Abstract
Recent technical advancements have facilitated the mapping of epigenomes at single-cell resolution; however, the throughput and quality of these methods have limited their widespread adoption. Here we describe a high-quality (105 nuclear fragments per cell) droplet-microfluidics-based method for single-cell profiling of chromatin accessibility. We use this approach, named ‘droplet single-cell assay for transposase-accessible chromatin using sequencing’ (dscATAC-seq), to assay 46,653 cells for the unbiased discovery of cell types and regulatory elements in adult mouse brain. We further increase the throughput of this platform by combining it with combinatorial indexing (dsciATAC-seq), enabling single-cell studies at a massive scale. We demonstrate the utility of this approach by measuring chromatin accessibility across 136,463 resting and stimulated human bone marrow-derived cells to reveal changes in the cis- and trans-regulatory landscape across cell types and under stimulatory conditions at single-cell resolution. Altogether, we describe a total of 510,123 single-cell profiles, demonstrating the scalability and flexibility of this droplet-based platform.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw sequencing files and processed files for all data generated in this study were deposited at Gene Expression Omnibus (GEO) under accession number GSE123581. UCSC genome browser tracks for the datasets generated in this study are available from the following websites: mouse brain, https://s3.us-east-2.amazonaws.com/jasonbuenrostro/2018_mouse_brain/hub.txt; BMMC dsciATAC-seq, https://s3.us-east-2.amazonaws.com/jasonbuenrostro/2018_BM_htsci/hub.txt; stimulated BMMC dsciATAC-seq, https://s3.us-east-2.amazonaws.com/jasonbuenrostro/2018_BM_htsci_stim/hub.txt.
Code availability
Complete code and documentation for the BAP software suite developed in this study is available at https://github.com/buenrostrolab/bap. Scripts corresponding to the analyses contained in this paper are provided at https://github.com/buenrostrolab/dscATAC_analysis_code.
References
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Spitz, F. & Furlong, E. E. M. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why? Mol. Cell 49, 825–837 (2013).
Weintraub, H. & Groudine, M. Chromosomal subunits in active genes have an altered conformation. Science 193, 848–856 (1976).
Boyle, A. P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Hesselberth, J. R. et al. Global mapping of protein–DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Plasschaert, L. W. et al. A single-cell atlas of the airway epithelium reveals the CFTR-rich pulmonary ionocyte. Nature 560, 377–381 (2018).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Wagner, A., Regev, A. & Yosef, N. Revealing the vectors of cellular identity with single-cell genomics. Nat. Biotechnol. 34, 1145–1160 (2016).
Kelsey, G., Stegle, O. & Reik, W. Single-cell epigenomics: recording the past and predicting the future. Science 358, 69–75 (2017).
Jin, W. et al. Genome-wide detection of DNase I hypersensitive sites in single cells and FFPE tissue samples. Nature 528, 142–146 (2015).
Rotem, A. et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nat. Biotechnol. 33, 1165–1172 (2015).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Cusanovich, D. A. et al. Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Cusanovich, D. A. et al. The cis-regulatory dynamics of embryonic development at single-cell resolution. Nature 555, 538–542 (2018).
Buenrostro, J. D. et al. Integrated single-cell analysis maps the continuous regulatory landscape of human hematopoietic differentiation. Cell 173, 1535–1548 (2018).
Corces, M. R. et al. An improved ATAC–seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Amini, S. et al. Haplotype-resolved whole-genome sequencing by contiguity-preserving transposition and combinatorial indexing. Nat. Genet. 46, 1343–1349 (2014).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat. Neurosci. 21, 432–439 (2018).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871 (2018).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. Chromvar: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Cusanovich, D. A. et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell 174, 1309–1324 (2018).
Chen, X., Miragaia, R. J., Natarajan, K. N. & Teichmann, S. A. A rapid and robust method for single cell chromatin accessibility profiling. Nat. Commun. 9, 5345 (2018).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 (2018).
Urban-Ciecko, J. & Barth, A. L. Somatostatin-expressing neurons in cortical networks. Nat. Rev. Neurosci. 17, 401–409 (2016).
Ullrich, B. & Südhof, T. C. Differential distributions of novel synaptotagmins: comparison to synapsins. Neuropharmacology 34, 1371–1377 (1995).
Meneses, A. Serotonin, neural markers, and memory. Front. Pharmacol. 6, 143 (2015).
Hatori, M. et al. Lhx1 maintains synchrony among circadian oscillator neurons of the SCN. eLife 3, e03357 (2014).
Visel, A. et al. A high-resolution enhancer atlas of the developing telencephalon. Cell 152, 895–908 (2013).
Kadkhodaei, B. et al. Nurr1 is required for maintenance of maturing and adult midbrain dopamine neurons. J. Neurosci. 29, 15923–15932 (2009).
Yap, E.-L. & Greenberg, M. E. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron 100, 330–348 (2018).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Bodenmiller, B. et al. Multiplexed mass cytometry profiling of cellular states perturbed by small-molecule regulators. Nat. Biotechnol. 30, 858–867 (2012).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. (2018). https://doi.org/10.1038/nbt.4314.
Essers, M. A. G. et al. IFNα activates dormant haematopoietic stem cells in vivo. Nature 458, 904–908 (2009).
Espín-Palazón, R. et al. Proinflammatory signaling regulates hematopoietic stem cell emergence. Cell 159, 1070–1085 (2014).
Petrillo, C. et al. Cyclosporine H overcomes innate immune restrictions to improve lentiviral transduction and gene editing in human hematopoietic stem cells. Cell Stem Cell 23, 820–832 (2018).
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Ulirsch, J.C. et al. Interrogation of human hematopoiesis at single-cell and single-variant resolution. Nat. Genet. 51, 683–693 (2019).
Lopez, R. D., Waller, E. K., Lu, P. H. & Negrin, R. S. CD58/LFA-3 and IL-12 provided by activated monocytes are critical in the in vitro expansion of CD56+ T cells. Cancer Immunol. Immunother. 49, 629–640 (2001).
Laroni, A. et al. Dysregulation of regulatory CD56bright NK cells/T cells interactions in multiple sclerosis. J. Autoimmun. 72, 8–18 (2016).
HCA Consortium. The Human Cell Atlas White Paper (2017).
Stuart, T. et al. Comprehensive integration of single cell data. Preprint at https://doi.org/10.1101/460147 (2018).
Han, X. et al. Mapping the mouse cell atlas by microwell-seq. Cell 173, 13071091– (2018).
Vitak, S. A. et al. Sequencing thousands of single-cell genomes with combinatorial indexing. Nat. Methods 14, 302–308 (2017).
Mulqueen, R. M. et al. Highly scalable generation of DNA methylation profiles in single cells. Nat. Biotechnol. 36, 428–431 (2018).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Rakyan, V. K., Down, T. A., Balding, D. J. & Beck, S. Epigenome-wide association studies for common human diseases. Nat. Rev. Genet. 12, 529–541 (2011).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Lander, E. S. & Waterman, M. S. Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics 2, 231–239 (1988).
Acknowledgements
We thank members of the Buenrostro lab for useful discussions and critical assessment of this work. We thank D. Norton at Bio-Rad for enabling the collaboration with the Broad Institute and Harvard University. We would also like to acknowledge other Bio-Rad colleagues for establishing and providing droplet-related consumables, including D. Do, B. Zhang, P. Pattamatta, S. Cater, L. Frenz, D. Greiner and J. Agresti. We also recognize L. Christiansen, A. Yunghans and L. Watson from the Illumina Assay Development team, in addition to F. Zhang and F. Schlesinger at Illumina, for their bioinformatics contributions. We are grateful to the Zhang lab (Broad Institute) for providing the Tn5 for combinatorial experiments. J.D.B., C.A.L., F.M.D. and V.K.K. acknowledge support by the Allen Distinguished Investigator Program through the Paul G. Allen Frontiers Group. This work was further supported by the Chan Zuckerberg Initiative. C.A.L. is supported by an NIH F31 grant (F31CA232670).
Author information
Authors and Affiliations
Contributions
F.M.D., J.G.C. and A.S.K. generated the data. C.A.L., V.K.K. and Z.D.B. analyzed the data. F.J.S. proposed the droplet scATAC-seq approach and oversaw the proof-of-concept studies performed by D.P. M.J.A. assisted in the development of computational resources. C.A.L., F.M.D. and J.D.B. wrote the manuscript with input from all authors. R.L. and J.D.B. jointly supervised this work.
Corresponding authors
Ethics declarations
Competing interests
Work by D.P. and F.J.S. was performed at Illumina. Work by J.G.C., Z.D.B., A.S.K. and R.L. was performed at Bio-Rad. J.D.B. holds patents related to ATAC-seq.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Optimization of Tn5 transposition for dscATAC-seq.
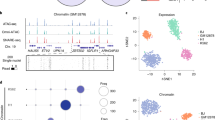
(a) Fraction of reads mapping to the nuclear genome for each of the Tn5 concentrations. The remaining reads map to the mitochondrial genome. Different volumes (2.5–10 μL) of the standard commercial Tn5 (TDE1) are compared against 3 replicates of a custom Tn5 concentration (2.5 μL) optimized for dscATAC-seq for K562 cells. (b) Number of unique reads mapping near transcription start sites (TSS) or (c) distal regulatory elements for the same Tn5 conditions. Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range. All three panels (a-c) show the top 500 cells sorted by library size. (d) Schematic of biochemical process leading to multiple fragments becoming tagged by multiple bead barcodes in the same droplet.
Supplementary Figure 2 Validation of bead merging computational approach.
(a) Browser shot of paired-end reads near the DIAPH1 and GAPDH loci. Reads are colored by bead barcode sequence. (b) Schematic of verification experiment where a library of random oligonucleotides was encapsulated into droplets together with Tn5 transposed cells and barcoded beads. The schematic shows a droplet containing a library of random oligos, a cell and two beads with different barcode sequences. (c) The expected number of beads per drop as a function of bead concentration. Inference of this line was determined by a maximum likelihood estimation for a double-truncated Poisson distribution. (d) Percent of drops with one or more beads as a function of bead concentration. Values are estimated using the probability density function of a Poisson distribution parameterized by the mean number of beads per drop from (c). (e) Jaccard index overlap metric for pairs of bead barcodes loaded at a concentration of 200 beads/μL. For each pair of bead barcodes observed, the Jaccard index was computed over the observed random oligonucleotide sequences. (f) The BAP overlap score computed from the dscATAC-seq data (agnostic to oligonucleotides) from the same experiment. In each panel, pairs of bead barcodes nominated for merging are highlighted in blue. Merged pairs were determined by computing a “knee” inflection point. The same two panels are shown in (g–j) but for increased bead concentration: (g, h) 800 beads/μL; (i,j) 5,000 beads/μL. (k) (left panel) Area under the receiver operating curve (AUROC) values for true positive bead merges nominated from the random oligonucleotide sequences. Four metrics are compared, including Pearson and Spearman correlation and the Jaccard index of reads in peaks per pair of bead barcodes. The final metric is our novel computational approach, termed BAP. Various bead concentrations per experimental condition are shown below the x-axis. (right panel) The same conditions and metrics but showing the area under the precision-recall curve (AUPRC). (l) %TSS enrichment scores for the same pool of cells processed at different bead concentrations. Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range. (m) Per-cell library complexities across a range of tested bead concentrations, the same as in panel (l). Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range. Both panels (l, m) show the top 500 cells sorted by library size. (n) Species mixing plots and collision rates (text) for the same experiment (800 beads/μL) with and without bead merging. (o) The same plots as in (n) but at a bead concentration of 5,000 beads/μL.
Supplementary Figure 3 Additional quality controls of dscATAC-seq.
(a) Species mixing plots and estimated collision rates for existing scATAC-seq methods. (b) Fraction of reads in peaks for the comparison in Fig. 1f. The chromatin accessibility peak set was obtained from ENCODE DNase-seq data for GM12878 and thus agnostic to the datasets compared here. Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range. (c) Number of cells (GM12878 only) compared in panel (b) and Fig. 1f. (d) Rank sorted variability across transcription factor motifs within the GM12878 dscATAC-seq profiles.
Supplementary Figure 4 Quality control information for the dscATAC-seq mouse brain dataset and comparison with existing data.
(a) Distribution of number of beads per cell identified across the two mice (bead input concentration = 5,000 beads/μL) for high-quality cells that pass quality controls. The corresponding bead merging curves are shown to the right for the twelve libraries. (b) Mouse brain cells in the t-SNE from Fig. 2a colored by number of bead barcodes detected per cell. The same coordinates are shown for (c) mouse donor, and (d) experimental well. (e) De novo embedding using latent semantic indexing (LSI). Colors match annotations from Fig. 2a. All plots show the same (n = 46,653) cells shown in Fig. 2a. (f) t-SNE of previously published sciATAC-seq data for mouse brain (Cusanovich et al., Cell 174(5), 1309–1324.e18, 2018) using the same 7-mer method (Louvain, t-SNE; compare to Fig. 2a; n = 5,744 cells). (g) Comparison of the percentage of reads mapping to the nuclear genome (separated into TSS-proximal or distal chromatin accessibility peaks) between whole mouse brain data generated using dscATAC-seq or a recently optimized sciATAC-seq method (Cusanovich et al., Cell 174(5), 1309–1324.e18, 2018). Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range. (h) Raw total number of reads mapping to distal chromatin accessibility peaks (see blue from panel (g) between dscATAC-seq and the sciATAC-seq method described in (g)). Boxplots summarize thousands of cells for each comparison. Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range.
Supplementary Figure 5 Chromatin accessibility scores for validation of cell clusters from mouse brain.
(a) Schematic demonstrating the approach used to define chromatin accessibility scores surrounding gene promoters. (b) t-SNE of cells by promoter region chromatin accessibility scores for all genes. The same colors and cells (n = 46,653) used in Fig. 2a are shown here. (c) Hierarchical clustering of chromatin accessibility scores calculated as shown in (a) for each cluster derived from the mouse brain dscATAC-seq dataset using Pearson correlation. 27 clusters from Fig. 2a are depicted. (d) Representative chromatin accessibility scores for known marker genes defining cell types in the mouse brain, plots are titled by the marker gene and defined cell type. (e) Mouse brain cells in the t-SNE from Fig. 2a colored by per-cell log10 library complexity (n = 46,653 cells). (f) Per-cell log10 library complexity for each cluster derived from the mouse brain dscATAC-seq dataset. Center line, median; box limits, first and third quartiles; whiskers, 1.5x interquartile range. (g) Per-cell ratio of total reads in peaks to TSS reads per cluster. For number of cells per cluster see Supplementary Table 2.
Supplementary Figure 6 Species mixing analysis of dsciATAC-seq.
(a, b) Species mixing analysis for human (K562) and mouse (3T3) cell mix generated using (a) 24 or (b) 48 Tn5 transposase barcodes. For each panel a schematic of the experimental procedure is included (left), and primary results from a cell titration plotting total mouse or human nuclear fragments (right). In these plots points are labeled as either low quality (black), mouse (red), human (blue) or mixed (purple).
Supplementary Figure 7 Quality control analysis of human bone marrow dsciATAC-seq data.
(a–f) Single-cell data derived from BMMCs colored by their (a) donor, (b) fraction of reads in peaks (FRiP), (c) log10 unique nuclear fragments, (d) log10 total aligned nuclear fragments, (e) log10 library size, and (f) fraction of reads with PCR duplicates. (g) De novo embedding and clustering of the human BMMC data using the 7-mer k-mer strategy. Colors represent Louvain clustering from the principal components of the 7-mer deviations. (h, i) Same coordinates as (g) but colored according to annotations defined in Fig. 4b, c, respectively. All panels show n = 60,495 cells.
Supplementary Figure 8 Cell types identified in the human bone marrow dsciATAC-seq data.
(a) Selected transcription factor deviation motifs shown for resting cells (n = 60,495 cells) profiled using dsciATAC-seq. (b) Embedded cells from isolated subtypes profiled using the standard dscATAC-seq platform (n = 52,873 cells). (c) UMAP embedding of single-cell data colored by clusters identified (compare to Fig. 4c). (d–f) Projection of additional single-cell data onto UMAP coordinates of the dsciATAC-seq bone marrow data, projecting (d) sorted progenitor subsets (Buenrostro et al., 173(6), 1535–1548.e16, 2018), (e) peripheral blood mononuclear cells (PBMCs) or (f) isolated subsets (shown individually in (b)).
Supplementary Figure 9 Stimulation of human bone marrow derived cells.
(a) Stimulated BMMC (n = 75,968) cells projected onto the UMAP coordinates defined by the non-stimulated control cells (n = 60,495 cells). (b,c) Cell-cell TF score variability for the stimulation and control cells showing (b) ex vivo culture and (c) ex vivo culture and LPS stimulation, only unique TF motifs are highlighted. (d, e) Cell-cell TF score variability for the control cells and variability of stimulation after normalizing to the control TF variability for (d) ex vivo culture and (e) ex vivo culture and LPS stimulation conditions, only unique TF motifs are highlighted. (f-j) Depictions of transcription factor deviation scores in resting cells (top) compared to the differential after stimulation (bottom) for selected motifs. A total of n = 60,495 cells are plotted. (k) Sample summary of differential peak analysis for the Mono-1 cluster. Each dot represents a chromatin accessibility peak found in at least 1% of cells. The overall % of cells with element are shown on the x-axis whereas the y-axis depicts the difference in the % of cells with the element accessible (stimulated - resting). Peaks found significantly different at a 1% FDR (two-sided binomial test; Benjamini Hochberg corrected) are colored in red and blue. (l) Overall summary statistics per-population from differential peak analysis showing the Z-statistic from the two-sided permutation test for differential accessibility. Each colored curve represents the overall Z-statistics for all peaks in the specified cluster.
Supplementary information
Supplementary Information
Supplementary Figures 1–9 and Supplementary Note
Supplementary Table 1
Mouse brain per cluster promoter region chromatin accessibility scores for marker genes from Saunders et al.
Supplementary Table 2
Optimal matches to mouse brain clusters from scRNA-seq datasets.
Supplementary Table 3
Cluster-specific peaks identified in the mouse brain.
Supplementary Table 4
Stimulus-responsive chromatin accessibility peaks determined using two-sided permutation test and adjusted using Benjamini–Hochberg.
Supplementary Table 5
Human peripheral blood and bone marrow cell donor information.
Supplementary Table 6
Oligonucleotides used in this study.
Rights and permissions
About this article
Cite this article
Lareau, C.A., Duarte, F.M., Chew, J.G. et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat Biotechnol 37, 916–924 (2019). https://doi.org/10.1038/s41587-019-0147-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0147-6
This article is cited by
-
txci-ATAC-seq: a massive-scale single-cell technique to profile chromatin accessibility
Genome Biology (2024)
-
scifi-ATAC-seq: massive-scale single-cell chromatin accessibility sequencing using combinatorial fluidic indexing
Genome Biology (2024)
-
Simultaneous single-cell analysis of 5mC and 5hmC with SIMPLE-seq
Nature Biotechnology (2024)
-
Cell-type-directed design of synthetic enhancers
Nature (2024)
-
Fast and flexible profiling of chromatin accessibility and total RNA expression in single nuclei using Microwell-seq3
Cell Discovery (2024)