Abstract
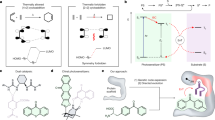
Naturally evolved enzymes, despite their astonishingly large variety and functional diversity, operate predominantly through thermochemical activation. Integrating prominent photocatalysis modes into proteins, such as triplet energy transfer, could create artificial photoenzymes that expand the scope of natural biocatalysis1,2,3. Here, we exploit genetically reprogrammed, chemically evolved photoenzymes embedded with a synthetic triplet photosensitizer that are capable of excited-state enantio-induction4,5,6. Structural optimization through four rounds of directed evolution afforded proficient variants for the enantioselective intramolecular [2+2]-photocycloaddition of indole derivatives with good substrate generality and excellent enantioselectivities (up to 99% enantiomeric excess). A crystal structure of the photoenzyme–substrate complex elucidated the non-covalent interactions that mediate the reaction stereochemistry. This study expands the energy transfer reactivity7,8,9,10 of artificial triplet photoenzymes in a supramolecular protein cavity and unlocks an integrated approach to valuable enantioselective photochemical synthesis that is not accessible with either the synthetic or the biological world alone.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available in the main text or the Supplementary Information. The crystal structure data of TPe3.0 and TPe3.0 in complex with substrate 1b have been deposited in the Protein Data Bank under accession numbers 7XUP and 7XUQ. Source data are provided with this paper.
References
Arnold, F. H. Innovation by evolution: bringing new chemistry to life (Nobel Lecture). Angew. Chem. Int. Ed. 58, 14420–14426 (2019).
Lovelock, S. L. et al. The road to fully programmable protein catalysis. Nature 606, 49–58 (2022).
Chen, K. & Arnold, F. H. Engineering new catalytic activities in enzymes. Nat. Catal. 3, 203–213 (2020).
Silvi, M. & Melchiorre, P. Enhancing the potential of enantioselective organocatalysis with light. Nature 554, 41–49 (2018).
Brimioulle, R., Lenhart, D., Maturi, M. M. & Bach, T. Enantioselective catalysis of photochemical reactions. Angew. Chem. Int. Ed. 54, 3872–3890 (2015).
Genzink, M. J., Kidd, J. B., Swords, W. B. & Yoon, T. P. Chiral photocatalyst structures in asymmetric photochemical synthesis. Chem. Rev. 122, 1654–1716 (2022).
Strieth-Kalthoff, F., James, M. J., Teders, M., Pitzer, L. & Glorius, F. Energy transfer catalysis mediated by visible light: principles, applications, directions. Chem. Soc. Rev. 47, 7190–7202 (2018).
Großkopf, J., Kratz, T., Rigotti, T. & Bach, T. Enantioselective photochemical reactions enabled by triplet energy transfer. Chem. Rev. 122, 1626–1653 (2022).
Zhou, Q.-Q., Zou, Y.-Q., Lu, L.-Q. & Xiao, W.-J. Visible-light-induced organic photochemical reactions through energy-transfer pathways. Angew. Chem. Int. Ed. 58, 1586–1604 (2019).
Strieth-Kalthoff, F. & Glorius, F. Triplet energy transfer photocatalysis: unlocking the next level. Chem 6, 1888–1903 (2020).
Poplata, S., Tröster, A., Zou, Y.-Q. & Bach, T. Recent advances in the synthesis of cyclobutanes by olefin [2+2] photocycloaddition reactions. Chem. Rev. 116, 9748–9815 (2016).
Münster, N., Parker, N. A., van Dijk, L., Paton, R. S. & Smith, M. D. Visible light photocatalysis of 6π heterocyclization. Angew. Chem. Int. Ed. 56, 9468–9472 (2017).
Becker, M. R., Wearing, E. R. & Schindler, C. S. Synthesis of azetidines via visible-light-mediated intermolecular [2+2] photocycloadditions. Nat. Chem. 12, 898–905 (2020).
Candish, L. et al. Photocatalysis in the life science industry. Chem. Rev. 122, 2907–2980 (2022).
Bach, T. & Hehn, J. Photochemical reactions as key steps in natural product synthesis. Angew. Chem. Int. Ed. 50, 1000–1045 (2011).
Kleinmans, R. et al. Intermolecular [2π+2σ]-photocycloaddition enabled by triplet energy transfer. Nature 606, 477–482 (2022).
Huang, M., Zhang, L., Pan, T. & Luo, S. Deracemization through photochemical E/Z isomerization of enamines. Science 375, 869–874 (2022).
Müller, C., Bauer, A. & Bach, T. Light-driven enantioselective organocatalysis. Angew. Chem. Int. Ed. 48, 6640–6642 (2009).
Li, X., Großkopf, J., Jandl, C. & Bach, T. Enantioselective, visible light mediated aza Paternò–Büchi reactions of quinoxalinones. Angew. Chem. Int. Ed. 60, 2684–2688 (2021).
Alonso, R. & Bach, T. A chiral thioxanthone as organocatalyst for enantioselective [2+2] photocycloaddition reactions induced by visible light. Angew. Chem. Int. Ed. 53, 4368–4371 (2014).
Hölzl-Hobmeier, A. et al. Catalytic deracemization of chiral allenes by sensitized excitation with visible light. Nature 564, 240–243 (2018).
Plaza, M., Großkopf, J., Breitenlechner, S., Bannwarth, C. & Bach, T. Photochemical deracemization of primary allene amides by triplet energy transfer: a combined synthetic and theoretical study. J. Am. Chem. Soc. 143, 11209–11217 (2021).
Skubi, K. L. et al. Enantioselective excited-state photoreactions controlled by a chiral hydrogen-bonding iridium sensitizer. J. Am. Chem. Soc. 139, 17186–17192 (2017).
Brenninger, C., Jolliffe, J. D. & Bach, T. Chromophore activation of α,β-unsaturated carbonyl compounds and its application to enantioselective photochemical reactions. Angew. Chem. Int. Ed. 57, 14338–14349 (2018).
Skubi, K. L., Blum, T. R. & Yoon, T. P. Dual catalysis strategies in photochemical synthesis. Chem. Rev. 116, 10035–10074 (2016).
Pecho, F. et al. Enantioselective [2+2] photocycloaddition via iminium ions: catalysis by a sensitizing chiral Brønsted acid. J. Am. Chem. Soc. 143, 9350–9354 (2021).
Blum, T. R., Miller, Z. D., Bates, D. M., Guzei, I. A. & Yoon, T. P. Enantioselective photochemistry through Lewis acid catalyzed triplet energy transfer. Science 354, 1391–1395 (2016).
Chapman, S. J. et al. Cooperative stereoinduction in asymmetric photocatalysis. J. Am. Chem. Soc. 144, 4206–4213 (2022).
Black, M. J. et al. Asymmetric redox-neutral radical cyclization catalysed by flavin-dependent ‘ene’-reductases. Nat. Chem. 12, 71–75 (2020).
Gao, X., Turek-Herman, J. R., Choi, Y. J., Cohen, R. D. & Hyster, T. K. Photoenzymatic synthesis of α-tertiary amines by engineered flavin-dependent “ene”-reductases. J. Am. Chem. Soc. 143, 19643–19647 (2021).
Biegasiewicz, K. F. et al. Photoexcitation of flavoenzymes enables a stereoselective radical cyclization. Science 364, 1166–1169 (2019).
Emmanuel, M. A., Greenberg, N. R., Oblinsky, D. G. & Hyster, T. K. Accessing non-natural reactivity by irradiating nicotinamide-dependent enzymes with light. Nature 540, 414–417 (2016).
Huang, X. et al. Photoenzymatic enantioselective intermolecular radical hydroalkylation. Nature 584, 69–74 (2020).
Harrison, W., Huang, X. & Zhao, H. Photobiocatalysis for abiological transformations. Acc. Chem. Res. 55, 1087–1096 (2022).
Burke, A. J. et al. Design and evolution of an enzyme with a non-canonical organocatalytic mechanism. Nature 570, 219–223 (2019).
Liu, X. et al. Genetically encoded photosensitizer protein facilitates the rational design of a miniature photocatalytic CO2-reducing enzyme. Nat. Chem. 10, 1201–1206 (2018).
Kang, F. et al. Rational design of a miniature photocatalytic CO2-reducing enzyme. ACS Catal. 11, 5628–5635 (2021).
Fu, Y. et al. Biocatalytic cross-coupling of aryl halides with a genetically engineered photosensitizer artificial dehalogenase. J. Am. Chem. Soc. 143, 617–622 (2021).
Madoori, P. K., Agustiandari, H., Driessen, A. J. M. & Thunnissen, A. W. H. Structure of the transcriptional regulator LmrR and its mechanism of multidrug recognition. EMBO J. 28, 156–166 (2009).
Roelfes, G. LmrR: a privileged scaffold for artificial metalloenzymes. Acc. Chem. Res. 52, 545–556 (2019).
Zhu, M., Zheng, C., Zhang, X. & You, S.-L. Synthesis of cyclobutane-fused angular tetracyclic spiroindolines via visible-light-promoted intramolecular dearomatization of indole derivatives. J. Am. Chem. Soc. 141, 2636–2644 (2019).
Xu, J. et al. Stereodivergent protein engineering of a lipase to access all possible stereoisomers of chiral esters with two stereocenters. J. Am. Chem. Soc. 141, 7934–7945 (2019).
Cozzi, F., Ponzini, F., Annuziata, R., Cinquini, M. & Siegel, J. S. Polar interactions between stacked π systems in fluorinated 1,8-diarylnaphthalenes: importance of quadrupole moments in molecular recognition. Angew. Chem. Int. Ed. 34, 1019 (1995).
Rui, J. et al. Directed evolution of nonheme iron enzymes to access abiological radical-relay C(sp3)−H azidation. Science 376, 869–874 (2022).
Drienovská, I. et al. A designer enzyme for hydrazone and oxime formation featuring an unnatural catalytic aniline residue. Nat. Chem. 10, 946–952 (2018).
Acknowledgements
We thank the National Key R&D Program of China (no. 2018YFA0903500), the National Natural Science Foundation of China (no. 22077042, 22107075) and the Natural Science Foundation of Top Talent of SZTU (20211061010013) for financial support. We thank the Analytical and Testing Centre of HUST, Analytical and Testing Centre of School of Chemistry and Chemical Engineering (HUST) and Research Core Facilities for Life Science (HUST) for instrument support. We thank X. Wan at the Shanghai Institute of Organic Chemistry for providing vibrational circular dichroism analysis, and the staff at beamlines BL02U1 and BL18U1 of the Shanghai Synchrotron Radiation Facility (SSRF) for assistance during X-ray crystal data collection. We thank T. Bach at Technical University of Munich and S. Xie at HUST for valuable discussions.
Author information
Authors and Affiliations
Contributions
Y.W. and F.Z. conceived the project and designed the experiments. N.S. and J.H. performed the experiments and interpreted the data. J.Q. and X.C. performed the crystallography study and interpreted the data. T.Z. and R.L. carried out the computational studies. J.G., L.T., W. Zhang and Y.D. assisted with the molecular biology experiments. G.W. assisted with the substrate synthesis. W. Zhao performed protein mass analysis. Y.W. and F.Z. wrote the manuscript with input from all of the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Docking of substrate 1a interacting with the LmrR dimer using AutoDock4.
The blue sticks show substrate 1a. The yellow balls and sticks show the W96 residue. The yellow dashes show the π-π interaction between W96 and the indole moiety of 1a. The red balls show the residues pointed to the inner pocket and surrounding W96, which are reserved in the first round of evolution. The green balls show the residues pointed to the inner pocket and with close spatial distances (<10 Å) to the C(2)-C(3) double bond of indole 1a, which were screened for BpA insertion.
Extended Data Fig. 2 The crystal structures of TPe3.0 and TPe3.0 in complex with substrate 1b.
a. The crystal structure of TPe3.0 cocrystalized with 1b (PDB code: 7XUQ). Two molecules forming a dimer are presented. The backbone is shown as grey cartoon. The BpA is shown as sticks with carbon atoms coloured in light blue. 1b is shown as sticks with carbon atoms coloured in orange. Oxygen and nitrogen atoms are shown in red and blue, respectively. The yellow dashes show the π-π interactions between BpA and the substrates with the distances (Å) labelled. b. The crystal structure of TPe3.0 (PDB code: 7XUP) in a monomeric form. c. Superimposition of the structure of TPe3.0 and the structure of TPe3.0 in complex with substrate 1b. TPe3.0 is shown as pink cartoon while TPe3.0 in complex with 1b is shown as grey cartoon. Interacting residues V15, L18, M89, A92, BpA and L96 are shown as sticks with carbon atoms coloured in light pink and grey respectively. Oxygen, nitrogen and sulfur atoms are shown in red, blue and yellow, respectively.
Extended Data Fig. 3 The reaction time course of photocycloaddition of 1b catalysed by TPe4.0_FBpA under different light intensity irradiation.
Light intensity is (A) 162 mW/cm2 and (B) 3.8 mW/cm2. Error bars denote the standard deviation from triplicate measurements, and they are not shown when smaller than the data point marker.
Supplementary information
Supplementary Information
Supplementary Sections 1–19, Figs. 1–19, Tables 1–10 and NMR spectra data.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, N., Huang, J., Qian, J. et al. Enantioselective [2+2]-cycloadditions with triplet photoenzymes. Nature 611, 715–720 (2022). https://doi.org/10.1038/s41586-022-05342-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05342-4
This article is cited by
-
Accessing ladder-shape azetidine-fused indoline pentacycles through intermolecular regiodivergent aza-Paternò–Büchi reactions
Nature Communications (2024)
-
Engineered enzymes for the synthesis of pharmaceuticals and other high-value products
Nature Synthesis (2024)
-
A light-driven enzymatic enantioselective radical acylation
Nature (2024)
-
A non-canonical nucleophile unlocks a new mechanistic pathway in a designed enzyme
Nature Communications (2024)
-
Remote stereocontrol with azaarenes via enzymatic hydrogen atom transfer
Nature Chemistry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.