Abstract
Nitrogen-use efficiency of indica varieties of rice is superior to that of japonica varieties. We apply 16S ribosomal RNA gene profiling to characterize root microbiota of 68 indica and 27 japonica varieties grown in the field. We find that indica and japonica recruit distinct root microbiota. Notably, indica-enriched bacterial taxa are more diverse, and contain more genera with nitrogen metabolism functions, than japonica-enriched taxa. Using genetic approaches, we provide evidence that NRT1.1B, a rice nitrate transporter and sensor, is associated with the recruitment of a large proportion of indica-enriched bacteria. Metagenomic sequencing reveals that the ammonification process is less abundant in the root microbiome of the nrt1.1b mutant. We isolated 1,079 pure bacterial isolates from indica and japonica roots and derived synthetic communities (SynComs). Inoculation of IR24, an indica variety, with an indica-enriched SynCom improved rice growth in organic nitrogen conditions compared with a japonica-enriched SynCom. The links between plant genotype and root microbiota membership established in this study will inform breeding strategies to improve nitrogen use in crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw sequence data reported in this paper have been deposited (PRJCA001214) in the Genome Sequence Archive in the BIG Data Center62, Chinese Academy of Sciences under accession codes CRA001372 for bacterial 16S rRNA gene sequencing data and CRA001362 for metagenomic sequencing data that are publicly accessible at http://bigd.big.ac.cn/gsa. All pure strains (Supplementary Table 11) are deposited in two national culture collection centers, the China Natural Gene Bank and the Agricultural Culture Collection of China. All information about these strains, such as the 16S rRNA gene sequences, taxonomy and isolation details, as well as any further updates are available at http://bailab.genetics.ac.cn/culture_collection/.
Code availability
Scripts employed in the computational analyses are available at https://github.com/microbiota/Zhang2019NBT.
References
Bakker, P., Pieterse, C. M. J., De Jonge, R. & Berendsen, R. L. The soil-borne legacy. Cell 172, 1178–1180 (2018).
Berendsen, R. L., Pieterse, C. M. & Bakker, P. A. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486 (2012).
Hacquard, S. et al. Microbiota and host nutrition across plant and animal kingdoms. Cell Host Microbe 17, 603–616 (2015).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Lundberg, D. S. et al. Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90 (2012).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Edwards, J. et al. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl Acad. Sci. USA 112, E911–E920 (2015).
Peiffer, J. A. et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl Acad. Sci. USA 110, 6548–6553 (2013).
Schlaeppi, K., Dombrowski, N., Oter, R. G., Ver Loren van Themaat, E. & Schulze-Lefert, P. Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc. Natl Acad. Sci. USA 111, 585–592 (2014).
Zarraonaindia, I. et al. The soil microbiome influences grapevine-associated microbiota. mBio 6, e02527–14 (2015).
Breidenbach, B., Pump, J. & Dumont, M. G. Microbial community structure in the rhizosphere of rice plants. Front. Microbiol. 6, 1537 (2016).
Castrillo, G. et al. Root microbiota drive direct integration of phosphate stress and immunity. Nature 543, 513–518 (2017).
Edwards, J. A. et al. Compositional shifts in root-associated bacterial and archaeal microbiota track the plant life cycle in field-grown rice. PLoS Biol. 16, e2003862 (2018).
Lebeis, S. L. et al. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349, 860–864 (2015).
Muller, D. B., Vogel, C., Bai, Y. & Vorholt, J. A. The plant microbiota: systems-level insights and perspectives. Annu. Rev. Genet. 50, 211–234 (2016).
Walters, W. A. et al. Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc. Natl Acad. Sci. USA 28, 7368–7373 (2018).
Zhang, J. Y. et al. Root microbiota shift in rice correlates with resident time in the field and developmental stage. Sci. China Life Sci. 61, 613–621 (2018).
Oka, H. I. Intervarietal variation and classification of cultivated rice. Ind. J. Genet. Plant. Breed. 18, 10 (1958).
Zhang, Q. F., Maroof, M. A., Lu, T. Y. & Shen, B. Z. Genetic diversity and differentiation of indica and japonica rice detected by RFLP analysis. Theor. Appl. Genet. 83, 495–499 (1992).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Morishima, H. & Oka, H. I. Phylogenetic differentiation of cultivated rice. XXVII. Numerical evaluations of the indica-japonica differentiation. Jpn. J. Breed. 31, 402–413 (1981).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Koutroubas, S. D. & Ntanos, D. A. Genotypic differences for grain yield and nitrogen utilization in Indica and Japonica rice under mediterranean conditions. Field Crop. Res. 83, 251–260 (2003).
Rakotoson, T. et al. Genetic variability of nitrogen use efficiency in rainfed upland rice. Field Crop. Res. 213, 194–203 (2017).
Hu, B. et al. Variation in NRT1.1B contributes to nitrate-use divergence between rice subspecies. Nat. Genet. 47, 834–838 (2015).
Wang, W. et al. Expression of the nitrate transporter OsNRT1.1A/OsNPF6.3 confers high yield and early maturation in rice. Plant Cell 30, 638–651 (2018).
Näsholm, T., Kielland, K. & Ganeteg, U. Uptake of organic nitrogen by plants. New Phytol. 182, 31–48 (2009).
Bloom, A. J. The increasing importance of distinguishing among plant nitrogen sources. Curr. Opin. Plant Biol. 25, 10–16 (2015).
Santos-Medellin, C., Edwards, J., Liechty, Z., Nguyen, B. & Sundaresan, V. Drought stress results in a compartment-specific restructuring of the rice root-associated microbiomes. mBio 8, e00764–00717 (2017).
Shenton, M., Iwamoto, C., Kurata, N. & Ikeo, K. Effect of wild and cultivated rice genotypes on rhizosphere bacterial community composition. Rice 9, 42 (2016).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R News 2, 18–22 (2002).
Louca, S., Parfrey, L. W. & Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science 353, 1272–1277 (2016).
Chanh, T. T., Tsutsumi, M. & Kurihara, K. Comparative study on the response of Indica and Japonica rice plants to ammonium and nitrate nitrogen. Soil Sci. Plant Nutr. 27, 83–92 (1981).
Chao, D. Y. & Lin, H. X. Nitrogen-use efficiency: transport solution in rice variations. Nat. Plants 1, 15096 (2015).
Ta, T. C. & Ohira, K. Effects of various environmental and medium conditions on the response of Indica and Japonica rice plants to ammonium and nitrate nitrogen. Soil Sci. Plant Nutr. 27, 347–355 (1981).
Kanehisa, M. et al. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res. 42, 199–205 (2014).
Gao, H. et al. Reduction of nitrate in Shewanella oneidensis depends on atypical NAP and NRF systems with NapB as a preferred electron transport protein from CymA to NapA. ISME J. 3, 966 (2009).
Malm, S. et al. The roles of the nitrate reductase NarGHJI, the nitrite reductase NirBD and the response regulator GlnR in nitrate assimilation of mycobacterium tuberculosis. Microbiology 155, 1332–1339 (2009).
Ho, C. H., Lin, S. H., Hu, H. C. & Tsay, Y. F. CHL1 functions as a nitrate sensor in plants. Cell 138, 1184–1194 (2009).
Nuan, G. et al. Study on the determination of organic matter in soil by the oven heating method. Acta Agricult. Boreali-Sinica 32, 343–346 (2017).
Zhang, L. H. et al. NRT1.1B improves selenium concentrations in rice grains by facilitating selenomethinone translocation. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13037 (2019).
Lindsay, E. L. & Norvell, W. A. Development of a DTPA soil test for zinc, iron, manganese, and copper. Soil Sci. Soc. Am. J. 42, 421–428 (1978).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, 590–596 (2012).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microb. 73, 5261–5267 (2007).
Bolyen, E. et al. QIIME 2: reproducible, interactive, scalable, and extensible microbiome data science. PeerJ Preprints 6, e27295v27291 (2018).
Wickham, H. ggplot2: Elegant Graphics For Data Analysis (Springer, 2016).
Chen, Y. et al. SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 7, 1–6 (2017).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Li, D. et al. MEGAHITv1. 0: A fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124 (2014).
Wang, H. et al. The power of inbreeding: NGS-based GWAS of rice reveals convergent evolution during rice domestication. Mol. Plant 9, 975–985 (2016).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. Preprint at https://doi.org/10.1101/081257 (2016).
Cole, J. R. et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642 (2014).
Asnicar, F., Weingart, G., Tickle, T. L., Huttenhower, C. & Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 3, e1029 (2015).
Members, B. I. G. D. C. Database resources of the BIG Data Center in 2018. Nucleic Acids Res. 46, D14–D20 (2018).
Acknowledgements
We thank P. Schulze-Lefert and S. Hacquard at the Max Planck Institute for Plant Breeding Research for their suggestions for improving the manuscript. This work was financially supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (grant nos. XDB11020700 to Y.B. and XDA08010104 to C.C.), the Key Research Program of Frontier Sciences of the Chinese Academy of Science (grant nos. QYZDB-SSW-SMC021 to Y.B. and QYZDJ-SSW-SMC014 to C.C.), the National Natural Science Foundation of China (grant nos. 31772400 to Y.B. and 31801945 to J.Z.), and the Key Research Program of the Chinese Academy of Sciences (grant no. KFZD-SW-219 to Y.B.). J.Z. is supported by the CPSF-CAS Joint Foundation for Excellent Postdoctoral Fellows (grant no. 2016LH00012). Y.B. is supported by the Thousand Youth Talents Plan (grant no. 2060299).
Author information
Authors and Affiliations
Contributions
C.C. and Y.B. conceived the study and supervised the project. J.Z. and N.Z. performed the experiments. Y.-X.L. analyzed the data of 16S rRNA gene profiling. B.H. and L.Y. coordinated field experiments and revised the manuscript. T.J., P.Y. and R.G.-O. analyzed the metagenomic data. X.Z., Y.Q. and G.F. were involved in the informatics analysis. J.H. performed the soil properties analysis. S.C., H.X., X.W., C.W., H.W. and B.Q. participated in growing plants and harvesting samples. X.G. optimized the protocol of library preparation for the 16S rRNA gene profiling. J.Z., C.C. and Y.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Coverage of members in the root bacterial microbiota by the representative indica and japonica varieties.
(a) Rarefaction curves of detected bacterial species of the root microbiota reach the saturation stage with increasing numbers of samples, indicating that the root microbiota in our population capture most root bacteria members from each rice subspecies. Indica and japonica varieties in two locations are shown separately. (b) Rarefaction curves of detected bacterial OTUs of the root microbiota from indica and japonica varieties reach saturation stage with increasing sequencing depth. Each vertical bar represents standard error. The numbers of replicated samples in this figure are as follows: in field I, indica (n = 201), japonica (n = 80), soil (n = 12); in field II, indica (n = 201), japonica (n = 81), soil (n = 12).
Supplementary Fig. 2 Comparison of the root microbiota of indica and japonica varieties.
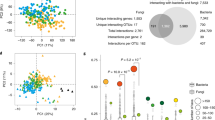
(a,b) Principal coordinate analysis with unweighted (a) and weighted (b) UniFrac distance show that the root microbiota of indica separate from that of japonica in field I in the first two axes, indicating that the root microbiota of indica are distinct from that of japonica (P < 0.001, PERMANOVA by Adonis). Ellipses cover 68% of the data for each rice subspecies. (c,d) Principal coordinate analysis with unweighted (c) and weighted (d) UniFrac distance showing that the root microbiota of indica separate from those of japonica in field II in the first two axes, revealing that root microbiota of indica are distinct from those of japonica (P < 0.001, PERMANOVA by Adonis). The numbers of replicated samples are as follows: in field I, indica (n = 201), japonica (n = 80); in field II, indica (n = 201), japonica (n = 81).
Supplementary Fig. 3 Comparison of the root microbiota in two fields.
(a,b) Unconstrained (a) and constrained (b) principal coordinate analysis of indica in field I, japonica in field I, indica in field II, and japonica in field II with Bray-Curtis distance. (c,d) Principal coordinate analysis of indica in field I, japonica in field I, indica in field II, and japonica in field II with unweighted (c) and weighted UniFrac (d) distance. Ellipses cover 68% of the data for each rice subspecies. The numbers of replicated samples are as follows: in field I, indica (n = 201), japonica (n = 80); in field II, indica (n = 201), japonica (n = 81).
Supplementary Fig. 4 Comparison of within-sample diversity (α-diversity) between indica and japonica.
(a,b) Chao 1 (a) and observed OTUs (b) of the root microbiota of indica, japonica, and corresponding unplanted bulk soils in two fields. The numbers of replicated samples are as follows: in field I, indica (n = 201), japonica (n = 80), soil (n = 12); in field II, indica (n = 201), japonica (n = 81), soil (n = 12). Data in two locations show the consistent trend that the root microbiota of indica show higher alpha diversity than those of japonica. The horizontal bars within boxes represent median. The tops and bottoms of boxes represent 75th and 25th percentiles, respectively. The upper and lower whiskers extend to data no more than 1.5 × the interquartile range from the upper edge and lower edge of the box, respectively.
Supplementary Fig. 5 Taxonomic composition of the indica- and japonica-enriched OTUs.
(a,b) The relative abundance of indica-enriched (a) and japonica-enriched (b) OTUs at the phylum level. Proteobacteria are shown at the class level.
Supplementary Fig. 6 Function and time-series shift of the indica- and japonica-enriched OTUs.
(a) Functional annotation of indica-enriched OTUs by FAPROTAX. The presence of functions is shown in red. (b) Shift of relative abundance of indica-enriched OTUs according to time-course data from the rice root microbiota in the field in Changping Farm17. (c) Functional annotation of japonica-enriched OTUs by FAPROTAX. The presence of functions is shown in red. (d) Shift of relative abundance of japonica-enriched OTUs according to time-course data from the rice root microbiota in the field in Changping Farm17.
Supplementary Fig. 7 Correlation between the natural variation of NRT1.1B and nitrogen-related functions in indica and japonica populations.
(a–e) The natural variation in NRT1.1B in indica and japonica populations is correlated with nitrogen-related functions in root microbiota from indica and japonica populations, including nitrite ammonification (a) (P = 2.2 × 10–16 in field I; P = 1.8 × 10–12 in field II, two-sided t-test), nitrate reduction (b) (P = 1.1 × 10–13 in field I; P = 2.1 × 10–8 in field II, two-sided t-test), respiration of nitrate (c) (P = 2.2 × 10–16 in field I; P = 6.1 × 10–13 in field II, two-sided t-test), nitrite respiration (d) (P = 2.4 × 10–16 in field I; P = 7.2 × 10–13 in field II, two-sided t-test) and nitrogen respiration (e) (P = 2.2 × 10–16 in field I; P = 6.8 × 10–13 in field II, two-sided t-test). NRT1.1Bindica harbors a “T” at 980 bp downstream of the ATG start codon and NRT1.1Bjaponica harbors a “C” at the same position, resulting in an amino acid substitution (p. Met327Thr). The numbers of replicated samples are as follows: in field I, indica (n = 192), japonica (n = 86); in field II, indica (n = 192), japonica (n = 87).
Supplementary Fig. 8 NRT1.1B and its natural variation modulate the assembly of the rice root microbiota.
(a,b) Principal coordinate analysis with unweighted (a) and weighted (b) UniFrac distance showing that the root microbiota of ZH11 (wild-type), nrt1.1b, Nipponbare NRT1.1Bindica, and Nipponbare NRT1.1Bjaponica separate in the first two axes. Ellipses cover 68% of the data for each genotype. (c) Constrained principal coordinate analysis showing that 57.1% of the root microbiota variations are explained by genotypes (ZH11, nrt1.1b, NRT1.1Bindica, and NRT1.1Bjaponica). The numbers of replicated samples are as follows: ZH11 (n = 16), nrt1.1b (n = 14), NRT1.1Bindica (n = 15), NRT1.1Bjaponica (n = 15). (d,e) A full factorial replication experiment validates the conclusion that NRT1.1B and its natural variation modulate the assembly of the rice root microbiota. Unconstrained (d) and constrained (e) principal coordinate analysis with Bray-Curtis distance showing that the root microbiota of ZH11 (wild-type), nrt1.1b, NRT1.1Bindica, and NRT1.1Bjaponica separate in the first two axes. The plants were grown in different fields and at different time from the samples in Fig. 4. Ellipses cover 68% of the data for each genotype. The numbers of replicated samples are as follows: ZH11 (n = 14), nrt1.1b (n = 10), NRT1.1Bindica (n = 15), NRT1.1Bjaponica (n = 15).
Supplementary Fig. 9 OTUs associated with NRT1.1B in the field condition related to Fig. 4.
(a) Enrichment and depletion of OTUs in the nrt1.1b mutant compared with wild-type ZH11. Each point represents an individual OTU, and the position along the x axis represents the abundance fold change between the nrt1.1b mutant and wild-type. (b) Heat map showing the relative abundance of the differential OTUs between the nrt1.1b mutant and wild-type ZH11. (c) OTUs enriched in NRT1.1Bindica or NRT1.1Bjaponica. Each point represents an individual OTU, and the position along the x axis represents the abundance fold change between NRT1.1Bindica or NRT1.1Bjaponica. (d) Heat map showing the relative abundance of the differential OTUs between NRT1.1Bindica and NRT1.1Bjaponica. The numbers of replicated samples are as follows: ZH11 (n = 16), nrt1.1b (n = 14), NRT1.1Bindica (n = 15), and NRT1.1Bjaponica (n = 15).
Supplementary Fig. 10 Experimental procedure for isolation and identification of rice root-associated bacteria.
Step 1–4 illustrate the isolation procedure; Step 5–11 illustrate the identification of cultivated rice root-associated bacteria by the improved two-step barcoded system.
Supplementary Fig. 11 The scheme for the previous high-throughput barcoding system to determine 16S rRNA gene sequences covering regions V5–V7 of bacterial isolates.
A previously published two-step barcoded pyrosequencing procedure (Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369, 2015). Please note that the second step PCR generates chimera sequences that will contain mislabeled plate and well barcodes for bacterial identification.
Supplementary Fig. 12 Plant growth with or without indica-enriched SynCom under inorganic nitrogen conditions.
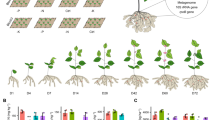
(a–c) IR24 (indica) rice plants were grown under different ratios of ammonium and nitrate (0:2, 2:0, and 1:1) with or without indica-enriched SynCom. After 2-week bacterial inoculation, rice plants were measured by root length (a), plant height (b), and shoot fresh weight (c). (d–f) Nipponbare (japonica) rice plants were grown under different ratios of ammonium and nitrate (0:2, 2:0, and 1:1) with or without indica-enriched SynCom. After 2-week bacterial inoculation, rice plants were measured by root length (d), plant height (e), and shoot fresh weight (f). Different letters indicate significantly different groups (P < 0.05, ANOVA, Tukey-HSD). Boxplots show combined data from three independent inoculation experiments with 4–5 technical replicates each (Supplementary Table 13). The horizontal bars within boxes represent medians. The tops and bottoms of boxes represent 75th and 25th percentiles, respectively. The upper and lower whiskers extend to data no more than 1.5 × the interquartile range from the upper edge and lower edge of the box, respectively.
Supplementary Fig. 13 Plant growth with indica- or japonica-enriched SynCom under inorganic nitrogen conditions.
IR24 rice plants were grown under the inorganic nitrogen condition with and without SynComs, including indica-enriched SynCom, japonica-enriched SynCom, and corresponding heat-killed bacteria as controls, respectively (Supplementary Table 13). After 2-week bacterial inoculation, rice plants were measured by root length (a), plant height (b), and shoot fresh weight (c). Different letters indicate significantly different groups (P < 0.05, ANOVA, Tukey-HSD). Boxplots show combined data from two independent inoculation experiments with 4–5 technical replicates each. The horizontal bars within boxes represent medians. The tops and bottoms of boxes represent 75th and 25th percentiles, respectively. The upper and lower whiskers extend to data no more than 1.5 × the interquartile range from the upper edge and lower edge of the box, respectively.
Supplementary information
Supplementary Figures
Supplementary Figs. 1–13
Supplementary Table 1
Soil properties and cultivation practices of field I and field II on Lingshui farm.
Supplementary Table 2
Information on indica and japonica varieties.
Supplementary Table 3
Metadata, OTU representative sequences, taxonomy annotation and OTU table.
Supplementary Table 4
Differential phyla and classes of Proteobacteria between indica and japonica.
Supplementary Table 5
Random-forest: accuracy of the random-forest model at each taxonomy level; feature importance at the family level; outcomes of prediction.
Supplementary Table 6
Differential abundance of OTUs between indica and japonica; details of OTUs in each part of the Venn diagrams in Fig. 3.
Supplementary Table 7
Abundance of OTUs in time-course data and functional annotation by FAPROTAX.
Supplementary Table 8
Differential abundances of OTUs between the nrt1.1b mutant and wild-type ZH11; differential abundances of OTUs between NRT1.1Bindica and NRT1.1Bjaponica; details of OTUs in each part of the Venn diagrams of Fig. 4.
Supplementary Table 9
KEGG orthology of metagenomes in ZH11 and the nrt1.1b mutant.
Supplementary Table 10
Detailed information of all cultivated CFUs and unique bacterial sequences from rice root.
Supplementary Table 11
Taxonomy and sequences of 1,079 bacterial stocks in rice root bacterial culture collection.
Supplementary Table 12
Experimental design of synthetic communities on germ-free plants related to the nitrogen assay.
Supplementary Table 13
Phenotypes of rice plants under inorganic nitrogen and organic nitrogen conditions with and without SynComs.
Supplementary Table 14
Primer sequences and experimental procedures for culture-independent community profiling and bacterial identification.
Rights and permissions
About this article
Cite this article
Zhang, J., Liu, YX., Zhang, N. et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat Biotechnol 37, 676–684 (2019). https://doi.org/10.1038/s41587-019-0104-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0104-4
This article is cited by
-
OsCIPK2 mediated rice root microorganisms and metabolites to improve plant nitrogen uptake
BMC Plant Biology (2024)
-
Selective regulation of endophytic bacteria and gene expression in soybean by water-soluble humic materials
Environmental Microbiome (2024)
-
Combined metagenomics and metabolomic analysis of microbial community structure and metabolic function in continuous soybean cropping soils of Songnen Plain, China
Chemical and Biological Technologies in Agriculture (2024)
-
Microbiome convergence enables siderophore-secreting-rhizobacteria to improve iron nutrition and yield of peanut intercropped with maize
Nature Communications (2024)
-
Exploitation of the microbiome for crop breeding
Nature Plants (2024)