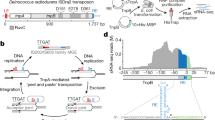
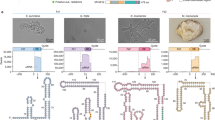
Abstract
The widespread TnpB proteins of IS200/IS605 transposon family have recently emerged as the smallest RNA-guided nucleases capable of targeted genome editing in eukaryotic cells1,2. Bioinformatic analysis identified TnpB proteins as the likely predecessors of Cas12 nucleases3,4,5, which along with Cas9 are widely used for targeted genome manipulation. Whereas Cas12 family nucleases are well characterized both biochemically and structurally6, the molecular mechanism of TnpB remains unknown. Here we present the cryogenic-electron microscopy structures of the Deinococcus radiodurans TnpB–reRNA (right-end transposon element-derived RNA) complex in DNA-bound and -free forms. The structures reveal the basic architecture of TnpB nuclease and the molecular mechanism for DNA target recognition and cleavage that is supported by biochemical experiments. Collectively, these results demonstrate that TnpB represents the minimal structural and functional core of the Cas12 protein family and provide a framework for developing TnpB-based genome editing tools.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data are available in the paper and the supplementary material. The atomic coordinates and cryo-EM density maps have been deposited in the Protein Data Bank and Electron Microscopy Data Bank under accession codes 8BF8/EMD-16016 (binary complex), 8EXA/EMD-28656 (ternary conformation 1 complex) and 8EX9/EMD-28655 (ternary conformation 2 complex). Source data are provided with this paper.
References
Karvelis, T. et al. Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease. Nature 599, 692–696 (2021).
Altae-Tran, H. et al. The widespread IS200/605 transposon family encodes diverse programmable RNA-guided endonucleases. Science 374, 57–65 (2021).
Kapitonov, V. V., Makarova, K. S. & Koonin, E. V. ISC, a novel group of bacterial and archaeal DNA transposons that encode Cas9 homologs. J. Bacteriol. 198, 797–807 (2016).
Shmakov, S. et al. Diversity and evolution of class 2 CRISPR–Cas systems. Nat. Rev. Microbiol. 15, 169–182 (2017).
Makarova, K. S. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Wang, J. Y., Pausch, P. & Doudna, J. A. Structural biology of CRISPR–Cas immunity and genome editing enzymes. Nat. Rev. Microbiol. 20, 641–656 (2022).
Koonin, E. V. Viruses and mobile elements as drivers of evolutionary transitions. Philos. Trans. R. Soc. B Biol. Sci. 371, 20150442 (2016).
Koonin, E. V., Makarova, K. S., Wolf, Y. I. & Krupovic, M. Evolutionary entanglement of mobile genetic elements and host defence systems: guns for hire. Nat. Rev. Genet. 21, 119–131 (2020).
Schuler, G., Hu, C. & Ke, A. Structural basis for RNA-guided DNA cleavage by IscB-ωRNA and mechanistic comparison with Cas9. Science 0, eabq7220 (2022).
Yamano, T. et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165, 949–962 (2016).
Gao, P., Yang, H., Rajashankar, K. R., Huang, Z. & Patel, D. J. Type V CRISPR-Cas Cpf1 endonuclease employs a unique mechanism for crRNA-mediated target DNA recognition. Cell Res. 26, 901–913 (2016).
Swarts, D. C., van der Oost, J. & Jinek, M. Structural basis for guide RNA processing and seed-dependent DNA targeting by CRISPR–Cas12a. Mol. Cell 66, 221–233.e4 (2017).
Stella, S., Alcón, P. & Montoya, G. Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage. Nature 546, 559–563 (2017).
Takeda, S. N. et al. Structure of the miniature type V-F CRISPR–Cas effector enzyme. Mol. Cell 81, 558–570 (2020).
Xiao, R., Li, Z., Wang, S., Han, R. & Chang, L. Structural basis for substrate recognition and cleavage by the dimerization-dependent CRISPR–Cas12f nuclease. Nucleic Acids Res. 49, 4120–4128 (2021).
Carabias, A. et al. Structure of the mini-RNA-guided endonuclease CRISPR-Cas12j3. Nat. Commun. 12, 4476 (2021).
Pausch, P. et al. DNA interference states of the hypercompact CRISPR–CasΦ effector. Nat. Struct. Mol. Biol. 28, 652–661 (2021).
Stella, S. et al. Conformational activation promotes CRISPR–Cas12a catalysis and resetting of the endonuclease activity. Cell 175, 1856–1871.e21 (2018).
Strohkendl, I., Saifuddin, F. A., Rybarski, J. R., Finkelstein, I. J. & Russell, R. Kinetic basis for DNA target specificity of CRISPR–Cas12a. Mol. Cell 71, 816–824.e3 (2018).
Swarts, D. C. & Jinek, M. Mechanistic insights into the cis- and trans-acting DNase activities of Cas12a. Mol. Cell 73, 589–600.e4 (2019).
Karvelis, T. et al. PAM recognition by miniature CRISPR–Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res. 48, 5016–5023 (2020).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Chen, J. S. et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Holm, L. Benchmarking fold detection by DaliLite v.5. Bioinforma. Oxf. Engl. 35, 5326–5327 (2019).
Bigelyte, G. et al. Miniature type V-F CRISPR–Cas nucleases enable targeted DNA modification in cells. Nat. Commun. 12, 6191 (2021).
Suzek, B. E. et al. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
Frickey, T. & Lupas, A. CLANS: a Java application for visualizing protein families based on pairwise similarity. Bioinformatics 20, 3702–3704 (2004).
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science 365, 48–53 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Yamano, T. et al. Structural basis for the canonical and non-canonical PAM recognition by CRISPR-Cpf1. Mol. Cell 67, 633–645.e3 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. Sect. Struct. Biol. 74, 519–530 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. Sect. Struct. Biol. 75, 861–877 (2019).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Al-Shayeb, B. et al. Clades of huge phages from across Earth’s ecosystems. Nature 578, 425–431 (2020).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Crooks, G. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
The research has been supported by the Novozyme Prize grant to V.S. and the Research Council of Lithuania grant no. S-MIP-21-8 to T.K. We thank the Danish Cryo-EM National Facility in CFIM at the University of Copenhagen and especially T. Pape and N. Sofos for support during cryo-EM data collection. We also thank I. Drulyte (Thermo Fisher Scientific) for invaluable help with cryo-EM sample preparation and data analysis, S. Erlendsson for preprocessing the initial data and G. Bigelyte for the help with plasmid construction. G.M. is part of the Novo Nordisk Foundation Center for Protein Research, which is supported financially by the Novo Nordisk Foundation (grant no. NNF14CC0001). This work was also supported by grant nos. NNF0024386 and NNF17SA0030214 and Distinguished Investigator grant no. NNF18OC0055061 to G.M., who is a member of the Integrative Structural Biology Cluster at the University of Copenhagen, and the Lundbeck Foundation (Lundbeckfonden) postdoctoral grant no. R380-2021-1448 to A.C.
Author information
Authors and Affiliations
Contributions
G.S., G.T., G.D., A.C., G.M., T.K. and V.S. designed the research. G.D. and A.S. performed the protein purifications. G.S., G.T. and A.C. collected cryo-EM data. G.S. and G.T. prepared cryo-EM samples. G.S., G.T., A.C. and G.M. solved the structures. G.D. performed DNA binding and cleavage assays. G.S., G.T., D.K., Č.V. and T.K. performed computational sequence and structure analyses. G.S., G.T., T.K. and V.S. wrote the manuscript with input from all authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
T.K. and V.S. are co-inventors on a patent application (PCT/IB2021/055958) filed by Vilnius University. V.S. is a Chairman of and has financial interest in CasZyme. G.M. is a shareholder and a member of Ensoma SAB. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks Yong-Sam Kim, Dipa Sashital and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification of ISDra2 TnpB RNP and design of DNA oligoduplexes.
a, Experimental workflow showing expression and multi-step purification of TnpB RNP complex. Addition of an HDV ribozyme sequence at the 3′-end ensured production of reRNA with a fixed 16-nt length guide sequence. b, SDS-PAGE analysis of the purified TnpB and TnpB (D191A) RNP complexes. Different amounts of purified protein samples were loaded on the gel. c, EMSA analysis of TnpB binding to the product-like partially single-stranded (left), substrate-like double-stranded (center) and nonspecific double-stranded (right) DNA oligoduplexes. d, Design of the head-to-head dual-end DNA oligoduplex that was used for structural analysis of TnpB-reRNA-DNA ternary complexes. e, EMSA analysis of TnpB binding to the dual-end DNA oligoduplex. Positions of complexes with one and two TnpB RNPs bound are indicated. f, Interpretation of cryo-EM data obtained with the dual-end DNA. During data processing, TnpB-reRNA-DNA complexes formed at each terminus of the dual-end DNA were interpreted as individual particles. The resultant cryo-EM reconstruction (bottom) in addition to the high quality map of the “primary complex” used for model building, also contained residual density corresponding to part of the second complex. Two copies of TnpB-reRNA-DNA complexes are shown fitted into both regions of the map. The angle between the long axes of these structures (blue lines) is 77°. Such fixed orientation of two TnpB RNPs bound to the dual-end DNA altered spatial distribution of the specimen, thereby alleviating the preferred orientation related problems. The sequences of the oligonucleotides are listed in Supplementary information Table 3. For uncropped gel images, see Supplementary information Fig. 1.
Extended Data Fig. 2 CryoEM single particle reconstruction of TnpB-reRNA binary and TnpB-reRNA-DNA ternary (conformation 1) complexes.
a, Workflow of the cryo-EM image processing and 3D reconstruction for the TnpB-reRNA binary complex. b, Workflow of the cryo-EM image processing and reconstruction for the TnpB-reRNA-DNA ternary conformation 1 (resolved RuvC domain) complex. The final electron density maps showing local resolution, masks from the local refinement jobs, directional distribution plots and FSC (Fourier shell correlation) plots are shown in black rectangles.
Extended Data Fig. 3 CryoEM single particle reconstruction of TnpB-reRNA-DNA ternary conformation 2 complex.
Two sets of data (untilted and tilted) were acquired and combined. The final electron density maps showing local resolution, mask from the local refinement job, directional distribution plots and FSC (Fourier shell correlation) plots are shown in a black rectangle.
Extended Data Fig. 4 Structural features of Deinococcus radiodurans ISDra2 TnpB ternary complexes.
a, Cryo-EM reconstruction (left) and cartoon representation (middle) of TnpB-reRNA-DNA ternary conformation 2 complex. Both sharpened (colored) and unsharpened (black outline) cryo-EM maps are shown. Bound nucleic acids and protein contacts in the complex are schematically depicted on the right. Dashed rectangular shapes encircle unresolved parts of the RNA/DNA. The dashed line on the cryo-EM map marks the boundary of the density corresponding to the primary (shown) and the secondary (cut-off) TnpB subunits, as explained in Extended Data Fig. 1f. b, Topology diagram of the ternary conformation 1 complex structure. Structural elements of the ZnF domains are based on an AlphaFold29 model. Helices and strands are shown as cylinders and arrows, respectively. Red circles mark the RuvC catalytic center residues D191, E278 and D361. c, Pairwise comparison of binary – ternary conformation 1, and ternary conformation 1 – ternary conformation 2 structures.
Extended Data Fig. 5 Effect of TAM mutations on binding and cleavage of dsDNA substrates.
Testing the TnpB RNP ability to bind (a and b, EMSA gels and their quantification plot, respectively), and cleave (c) dsDNA substrates, containing single nucleotide mutations in the TAM sequence. The dots in the line graph represent the mean of three independent reaction replicates (n = 3) ± standard deviation (s.d.). TAM and target sequences are highlighted in magenta and black, respectively. TS – target strand; NTS – non-target strand. P – cleavage products. D – catalytically inactive TnpB RNP complex (dTnpB – D191A). M – radiolabeled ssDNA ladder. For uncropped gel images, see Supplementary information Fig. 1.
Extended Data Fig. 6 TnpB-reRNA interactions with DNA.
a, Protein contacts to the first three base pairs of the RNA-DNA heteroduplex. b, The “lid” subdomain contacts. The “lid” subdomain sterically blocks the RuvC catalytic center and makes contacts to the C7:dG-7 and U8:dA-8 base pairs in RNA-DNA heteroduplex from the minor groove side. dsDNA oligonucleotide binding (c) and plasmid cleavage (d) by TnpB complex containing unmodified reRNA (TnpB-reRNA) and reRNA with altered targeting sequence (TnpB-reRNA_AG). Schematic representation of the reRNA and target sequences are provided at the top of the (c) panel. The sequences of the plasmids and oligonucleotides are listed in Supplementary information Table 1 and Supplementary information Table 3, respectively. For uncropped gel images, see Supplementary information Fig. 1.
Extended Data Fig. 7 Effect of single nucleotide mismatches on binding and cleavage of dsDNA substrates.
Testing the TnpB RNP ability to bind (a), and cleave (b) dsDNA substrates, containing single nucleotide mismatches at various positions (the number following “MM”) in the target sequence. The cleavage reactions were quenched at 0, 5, 15, and 60 min time points. TAM and target sequences are highlighted in magenta and black, respectively. NTS – non-target strand, P – cleavage products, Target – dsDNA substrate without mismatches, M – radiolabeled ssDNA marker. For uncropped gel images, see Supplementary information Fig. 1.
Extended Data Fig. 8 Structural and biochemical features of TnpB reRNA.
a, Schematic representation of unmodified (reRNA, left) and truncated (reRNA_trunc, right) reRNAs. The tetraloop introduced in truncated reRNA is shown in blue. b, Plasmid DNA cleavage by TnpB complex containing unmodified reRNA (TnpB-reRNA) and reRNA with truncated stem 1 (TnpB-reRNA_trunc). c, FQ reporter cleavage by TnpB-reRNA and TnpB-reRNA_trunc variants. The dots represent the mean of three independent reaction replicates (n = 3) ± standard deviation (s.d.). d, Comparison of RNA-DNA bound to ISDra2 TnpB and UnCas12f. Note that in TnpB ternary complex stem 1 is parallel to the RNA-DNA heteroduplex, but its UnCas12f (PDB ID: 7C7L)14 equivalent stem 4 is perpendicular to RNA-DNA, and position of TnpB stem 1 is occupied by the RuvC domain of the 2nd protein subunit. For uncropped gel images, see Supplementary information Fig. 1.
Extended Data Fig. 9 Conserved motifs in Cas12/TnpB RuvC domain.
Cas12/TnpB can be grouped into two supergroups ((Q/D)RD and (N/H)AD) according to the conserved residues (shown in a gray rectangle) in their RuvC-III motif. ISDra2 TnpB RuvC catalytic center residues D191, E278 and D361 are marked with red rectangles.
Supplementary information
Supplementary Information
Supplementary Figs. 1 (uncropped gel images for the Extended Data figures) and 2 (full graphs for collateral FQ-reporter trans-cleavage assay) and Supplementary Tables 1–3.
Supplementary Data
This file contains source data for Supplementary Fig. 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sasnauskas, G., Tamulaitiene, G., Druteika, G. et al. TnpB structure reveals minimal functional core of Cas12 nuclease family. Nature 616, 384–389 (2023). https://doi.org/10.1038/s41586-023-05826-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-05826-x
This article is cited by
-
Discovery and structural mechanism of DNA endonucleases guided by RAGATH-18-derived RNAs
Cell Research (2024)
-
Engineering a transposon-associated TnpB-ωRNA system for efficient gene editing and phenotypic correction of a tyrosinaemia mouse model
Nature Communications (2024)
-
Targeted mutagenesis in mice via an engineered AsCas12f1 system
Cellular and Molecular Life Sciences (2024)
-
Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors
Nature Biotechnology (2023)
-
Mechanistic and evolutionary insights into a type V-M CRISPR–Cas effector enzyme
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.