015).
Man, K. et al. Transcription factor IRF4 promotes CD8+ T cell exhaustion and limits the development of memory-like T cells during chronic infection. Immunity 47, 1129–1141.e5 (2017).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Gu-Trantien, C. et al. CD4+ follicular helper T cell infiltration predicts breast cancer survival. J. Clin. Invest. 123, 2873–2892 (2013).
Zappasodi, R. et al. Non-conventional inhibitory CD4+ Foxp3-PD-1hi T cells as a biomarker of immune checkpoint blockade activity. Cancer Cell 33, 1017–1032.e7 (2018).
Locci, M. et al. Activin a programs the differentiation of human TFH cells. Nat. Immunol. 17, 976–984 (2016).
Baumjohann, D. et al. Persistent antigen and germinal center B cells sustain T follicular helper cell responses and phenotype. Immunity 38, 596–605 (2013).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Crawford, A. et al. Molecular and transcriptional basis of CD4+ T cell dysfunction during chronic infection. Immunity 40, 289–302 (2014).
Elsaesser, H., Sauer, K. & Brooks, D. G. IL-21 is required to control chronic viral infection. Science 324, 1569–1572 (2009).
Yi, J. S., Du, M. & Zajac, A. J. A vital role for interleukin-21 in the control of a chronic viral infection. Science 324, 1572–1576 (2009).
Fröhlich, A. et al. IL-21R on T cells is critical for sustained functionality and control of chronic viral infection. Science 324, 1576–1580 (2009).
Denny, S. K. et al. Nfib promotes metastasis through a widespread increase in chromatin accessibility. Cell 166, 328–342 (2016).
Yang, D. et al. Intertumoral heterogeneity in SCLC is influenced by the cell type of origin. Cancer Discov. 8, 1316–1331 (2018).
Ay, F., Bailey, T. L. & Noble, W. S. Statistical confidence estimation for Hi-C data reveals regulatory chromatin contacts. Genome Res. 24, 999–1011 (2014).
Acknowledgements
We thank members of the Chang and Greenleaf laboratories and 10x Genomics for helpful discussions. We thank the following people at 10x Genomics: A. Puleo for sorting cells, J. Chevillet for training, Z. Bent and M. Dodge for reagents development, R. Gerver and W. Wang for microfluidics and A. Gallegos, A. Gonzales, N. Keivanfar, S. Maheshwari, P. Marks, J. Mellen, R. Rico and K. Wu for computational and software support. We thank X. Ji, D. Wagh and J. Coller at the Stanford Functional Genomics Facility and C. Bruce at 10x Genomics for sequencing support, and A. Valencia for assistance with clinical specimen processing. This work was supported by the National Institutes of Health grant nos. P50HG007735 (H.Y.C. and W.J.G.), K08CA230188 (A.T.S.), K99-AG059918 (M.R.C.), UM1HG009442 (H.Y.C. and W.J.G.) and S10OD018220 (Stanford Functional Genomics Facility), the Parker Institute for Cancer Immunotherapy (A.T.S. and H.Y.C.), the Michelson Foundation (A.T.S.) and the Scleroderma Research Foundation (H.Y.C.). A.T.S. was supported by a Bridge Scholar Award from the Parker Institute for Cancer Immunotherapy, a Career Award for Medical Scientists from the Burroughs Wellcome Fund and the Human Vaccines Project Michelson Prize for Human Immunology and Vaccine Research. K.E.Y. was supported by the National Science Foundation Graduate Research Fellowship Program (NSF DGE-1656518) and a Stanford Graduate Fellowship. W.J.G. is a Chan Zuckerberg Biohub investigator and acknowledges grant nos. 2017–174468 and 2018–182817 from the Chan Zuckerberg Initiative. H.Y.C. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.T.S., J.M.G., G.X.Y.Z., W.J.G. and H.Y.C. conceived the project. A.T.S., J.M.G., Y.Q., K.E.Y., M.R.C., M.R.M., S.E.P., F.M., G.P.M., J.C.B., D.J., C.M.N., J.W. and Y.Y. performed experiments. J.M.G. led the analysis of scATAC-seq data. B.N.O., P.S. and L.W. contributed to the Cell Ranger ATAC software and contributed to data analysis with P.G.G. A.L.S.C. obtained clinical specimens. A.T.S., H.Y.C., W.J.G. and G.X.Y.Z. guided experiments and data analysis. A.T.S., J.M.G., G.X.Y.Z., W.J.G. and H.Y.C. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
H.Y.C. is a cofounder of Accent Therapeutics and Epinomics and is an adviser to 10x Genomics and Spring Discovery. W.J.G. is a cofounder of Epinomics and an adviser to 10x Genomics, Guardant Health and Centrillion. A.T.S. is an advisor to Immunai. F.M., G.P.M., B.N.O., P.S., J.C.B., D.J., C.M.N., J.W., L.W., Y.Y., P.G.G. and G.Y.Z. are employees of 10x Genomics. A.L.S.C. was an advisory board member and clinical investigator for studies sponsored by Merck, Regeneron, Novartis, Galderma and Genentech Roche. Stanford University holds patents on ATAC-seq, on which P.G., W.J.G. and H.Y.C. are named as inventors.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
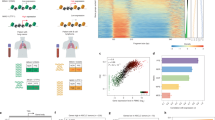
Supplementary Figure 1 Droplet-based scATAC-seq workflow and quality control measurements.
(a) Protocol steps for scATAC-seq in droplets. (b) Genome tracks showing the comparison of aggregate scATAC-seq profiles from A20 B lymphocytes (top panel). scATAC-seq profiles were obtained from four independent experiments, as indicated. The bottom panel shows accessibility profiles of 100 random single A20 cells from two cell mixing experiments. Each pixel represents a 100bp region. (c) Left plot: Pearson correlation heatmaps of log-normalized reads in bulk GM12878 Omni-ATAC-seq peaks in aggregate scATAC-seq profiles generated from varying numbers of single cells, or from published Omni-ATAC profiles5 (n=100,000 ATAC-seq peaks). Right plot: Pearson correlation heatmaps of log-normalized reads in aggregate scATAC-seq profiles from A20 cells (n=100,000 ATAC-seq peaks, identified in an aggregate profile from all cells). Numbers in parentheses indicate the cell loading concentration. (d) Peak recovery analysis with subsampled cells and unique fragments as determined by x-axis and colors, respectively. scATAC-seq cells were subsampled to the indicated unique fragments, and the proportion of peaks recovered from the aggregate profile was calculated as a function of number of cells analyzed. GM12878 cells generated a median of 29,451 unique nuclear fragments per cell (top level of down-sampling was 25,000) while A20 cells generated a median of 20,809 unique nuclear fragments per cell (top level of down-sampling was 20,000). The center line represents the Loess fit, and shaded regions indicate 95% confidence interval (n=16 sub-sampled profiles at each read depth). (e) Pearson correlation analysis with subsampled cells and unique fragments as determined by x-axis and colors, respectively. scATAC-seq profiles were subsampled to the indicated unique fragments, and Pearson correlation to the aggregate profile was calculated as a function of number of cells analyzed (n=16 sub-sampled profiles at each read depth). (f) Analysis workflow for scATAC-seq data in this study.
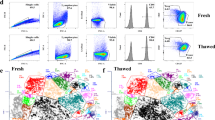
Supplementary Figure 2 scATAC-seq performance in frozen cells and synthetic cell mixtures.
(a) Synthetic immune cell mixture quality control experiments. Sorted human monocytes or T cells were mixed at the indicated ratio and analyzed with scATAC-seq. Plots show the UMAP of scATAC-seq profiles (top), and gene scores for monocyte- or T cell-associated cardinal genes (see Methods) in each single cell (middle and bottom). Dashed circles indicate monocyte and T cell identity of single cells as determined by ATAC-seq profiles. Colors indicate cluster identity defined de novo. (b) Sorted human monocytes or T cells were mixed at the indicated ratios and analyzed with scATAC-seq and analyzed as described in (a). (c) Comparison of data quality from fresh and frozen PBMCs, and frozen PBMCs sorted for live cells. Representative ATAC-seq data quality control filters by sample source. Shown are the number of unique ATAC-seq nuclear fragments in each single cell (each dot) compared to TSS enrichment of all fragments in that cell. Dashed lines represent the filters for high-quality single-cell data (1,000 unique nuclear fragments and TSS score greater than or equal to 8). (d) One-to-one plots of log-normalized reads in aggregated scATAC-seq in profiles generated from the indicated cell source (fresh, frozen, or frozen sorted PBMCs). Peaks were defined in fresh samples. Numbers indicate Pearson correlation value. (e) ROC (top) and Precision-vs-Recall (bottom) curves showing recovery of fresh PBMC peaks with frozen or frozen sorted cells. True positive peaks were defined as those identified in fresh PBMC scATAC-seq profiles. (f) Integrated UMAP of all scATAC-seq profiles from monocyte/T cell mixing experiments in (a-b). This indicates that strong clustering batch effects are not seen between experiments. (g) UMAP and PCA analysis of of scATAC-seq profiles (left) and clusters (right) identified in fresh, frozen, or frozen-sorted PBMCs using fresh PBMC peaks.
Supplementary Figure 3 Sample descriptions and quality control of scATAC-seq hematopoiesis data.
(a) UMAP projection of 63,882 scATAC-seq profiles of bone marrow and peripheral blood immune cell types. Dots represent individual cells, and colors indicate the experimental source of each cluster, as labeled on the right of the plot (see Methods). (b) Bar plots indicate the number of scATAC-seq profiles obtained from each experimental source of cells (left), and the median number of unique nuclear fragments in single cells (right). (c) UMAP projection of 63,882 scATAC-seq profiles of bone marrow and peripheral blood immune cell types. Colors represent the log10 number of unique nuclear fragments per single cell. (d) Representative scATAC-seq data quality control filters by sample source. Shown are the number of unique ATAC-seq nuclear fragments in each single cell (each dot) compared to TSS enrichment of all fragments in that cell. Dashed lines represent the filters for high-quality single-cell data (1,000 unique nuclear fragments and TSS score greater than or equal to 8). (e) Single-cell ATAC-seq data quality control filters in profiles generated using the C1 microfluidic system11 (Fluidigm; left) or sci-ATAC-seq12 (middle and right panels). (f) Peak recovery analysis with subsampled cells and unique fragments as determined by x-axis and colors, respectively. scATAC-seq cells were subsampled to the indicated unique fragments, and the proportion of peaks recovered from the aggregate profile was calculated as a function of number of cells analyzed. The center line represents the Loess fit, and shaded regions indicate 95% confidence interval (n=10 sub-sampled profiles at each read depth).
Supplementary Figure 4 Cis-regulatory elements in hematopoiesis and co-accessibility validation.
(a) Genome tracks of aggregate scATAC-seq data, clustered as indicated in Figure 2b. Arrows indicate the position and distance (in kb) of intragenic or distal enhancers in each gene locus. (b) MetaV4C plot of H3K27ac HiChIP data demonstrating HiChIP signal at Cicero-identified co-accessible cis-elements (linked to promoter elements). Each plot shows the aggregate HiChIP signal (from 3 T cell types, n=2 biologically independent HiChIP profiles per cell type) between linked cis-elements identified in scATAC-seq data. Each link is scaled so that the 0% position indicates the promoter site and the 100% position indicates the linked cis-element site. The peak indicates an enrichment of HiChIP signal at the linked peaks compared to surrounding genomic regions. Biased links are identified by differential peak analysis in the indicated scATAC-seq clusters. The center line represents the Loess fit, and shaded regions indicate 95% confidence interval. (c) Support for Cicero-identified co-accessible cis-elements by GTEX eQTL data. Shown is the mean enrichment of eQTL signal (determined in the indicated tissue type, bars indicate standard deviation from n=250 simulations) in co-accessible sites linked to promoter elements described in GTEX vs 250 permutations of ATAC-seq peak to genes. Greater enrichment is observed in immune tissues because scATAC-seq data profiled the relevant cell types. (d) Heatmap of log-normalized gene scores for the indicated genes. (e) UMAP projection colored by log normalized gene scores demonstrating the accessibility of cis-regulatory elements linked (using Cicero) to the indicated gene.
Supplementary Figure 5 TF motif accessibility in hematopoiesis and cell type-specific GWAS enrichment.
(a) Example TF footprints and motifs in the indicated scATAC-seq clusters identified in Fig. 2b. The Tn5 insertion bias track is shown below. (b) UMAP projection of scATAC-seq profiles colored by chromVAR TF motif bias-corrected deviations for the indicated factors. (c) Analysis workflow for GWAS enrichment scores using Cicero co-accessibility. (d) Heatmap showing GWAS deviation scores for PICS SNPs associated with the indicated diseases. PICS SNPs were identified previously21. (e) Example of increased ATAC-seq signal in a GWAS-containing cis-element in NK and T cell scATAC-seq clusters. The HiChIP plot (top) demonstrates increased H3K27ac HiChIP signal between the STAT4 promoter and the highlighted cis-elements. The center line represents the Loess fit, and shaded regions indicate 95% confidence interval (n=2 biologically independent HiChIP profiles per cell type).
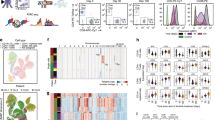
Supplementary Figure 6 Sample descriptions and quality control of scATAC-seq profiles of the BCC TME.
(a) UMAP projection of 37,818 scATAC-seq profiles of BCC TME cell types. Dots represent individual cells, and colors indicate the experimental source of each cluster, as labeled on the right of the plot (see Methods). ‘Total’ samples were sorted as all live cells in a single BCC biopsy. ‘T cell’ samples were sorted as CD45+CD3+ cells in the biopsy. ‘Immune’ samples were sorted as CD45+CD3- cells in the biopsy. ‘Stromal’ samples were sorted as CD45-CD3- cells in the biopsy. (b) Bar plots indicate the number of scATAC-seq profiles obtained from each experimental source of cells (left), and the median number of unique nuclear fragments in single cells (right). (c) UMAP projection of 37,818 scATAC-seq TME profiles. Colors represent the log10 number of unique nuclear reads per single cell. (d) Representative ATAC-seq data quality control filters by sample source. Shown are the number of unique ATAC-seq nuclear fragments in each single cell (each dot) compared to TSS enrichment of all fragments in that cell. Dashed lines represent the filters for high-quality single-cell data (1,000 unique nuclear fragments and TSS score greater than or equal to 8). (e) Genome tracks of aggregate scATAC-seq data, clustered as indicated in Figure 4b. Arrows indicate the position and distance (in kb) of intragenic or distal enhancers in each gene locus. (f) Bar plots indicating the relative proportion of cells from each patient detected in each cluster (top) and the relative proportion of cells from each cluster detected in each patient (bottom).
Supplementary Figure 7 TF motif accessibility in the BCC TME.
(a) Example TF footprints and motifs in the indicated scATAC-seq clusters identified in Fig. 4b. The Tn5 insertion bias track is shown below. (e) UMAP projection of scATAC-seq profiles colored by chromVAR TF motif bias-corrected deviations for the indicated factors.
Supplementary Figure 8 Regulatory landscapes of tumor-infiltrating T cell subsets.
(a) UMAP projection of intratumoral T cell scATAC-seq data colored by log normalized gene scores, demonstrating the accessibility of cis-regulatory elements linked (using Cicero) to the indicated CD8+ T cell signature genes. (b) UMAP projection of intratumoral T cell scATAC-seq data colored by log normalized gene scores, demonstrating the accessibility of cis-regulatory elements linked (using Cicero) to the indicated CD4+ T cell signature genes. (c) Genome tracks of Tfh signature genes in aggregate scATAC-seq data, clustered as indicated in Figure 5a. Arrows indicate the position and distance (in kb) of intragenic or distal enhancers in each gene locus. (d) Heatmap of Z-scores of 35,147 cis-regulatory elements in scATAC-seq clusters derived from (b). Labels indicate cell type-specific accessibility of regulatory elements. (e) Genome tracks of CD8+ TEx signature genes in aggregate scATAC-seq data, demonstrating the overlap of TEx and Tfh regulatory elements. Arrows indicate the position and distance (in kb) of intragenic or distal enhancers in each gene locus. Selected TF binding motifs present in the +5kb enhancer of PDCD1 are shown (bottom). Lines indicate the binding motif location. (f) Heatmap representation of ATAC-seq chromVAR bias-corrected deviations in the 250 most variable TFs across all intratumoral T cell scATAC-seq clusters, as identified in Figure 5a. Cluster identities are indicated at the bottom of the plot.
Supplementary Figure 9 T cell cluster dynamics pre- and post-therapy according to clinical outcome.
(a) Shown is the pre- and post-therapy frequency (left) and cell number (right) for each T cell cluster identified in Figure 5. Each plot is generated from all responder patient samples aggregated together. (b) Pre- and post-therapy frequency (left) and cell number (right) for each T cell cluster in non-responder patients.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9
Supplementary Table 1
List of single-cell barcodes. Sequences of 16-bp barcodes to index single cells in GEMs.
Supplementary Table 2
Quality control sample characteristics. Sources and single-cell numbers for each sample analyzed in QC experiments. Related to Fig. 1 and Supplementary Figs. 1–2.
Supplementary Table 3
Cell frequencies in monocyte/T cell mixing experiments. Expected and observed ratios of monocytes and T cells in cell-mixing experiments. Related to Fig. 1 and Supplementary Figs. 1–2.
Supplementary Table 4
Hematopoiesis sample characteristics. The source, isolation method and single-cell numbers for each sample analyzed by scATAC-seq in the study of immune cell development. Related to Figs. 2–3 and Supplementary Figs. 3–5.
Supplementary Table 5
BCC sample characteristics. Patient and cell characteristics for all BCC scATAC-seq samples. Response to anti-PD-1 therapy was categorized using RECIST scoring, as described in the Methods. Related to Figs. 4–5 and Supplementary Figs. 6–8.
Rights and permissions
About this article
Cite this article
Satpathy, A.T., Granja, J.M., Yost, K.E. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat Biotechnol 37, 925–936 (2019). https://doi.org/10.1038/s41587-019-0206-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0206-z
This article is cited by
-
Histone lactylation couples cellular metabolism with developmental gene regulatory networks
Nature Communications (2024)
-
Single-cell sequencing technology applied to epigenetics for the study of tumor heterogeneity
Clinical Epigenetics (2023)
-
Single-nucleus genomics in outbred rats with divergent cocaine addiction-like behaviors reveals changes in amygdala GABAergic inhibition
Nature Neuroscience (2023)
-
CD4+ T cells in cancer
Nature Cancer (2023)
-
A fast chemical reprogramming system promotes cell identity transition through a diapause-like state
Nature Cell Biology (2023)