Abstract
In mice and humans, sleep quantity is governed by genetic factors and exhibits age-dependent variation1,2,3. However, the core molecular pathways and effector mechanisms that regulate sleep duration in mammals remain unclear. Here, we characterize a major signalling pathway for the transcriptional regulation of sleep in mice using adeno-associated virus-mediated somatic genetics analysis4. Chimeric knockout of LKB1 kinase—an activator of AMPK-related protein kinase SIK35,6,7—in adult mouse brain markedly reduces the amount and delta power—a measure of sleep depth—of non-rapid eye movement sleep (NREMS). Downstream of the LKB1–SIK3 pathway, gain or loss-of-function of the histone deacetylases HDAC4 and HDAC5 in adult brain neurons causes bidirectional changes of NREMS amount and delta power. Moreover, phosphorylation of HDAC4 and HDAC5 is associated with increased sleep need, and HDAC4 specifically regulates NREMS amount in posterior hypothalamus. Genetic and transcriptomic studies reveal that HDAC4 cooperates with CREB in both transcriptional and sleep regulation. These findings introduce the concept of signalling pathways targeting transcription modulators to regulate daily sleep amount and demonstrate the power of somatic genetics in mouse sleep research.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All reagents and raw data generated in this study are available from the corresponding author upon reasonable request. The raw and processed RNA-seq data have been deposited at NCBI Gene Expression Omnibus under accession GSE182907. The raw and processed ChIP–seq data have been deposited at NCBI Gene Expression Omnibus under accession GSE182805. Because of their large size, the source data for mouse sleep phenotype analysis will be provided upon reasonable request. Source data are provided with this paper.
Code availability
The automated SleepV software for video-based analysis of sleep–wake behaviours4 has been deposited at GitHub (https://github.com/wochiguodong/SleepV.git.)4. The custom-designed C++ language-based automated sleep staging software for EEG/EMG data has been deposited at Zenodo (https://doi.org/10.5281/zenodo.7102778).
References
Sehgal, A. & Mignot, E. Genetics of sleep and sleep disorders. Cell 146, 194–207 (2011).
Webb, J. M. & Fu, Y.-H. Recent advances in sleep genetics. Curr. Opin. Neurobiol. 69, 19–24 (2021).
Zheng, L. & Zhang, L. The molecular mechanism of natural short sleep: a path towards understanding why we need to sleep. Brain Sci. Adv. 8, 165–172 (2022).
Wang, G. et al. Somatic genetics analysis of sleep in adult mice. J. Neurosci. 42, 5617–5640 (2022).
Lizcano, J. M. et al. LKB1 is a master kinase that activates 13 kinases of the AMPK subfamily, including MARK/PAR-1. EMBO J. 23, 833–843 (2004).
Walkinshaw, D. R. et al. The tumor suppressor kinase LKB1 activates the downstream kinases SIK2 and SIK3 to stimulate nuclear export of class IIa histone deacetylases. J. Biol. Chem. 288, 9345–9362 (2013).
Funato, H. et al. Forward-genetics analysis of sleep in randomly mutagenized mice. Nature 539, 378–383 (2016).
Honda, T. et al. A single phosphorylation site of SIK3 regulates daily sleep amounts and sleep need in mice. Proc. Natl Acad. Sci. USA 115, 10458–10463 (2018).
Li, X., Song, S., Liu, Y., Ko, S.-H. & Kao, H.-Y. Phosphorylation of the histone deacetylase 7 modulates its stability and association with 14-3-3 proteins. J. Biol. Chem. 279, 34201–34208 (2004).
Wang, A. H. et al. Regulation of histone deacetylase 4 by binding of 14-3-3 proteins. Mol. Cell. Biol. 20, 6904–6912 (2000).
Chen, Z. et al. The role of histone deacetylase 4 during chondrocyte hypertrophy and endochondral bone development. Bone Joint Res. 9, 82–89 (2020).
Haberland, M., Montgomery, R. L. & Olson, E. N. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat. Rev. Genet. 10, 32–42 (2009).
Grubbs, J. J., Lopes, L. E., van der Linden, A. M. & Raizen, D. M. A salt-induced kinase is required for the metabolic regulation of sleep. PLoS Biol. 18, e3000220 (2020).
Banks, G. T. et al. Forward genetics identifies a novel sleep mutant with sleep state inertia and REM sleep deficits. Sci. Adv. 6, eabb3567 (2020).
Sunagawa, G. A. et al. Mammalian reverse genetics without crossing reveals Nr3a as a short-sleeper gene. Cell Rep. 14, 662–677 (2016).
Funato, H. & Yanagisawa, M. Hunt for mammalian sleep-regulating genes. Brain Sci. Adv. 7, 75–96 (2021).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Xu, J. et al. Regulation of sleep quantity and intensity by long and short isoforms of SLEEPY kinase. Sleep https://doi.org/10.1093/sleep/zsac198 (2022).
Gruffat, H., Manet, E. & Sergeant, A. MEF2-mediated recruitment of class II HDAC at the EBV immediate early gene BZLF1 links latency and chromatin remodeling. EMBO Rep. 3, 141–146 (2002).
Zhu, Y. et al. Class IIa HDACs regulate learning and memory through dynamic experience-dependent repression of transcription. Nat. Commun. 10, 3469 (2019).
Mihaylova, M. M. et al. Class IIa histone deacetylases are hormone-activated regulators of FOXO and mammalian glucose homeostasis. Cell 145, 607–621 (2011).
Chen, B. & Cepko, C. L. HDAC4 regulates neuronal survival in normal and diseased retinas. Science 323, 256–259 (2009).
Guo, X. et al. A short N-terminal domain of HDAC4 preserves photoreceptors and restores visual function in retinitis pigmentosa. Nat. Commun. 6, 8005 (2015).
LeGates, T. A., Fernandez, D. C. & Hattar, S. Light as a central modulator of circadian rhythms, sleep and affect. Nat. Rev. Neurosci. 15, 443–454 (2014).
Price, V., Wang, L. & D’Mello, S. R. Conditional deletion of histone deacetylase-4 in the central nervous system has no major effect on brain architecture or neuronal viability. J. Neurosci. Res. 91, 407–415 (2013).
Fischle, W. et al. Enzymatic activity associated with class II HDACs is dependent on a multiprotein complex containing HDAC3 and SMRT/N-CoR. Mol. Cell 9, 45–57 (2002).
Gvilia, I., Xu, F., McGinty, D. & Szymusiak, R. Homeostatic regulation of sleep: a role for preoptic area neurons. J. Neurosci. 26, 9426–9433 (2006).
Kroeger, D. et al. Galanin neurons in the ventrolateral preoptic area promote sleep and heat loss in mice. Nat. Commun. 9, 4129 (2018).
Yoo, S. H. et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl Acad. Sci. USA 101, 5339–5346 (2004).
Zhang, E. E. et al. A genome-wide RNAi screen for modifiers of the circadian clock in human cells. Cell 139, 199–210 (2009).
Okamoto, S. et al. Dominant-interfering forms of MEF2 generated by caspase cleavage contribute to NMDA-induced neuronal apoptosis. Proc. Natl Acad. Sci. USA 99, 3974–3979 (2002).
Kozhemyakina, E., Cohen, T., Yao, T.-P. & Lassar, A. B. Parathyroid hormone-related peptide represses chondrocyte hypertrophy through a protein phosphatase 2A/histone deacetylase 4/MEF2 pathway. Mol. Cell. Biol. 29, 5751 (2009).
Lundell, L. S., Massart, J., Altintas, A., Krook, A. & Zierath, J. R. Regulation of glucose uptake and inflammation markers by FOXO1 and FOXO3 in skeletal muscle. Mol. Metab. 20, 79–88 (2019).
Ahn, S. et al. A dominant-negative inhibitor of CREB reveals that it is a general mediator of stimulus-dependent transcription of c-fos. Mol. Cell Biol. 18, 967–977 (1998).
Graves, L. A. et al. Genetic evidence for a role of CREB in sustained cortical arousal. J. Neurophysiol. 90, 1152–1159 (2003).
Hendricks, J. C. et al. A non-circadian role for cAMP signaling and CREB activity in Drosophila rest homeostasis. Nat. Neurosci. 4, 1108–1115 (2001).
Wimmer, M. E., Cui, R., Blackwell, J. M. & Abel, T. Cyclic AMP response element-binding protein is required in excitatory neurons in the forebrain to sustain wakefulness. Sleep 44, zsaa267 (2020).
Lamph, W. W., Dwarki, V. J., Ofir, R., Montminy, M. & Verma, I. M. Negative and positive regulation by transcription factor cAMP response element-binding protein is modulated by phosphorylation. Proc. Natl Acad. Sci. USA 87, 4320–4324 (1990).
Li, J. et al. Nuclear accumulation of HDAC4 in ATM deficiency promotes neurodegeneration in ataxia telangiectasia. Nat. Med. 18, 783–790 (2012).
Sen, T. & Sen, N. Isoflurane-induced inactivation of CREB through histone deacetylase 4 is responsible for cognitive impairment in developing brain. Neurobiol. Dis. 96, 12–21 (2016).
Zada, D., Bronshtein, I., Lerer-Goldshtein, T., Garini, Y. & Appelbaum, L. Sleep increases chromosome dynamics to enable reduction of accumulating DNA damage in single neurons. Nat. Commun. 10, 895 (2019).
Zada, D. et al. Parp1 promotes sleep, which enhances DNA repair in neurons. Mol. Cell 81, 4979–4993.e4977 (2021).
Brüning, F. et al. Sleep-wake cycles drive daily dynamics of synaptic phosphorylation. Science 366, eaav3617 (2019).
Wang, Z. et al. Quantitative phosphoproteomic analysis of the molecular substrates of sleep need. Nature 558, 435–439 (2018).
Benito, E., Valor, L. M., Jimenez-Minchan, M., Huber, W. & Barco, A. cAMP response element-binding protein is a primary hub of activity-driven neuronal gene expression. J. Neurosci. 31, 18237–18250 (2011).
Sando, R. 3rd et al. HDAC4 governs a transcriptional program essential for synaptic plasticity and memory. Cell 151, 821–834 (2012).
Noya, S. B. et al. The forebrain synaptic transcriptome is organized by clocks but its proteome is driven by sleep. Science 366, eaav2642 (2019).
Kim, S. J. et al. Kinase signalling in excitatory neurons regulates sleep quantityand depth. Nature https://doi.org/10.1038/s41586-022-05450-1 (2022).
Lu, J., McKinsey, T. A., Zhang, C.-L. & Olson, E. N. Regulation of skeletal myogenesis by association of the MEF2 transcription factor with class II histone deacetylases. Mol. Cell 6, 233–244 (2000).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Yardeni, T., Eckhaus, M., Morris, H. D., Huizing, M. & Hoogstraten-Miller, S. Retro-orbital injections in mice. Lab. Anim. 40, 155–160 (2011).
Zhao, H. et al. Absence of stress response in dorsal raphe nucleus in modulator of apoptosis 1-deficient mice. Mol. Neurobiol. 56, 2185–2201 (2019).
Christakis, D. A., Ramirez, J. S. & Ramirez, J. M. Overstimulation of newborn mice leads to behavioral differences and deficits in cognitive performance. Sci. Rep. 2, 546 (2012).
Horsch, M. et al. Requirement of the RNA-editing enzyme ADAR2 for normal physiology in mice. J. Biol. Chem. 286, 18614–18622 (2011).
Liu, Q., Xu, Y., Wan, W. & Ma, Z. An unexpected improvement in spatial learning and memory ability in alpha-synuclein A53T transgenic mice. J. Neural Transm. 125, 203–210 (2018).
Peters, J., Dieppa-Perea, L. M., Melendez, L. M. & Quirk, G. J. Induction of fear extinction with hippocampal-infralimbic BDNF. Science 328, 1288–1290 (2010).
Soria-Gomez, E. et al. Habenular CB1 receptors control the expression of aversive memories. Neuron 88, 306–313 (2015).
Zhang, J. et al. Presynaptic Excitation via GABAB receptors in habenula cholinergic neurons regulates fear memory expression. Cell 166, 716–728 (2016).
Patti, C. L. et al. Effects of sleep deprivation on memory in mice: role of state-dependent learning. Sleep 33, 1669–1679 (2010).
Savelyev, S. A., Larsson, K. C., Johansson, A. S. & Lundkvist, G. B. Slice preparation, organotypic tissue culturing and luciferase recording of clock gene activity in the suprachiasmatic nucleus. J. Vis. Exp. 48, 2439 (2011).
Mei, L. et al. Long-term in vivo recording of circadian rhythms in brains of freely moving mice. Proc. Natl Acad. Sci. USA 115, 4276–4281 (2018).
Yang, J. et al. A quick protocol for the preparation of mouse retinal cryosections for immunohistochemistry. Open Biol. 11, 210076 (2021).
Castel, M., Belenky, M., Cohen, S., Wagner, S. & Schwartz, W. J. Light-induced c-Fos expression in the mouse suprachiasmatic nucleus: immunoelectron microscopy reveals co-localization in multiple cell types. Eur. J. Neurosci. 9, 1950–1960 (1997).
Xu, P. et al. NPAS4 regulates the transcriptional response of the suprachiasmatic nucleus to light and circadian behavior. Neuron 109, 3268–3282 (2021).
Ju, D. et al. Chemical perturbations reveal that RUVBL2 regulates the circadian phase in mammals. Sci. Transl. Med. 12, eaba0769 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357 (2012).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Acknowledgements
We thank M. Luo and F. Shao for sharing reagents; Y. Zhou, M. Zhao, Z. Li, W. Min, Y. Zhuang, Y. Sun, H. Huang, J. Wang and Y. Yin for technical assistance; and X. Wang, F. Shao, T. Chen, T. Han and M. Xu for discussion and comments on the manuscript. This work was supported by the National Major Project of China Science and Technology Innovation 2030 for Brain Science and Brain-Inspired Technology (2021ZD0203400 to Q.Liu), the innovation grant (Z181100001318004 to Q.Liu), and institution grants from the Beijing Municipal Commission of Science and Technology Commission and Chinese Ministry of Science and Technology.
Author information
Authors and Affiliations
Contributions
R.Z., G.W., Q.Li and J.X. executed most of the experiments with input from Q.Liu, E.E.Z., H.F. and M.Y. F.W. helped to generate multiple mouse strains. X.G., S.Z., Z.C., L.C. and B.S. performed EEG/EMG recording and analysis. R.Z. and D.J. performed RNA-seq and ChIP–seq analyses with help from B.L., R.G. and T.C. D.S. performed ex vivo recording of SCN of Per2::Luc mice. Q.S., Y.G. and C.W. performed AAV packaging and purification. F.M. and C.L. performed stereotactic injection of AAVs in the hypothalamus and thalamus. H.W., X.W. and H.L. helped with mouse husbandry. R.Z., G.W. and Q.Liu prepared the figures and Q.Liu wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 LKB1-SIK3 signalling regulates NREMS by suppressing HDAC4/5.
a, Representative images showing co-immunostaining of LKB1 (green) and mCherry or Cre (HA, red) neurons in the cortex, hippocampus, thalamus, midbrain and medulla of AAV-hSyn-mCherry or AAV-hSyn-cre injected Lkb1flox/flox mice. Scale bars, 50 μm. b and c, REMS (b) or Wake (c) amount of AAV-hSyn-mCherry (control (Ctrl), n = 11) or AAV-hSyn-cre (Lkb1KO, n = 12) injected Lkb1flox/flox mice. d-g, Hourly NREMS time (d), EEG power spectra analysis (e), and REMS (f) or Wake (g) amount of Lkb1flox/flox mice co-injected with AAV-hSyn-cre and AAV-hSyn-mCherry (Lkb1KO/mCherry, n = 7), AAV-hSyn-SIK3-ST221E (Lkb1KO/SIK3-S(T221E), n = 7), or AAV-hSyn-SLP-ST221E (Lkb1KO/SLP-S(T221E), n = 8). h, Schematic of H4N/H5N-VP16 interfering with transcriptional activities of endogenous HDAC4/5 proteins. i, Representative images (two independent experiments with similar results) showing the effects of H4N-VP16 and H5N-VP16 on expression of co-transfected 3x MEF2 response element (MRE)-copGFP reporter gene in AAVpro 293T cells. Scale bars, 200 μm. j-m, Hourly NREMS time (j), EEG power spectra analysis (k), and REMS (l) or Wake (m) amount of Lkb1flox/flox mice injected with AAV-hSyn-cre and AAV-hSyn-mCherry (Lkb1KO/mCherry, n = 7) or AAV-hSyn-H4N/H5N-VP16 (Lkb1KO/H4N/H5N-VP16, n = 6). Data are mean ± s.e.m. Two-way ANOVA with Sidak’s test (b, c, l and m); mixed-effects model (d and j); one-way ANOVA with Dunnett’s test (e); two-way ANOVA with Dunnett’s test (f and g); and two-tailed unpaired t-test (k). n refers to the number of biological replicates.
Extended Data Fig. 2 ABC knockout of HDAC4/5 increases NREMS amount and delta power.
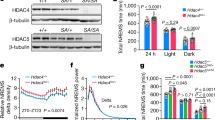
a-d, Hourly NREMS time (a), EEG power spectra analysis (b), and REMS (c) or Wake (d) amount of C57BL/6J mice injected with AAV-hSyn-mCherry (n = 15), AAV-hSyn-H4N-VP16 (n = 7) or AAV-hSyn-H4N/H5N-VP16 (n = 7). e, Schematic of ABC-KO of Hdac4 or Hdac5 by triple-target CRISPR. f and g, Immunoblotting (f) and quantification (g) of HDAC4 and HDAC5 proteins in whole brain lysates of Cas9 mice (n = 3 per group) injected with AAV-sgRNANT (ABC-NT), AAV-sgRNAHD4 (ABC-Hdac4KO), AAV-sgRNAHD5 (ABC-Hdac5KO), or AAV-sgRNAHD4 and AAV-sgRNAHD5 (ABC-Hdac4/5DKO). h-m, NREMS (h), REMS (i) or Wake (j) amount, hourly NREMS time (k), delta power density (l) and EEG power spectra analysis (m) of ABC-NT, ABC-Hdac4KO, ABC-Hdac5KO, or ABC- Hdac4/5DKO mice (n = 8 per group). n-q, Hourly NREMS time (n), EEG power spectra analysis (o), and REMS (p) or Wake (q) amount of AAV-hSyn-mCherry (ABC-Hdac4WT, n = 10) or AAV-hSyn-cre (ABC-Hdac4KO, n = 9) injected Hdac4flox/flox mice and AAV-hSyn-mCherry (ABC-Hdac5WT, n = 11) or AAV-hSyn-cre (ABC-Hdac5KO, n = 8) injected Hdac5flox/flox mice. Data are mean ± s.e.m. Mixed-effects model (a, k, l and n); one-way ANOVA with Dunnett’s test (b, g and m); two-way ANOVA with Sidak’s test (c-d, h—j and p-q); and two-tailed unpaired t-test (o). See Supplementary Fig. 1 for gel source data. n refers to the number of biological replicates.
Extended Data Fig. 3 ABC-Hdac4KO mice display normal vision and behaviors.
a-d, Hourly NREMS time (a), EEG power spectra analysis (b), and REMS (c) or Wake (d) amount of AAV-hSyn-rTTA and AAV-TRE-cre co-injected Hdac4flox/flox (iHdac4KO) mice without (-Dox, n = 6) or with (+Dox, n = 8) doxycycline treatment. e, Representative images (two independent experiments with similar results) showing co-immunostaining of S-Opsin (green) and Rhodopsin (red) in the retina of AAV-hSyn-mCherry (ABC-Ctrl) or AAV-hSyn-cre (ABC-Hdac4KO) injected Hdac4flox/flox mice. Scale bars, 20 μm. f, Representative images showing immunostaining of c-Fos (green) in the SCN of ABC-Ctrl or ABC-Hdac4KO mice after 15 min of light exposure during the dark phase. Scale bars, 50 μm. g, Quantification of light-induced c-Fos expression in (f) (n = 3). h-l, Fear conditioning (h), open field (i, j), sucrose preference (k), and rotarod (l) behavioral tests of ABC-Ctrl (n = 7) and ABC-Hdac4KO (n = 8) mice. Data are mean ± s.e.m. Mixed-effects model (a); two-way ANOVA with Sidak’s test (c-d, i, and k-l); and two-tailed unpaired t-test, two tailed (b, g-h and j). n refers to the number of biological replicates.
Extended Data Fig. 4 ABC expression HDAC4(CN) or HDAC5(CN) reduces NREMS amount and delta power.
a-d, Hourly NREMS time (a), EEG power spectra analysis (b), and REMS (c) or Wake (d) amount of AAV-hSyn-mCherry (n = 9), AAV-hSyn-HDAC4WT (n = 10) or AAV-hSyn-HDAC4CN (n = 8) injected C57BL/6J mice. e-h, Hourly NREMS time (e), EEG power spectra analysis (f), and REMS (g) or Wake (h) amount of AAV-hSyn-mCherry (n = 18), AAV-hSyn-HDAC5WT (n = 10) or AAV-hSyn-HDAC5CN (n = 9) injected C57BL/6J mice. i-l, Hourly NREMS time (i), EEG power spectra analysis (j), and REMS (k) or Wake (l) amount of AAV-hSyn-mCherry (n = 8), AAV-hSyn-HDAC7WT (n = 8) or AAV-hSyn-HDAC7CN (n = 9) injected C57BL/6J mice. m-p, Hourly NREMS time (m), EEG power spectra analysis (n), and REMS (o) or Wake (p) amount of AAV-hSyn-mCherry (n = 8), AAV-hSyn-HDAC4CN (n = 9), AAV-hSyn-HDAC4CN/D934N (n = 9) or AAV-hSyn-HDAC4CN/D840N (n = 6) injected C57BL/6J mice. Data are mean ± s.e.m. Mixed-effects model (a, e and m); one-way ANOVA with Dunnett’s test (b, f, j and n); and two-way ANOVA with Dunnett’s test (c-d, g-h, k-l and o-p). n refers to the number of biological replicates.
Extended Data Fig. 5 Regulation of HDAC4/5 by sleep-wake cycle and homeostatic sleep response of ABC-Hdac4KO and ABC-Hdac4CN mice.
a and b, Immunoblotting (a) and quantification (b) of S245 phosphorylation of HDAC4/5 in total lysates of cortex from C57BL/6J mice at ZT2 and ZT14 (n = 3). c and d, Immunoblotting (c) and quantification (d) of nuclear:cytoplasmic ratio of HDAC4/5 in cytoplasmic and nuclear extracts of cortex from C57BL/6J mice at ZT2 and ZT14 (n = 3). e and i, Immunoblotting of total lysates of cortex (e) or hypothalamus (i) from 6-h slept (S6) and sleep-deprived (SD6) C57BL/6J mice. f and j, Quantification of S245 phosphorylation of HDAC4/5 in (e) and (i) (n = 3). g and k, Immunoblotting of cytoplasmic and nuclear extracts of cortex (g) or hypothalamus (k) from S6 and SD6 mice. h and l, Quantification of nuclear:cytoplasmic ratio of HDAC4/5 in (g) and (k) (n = 3). m-p, Hourly NREMS time (m) or delta power density (o) and accumulated NREMS time (n) or delta power (p) change of AAV-hSyn-GFP (Ctrl, n = 8) or AAV-hSyn-cre (Hdac4KO, n = 7) injected Hdac4flox/flox mice before (BS) and after sleep deprivation (SD). q-t, Hourly NREMS time (q) or delta power density (s) and accumulated NREMS time (r) or delta power change (t) of AAV-hSyn-mCherry (n = 9), AAV-hSyn-HDAC4WT (n = 10) or AAV-hSyn-HDAC4CN (n = 7) injected C57BL/6J mice before (BS) and after sleep deprivation (SD). Data are mean ± s.e.m. Two-tailed unpaired t-test (b, f, j and p); two-way ANOVA with Sidak’s test (d, h and l); mixed-effects model (m and q); and one-way ANOVA with Dunnett’s test (t). See Supplementary Fig. 1 for gel source data. n refers to the number of biological replicates.
Extended Data Fig. 6 HDAC4/5 regulates NREMS amount in posterior hypothalamus without altering circadian clock.
a—d, Hourly NREMS time (a), EEG power spectra analysis (b), and REMS (c) or Wake (d) amount of C57BL/6J mice injected with AAV-hSyn-mCherry (n = 6), AAV-hSyn-HDAC4CN (n = 7) or AAV-hSyn-H4N/H5N-VP16 (n = 7) in posterior hypothalamus. e and f, Schematic and representative images showing HDAC4(CN) expression in SCN following stereotactic injection of AAV-hSyn-HDAC4CN in POA (n = 6) or posterior hypothalamus (n = 7). Scale bars, 1 mm. g, Representative ex vivo SCN slice bioluminescence recording of AAV-hSyn-mCherry, AAV-hSyn-HDAC4CN, or AAV-hSyn-H4N/H5N-VP16VP16 injected Per2::Luc mice (n = 3), and the short period control Cry1−/−;Per2::Luc mice (n = 1). Circadian time 0 was set as 24 h post medium change. h, Quantitative analysis of circadian periods in (g). Solid line represents the mean of SCN bioluminescence recording of AAV-hSyn-mCherry injected Per2::Luc mice, whereas dashed lines represent cut-offs of abnormality (≥ three standard deviations). i, RNAi knockdown of all four genes caused no clock phenotype in Bmal1:dLuc U2OS cells. Data were re-analyzed from a previous study30. siGL2 serves as the negative and siCRY2 as the positive controls for period changes. Circadian time 0 was set as 24 h post-medium change. j, Quantitative analyses of each knockdown result in (i) using Multicycle software. The solid line represents the mean of negative control, and dash lines indicate the cut-offs of abnormality (≥ three standard deviations). Each dot represents one experiment and duplicate experiments are applied for each gene. Data are mean ± s.e.m. Mixed-effects model (a); two-way ANOVA with Dunnett’s test (c and d); and one-way ANOVA with Dunnett’s test (h and j). n refers to the number of biological replicates.
Extended Data Fig. 7 HDAC4(CN) regulates NREMS amount in a CREB-dependent manner.
a, Schematics of wild-type and mutant MEF2A, MEF2C,MEF2D, FOXO1 and FOXO3. b-e, Hourly NREMS time (b), EEG power spectra analysis (c), and REMS (d) or Wake (e) amount of AAV-hSyn-eGFP (n = 21), AAV-hSyn-CREB (n = 14), or AAV-hSyn-A-CREB (n = 21) injected C57BL/6J mice. f, Representative images showing co-immunostaining of A-CREB-T7 (purple), HDAC4(CN)-HA (red) and NeuN (green) of cortical and thalamic sections from C57BL/6J mice co-injected with AAV-hSyn-HDAC4CN and AAV-hSyn-A-CREB. Scale bars, 20 μm. g, Quantification of the percentage of HDAC4(CN)+ neurons that co-express A-CREB and vice versa in the cortex and thalamus in (f) (n = 6). h-k, Hourly NREMS time (h), EEG power spectra analysis (i), and REMS (j) or Wake (k) amount of C57BL/6J mice injected with AAV-hSyn-mCherry (n = 6), AAV-hSyn-HDAC4CN and AAV-hSyn-mCherry (HD4(CN)+mCherry, n = 7), or AAV-hSyn-HDAC4CN and AAV-hSyn-A-CREB (HD4(CN)+A-CREB, n = 6). Data are mean ± s.e.m. Mixed-effects model (b and h); and two-way ANOVA with Dunnett’s test (d-e and j-k). n refers to the number of biological replicates.
Extended Data Fig. 8 HDAC4 and CREB coordinately regulate sleep downstream of SIK3 kinase.
a, Immunoblotting of brain lysates from AAV-hSyn-eGFP injected wild-type (WT+eGFP) mice and AAV-hSyn-eGFP (SLP+eGFP), AAV-hSyn-CREB (SLP+CREB), or AAV-hSyn-HDAC4CN (SLP+HD4(CN)) injected Sik3E13∆/+ mice. b, Daily sleep time in WT+eGFP (n = 10), SLP+eGFP (n = 9), SLP+CREB (n = 9), or SLP+HD4(CN) (n = 8) mice. c-f, Hourly NREMS time (c), EEG power spectra analysis (d), and REMS (e) or Wake (f) amount of WT+eGFP (n = 10), SLP+eGFP (n = 9), SLP+CREB (n = 6), or SLP+HD4(CN) (n = 8) mice. g, Immunoblotting of brain lysates from AAV-hSyn-eGFP (SLP+eGFP), AAV-hSyn-HDAC4WT (SLP+HD4(WT)), or AAV-hSyn-HDAC4S245A (SLP+HD4(S245A)) injected Sik3E13∆/+ mice (two independent experiments with similar results). h-m, Hourly NREMS time (h) and delta power density (i), EEG power spectra analysis (j), and NREMS (k), REMS (l), or Wake (m) amount of SLP+eGFP (n = 12), SLP+HD4(WT) (n = 8), or SLP+HD4(S245A) (n = 8) mice. n, Immunoblotting of brain lysates from AAV-hSyn-eGFP (SLP+eGFP), AAV-hSyn-HDAC5WT (SLP+HD5(WT)), or AAV-hSyn-HDAC5CN (SLP+HD5(CN)) injected Sik3E13∆/+ mice (two independent experiments with similar results). o-t, Hourly NREMS time (o) and delta power density (p), EEG power spectra analysis (q), and NREMS (r), REMS (s) or Wake (t) amount of SLP+eGFP (n = 8), SLP+HD5(WT) (n = 8), or SLP+HD5(CN) (n = 9) mice. Data are mean ± s.e.m. One-way ANOVA with Dunnett’s test (b, d and j); mixed-effects model (c, h, i and o); and two-way ANOVA with Dunnett’s test (e-f, k-m and r-t). See Supplementary Fig. 1 for gel source data. n refers to the number of biological replicates.
Extended Data Fig. 9 HDAC4 and CREB specifically rescue transcriptomic changes in the cortex and hypothalamus of Sik3E13∆/+ mice.
a, Venn diagram showing the common and unique sets of DEGs among the cortex (CTX), thalamus (THA) and hypothalamus (HPT) of SLP+eGFP mice. b, Venn diagrams showing the numbers of unique and overlapped DEGs among the SLP+eGFP, SLP+CREB and SLP+HDAC4(CN) mouse cortex, thalamus and hypothalamus, respectively. To identify DEGs, we compare the transcriptomes of WT+eGFP vs SLP+eGFP samples as well as the transcriptomes of SLP+eGFP vs SLP+CREB or SLP+HDAC4(CN) samples (n = 4 per group). c, A table listing overlapped DEGs in the cortex, thalamus and hypothalamus, respectively, among the SLP+eGFP (normalized to WT+eGFP) and SLP+CREB and SLP+HDAC4(CN) (normalized to SLP+eGFP) brain samples.
Extended Data Fig. 10 HDAC4 and CREB function cooperatively in transcriptional regulation.
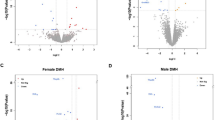
a, Volcano plots of RNA-seq data from ABC-HDAC4(CN) or ABC-CREB mouse brain tissues (n = 4 for thalamus or hypothalamus, n = 3 for cortex) by binomial test via edgeR. The horizontal dashed line shows the two-sided unadjusted P = 0.05 as cutoff. The vertical dashed lines show +/-0.38-log2 fold changes as cutoff. Orange and blue dots indicate up- or down-regulated DEGs. b, RNA-seq heatmaps showing DEGs in HDAC4(CN) brain samples that are documented as target genes of MEF2, RUNX2, FOXO, or SRF, respectively. c, Percentage of CREB and/or HDAC4(CN)-bound genes (by ChIP-seq) in the DEGs of ABC-HDAC4(CN), ABC-CREB or both brain samples (by RNA-seq). d, Venn diagrams showing the numbers of unique and overlapped DEGs by comparing the RNA-seq and ChIP-seq results. e, Circadian oscillation of the overlapped DEGs (by RNA-seq) or HDAC4(CN)/CREB-bound genes (by ChIP-seq) based on previous circadian transcriptome analysis of the synaptosome or forebrain47. f, A model for transcriptional regulation of sleep amount in mice.
Supplementary information
Supplementary Fig. 1
The unprocessed Immunoblots associated with the data presented in main figures and extended data figures.
Supplementary Table 1
The DEGs of ABC-HDAC4CN or ABC-CREB mouse brain tissues (relative to the corresponding ABC-mCherry mouse brain tissues). Linked to Fig. 6.
Supplementary Table 2
The overlapping DEGs in the cortex (CTX), hypothalamus (HPT) and thalamus (THA) of ABC-HDAC4CN and ABC-CREB mouse brains (relative to the corresponding ABC-mCherry mouse brain tissues). Linked to Fig. 6.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, R., Wang, G., Li, Q. et al. A signalling pathway for transcriptional regulation of sleep amount in mice. Nature 612, 519–527 (2022). https://doi.org/10.1038/s41586-022-05510-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05510-6
This article is cited by
-
40 Hz light flickering promotes sleep through cortical adenosine signaling
Cell Research (2024)
-
Curcumin regulates autophagy through SIRT3-SOD2-ROS signaling pathway to improve quadriceps femoris muscle atrophy in KOA rat model
Scientific Reports (2024)
-
Knockdown of SIK3 in the CA1 Region can Reduce Seizure Susceptibility in Mice by Inhibiting Decreases in GABAAR α1 Expression
Molecular Neurobiology (2024)
-
SIK3-HDAC4 signaling pathway: the switch for transition between sleep and wakefulness
Molecular Biomedicine (2023)
-
Biochemical pathways of sleep
Cell Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.