Abstract
Although melanoma is notorious for its high degree of heterogeneity and plasticity1,2, the origin and magnitude of cell-state diversity remains poorly understood. Equally, it is unclear whether growth and metastatic dissemination are supported by overlapping or distinct melanoma subpopulations. Here, by combining mouse genetics, single-cell and spatial transcriptomics, lineage tracing and quantitative modelling, we provide evidence of a hierarchical model of tumour growth that mirrors the cellular and molecular logic underlying the cell-fate specification and differentiation of the embryonic neural crest. We show that tumorigenic competence is associated with a spatially localized perivascular niche, a phenotype acquired through an intercellular communication pathway established by endothelial cells. Consistent with a model in which only a fraction of cells are fated to fuel growth, temporal single-cell tracing of a population of melanoma cells with a mesenchymal-like state revealed that these cells do not contribute to primary tumour growth but, instead, constitute a pool of metastatic initiating cells that switch cell identity while disseminating to secondary organs. Our data provide a spatially and temporally resolved map of the diversity and trajectories of melanoma cell states and suggest that the ability to support growth and metastasis are limited to distinct pools of cells. The observation that these phenotypic competencies can be dynamically acquired after exposure to specific niche signals warrant the development of therapeutic strategies that interfere with the cancer cell reprogramming activity of such microenvironmental cues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw data of scRNA-seq (and output files from CellRanger) and spatial transcriptomics (Visium and stereo-seq) are deposited at the GEO under accession number GSE207592. Processed count matrices and cell annotations (scRNA-seq and Visium) are available online (https://marinelab.sites.vib.be/en). Molecular cartography data are available at Zenodo (https://zenodo.org/record/6856193#.Ytj-fnZBz-g). All other data supporting the findings of this study are available on reasonable request. Source data are provided with this paper.
Code availability
Scripts and annotation files for the study have been deposited at GitHub (https://github.com/MarineLab/Karras-et-al). The raw list of clone sizes and scripts to extract the model parameters, as well as running the stochastic simulations of the two-compartment model, are available at GitHub (https://github.com/ibordeu/scripts_Karras_et_al_2022_git). A brief description of how to reproduce the analysis workflows and the figures presented in this paper is included there.
Change history
19 October 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41586-022-05415-4
References
Rambow, F., Marine, J. C. & Goding, C. R. Melanoma plasticity and phenotypic diversity: therapeutic barriers and opportunities. Genes Dev. 33, 1295–1318 (2019).
Arozarena, I. & Wellbrock, C. Phenotype plasticity as enabler of melanoma progression and therapy resistance. Nat. Rev. Cancer 19, 377–391 (2019).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Rambow, F. et al. Toward minimal residual disease-directed therapy in melanoma. Cell 174, 843–855 (2018).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Wouters, J. et al. Robust gene expression programs underlie recurrent cell states and phenotype switching in melanoma. Nat. Cell Biol. 22, 986–998 (2020).
Patton, E. E. et al. Melanoma models for the next generation of therapies. Cancer Cell 39, 610–631 (2021).
Ackermann, J. et al. Metastasizing melanoma formation caused by expression of activated N-RasQ61K on an INK4a-deficient background. Cancer Res. 65, 4005–4011 (2005).
Serrano, M. et al. Role of the INK4a locus in tumor suppression and cell mortality. Cell 85, 27–37 (1996).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997 (2018).
Rambow, F. et al. New functional signatures for understanding melanoma biology from tumor cell lineage-specific analysis. Cell Rep. 13, 840–853 (2015).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013 (2018).
Fan, J. et al. Linking transcriptional and genetic tumor heterogeneity through allele analysis of single-cell RNA-seq data. Genome Research 28, 1217–1227 (2018).
Goding, C. R. & Arnheiter, H. MITF—the first 25 years. Genes Dev. 33, 983–1007 (2019).
Hoek, K. S. & Goding, C. R. Cancer stem cells versus phenotype-switching in melanoma. Pigment Cell Melanoma Res. 23, 746–759 (2010).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Soldatov, R. et al. Spatiotemporal structure of cell fate decisions in murine neural crest. Science 364, eaas9536 (2019).
Kerosuo, L. & Bronner, M. E. cMyc regulates the size of the premigratory neural crest stem cell pool. Cell Rep. 17, 2648–2659 (2016).
Tsoi, J. et al. Multi-stage differentiation defines melanoma subtypes with differential vulnerability to drug-induced iron-dependent oxidative stress. Cancer Cell 33, 890–904 (2018).
Köhler, C. et al. Mouse cutaneous melanoma induced by mutant BRaf arises from expansion and dedifferentiation of mature pigmented melanocytes. Cell Stem Cell 21, 679–693 (2017).
Pozniak, J. et al. A TCF4/BRD4-dependent regulatory network confers cross-resistance to targeted and immune checkpoint therapy in melanoma. Preprint at bioRxiv https://doi.org/10.1101/2022.08.11.502598 (2022).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Reeves, M. Q., Kandyba, E., Harris, S., Del Rosario, R. & Balmain, A. Multicolour lineage tracing reveals clonal dynamics of squamous carcinoma evolution from initiation to metastasis. Nat. Cell Biol. 20, 699–709 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Calabrese, C. et al. A perivascular niche for brain tumor stem cells. Cancer Cell 11, 69–82 (2007).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Wei, K. et al. Notch signalling drives synovial fibroblast identity and arthritis pathology. Nature 582, 259–264 (2020).
Takano, S. et al. Prrx1 isoform switching regulates pancreatic cancer invasion and metastatic colonization. Genes Dev. 30, 233–247 (2016).
Ocaña, O. H. et al. Metastatic colonization requires the repression of the epithelial-mesenchymal transition inducer Prrx1. Cancer Cell 22, 709–724 (2012).
Hoek, K. S. et al. In vivo switching of human melanoma cells between proliferative and invasive states. Cancer Res. 68, 650–656 (2008).
Verfaillie, A. et al. Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state. Nat. Commun. https://doi.org/10.1038/ncomms7683 (2015).
Widmer, D. S. et al. Systematic classification of melanoma cells by phenotype-specific gene expression mapping. Pigment Cell Melanoma Res. 25, 343–353 (2012).
Kawanami, A., Matsushita, T., Chan, Y. Y. & Murakami, S. Mice expressing GFP and CreER in osteochondro progenitor cells in the periosteum. Biochem. Biophys. Res. Commun. 386, 477–482 (2009).
Boiko, A. D. et al. Human melanoma-initiating cells express neural crest nerve growth factor receptor CD271. Nature 466, 133–137 (2010).
Roesch, A. et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell 141, 583–594 (2010).
Schatton, T. et al. Identification of cells initiating human melanomas. Nature 451, 345–349 (2008).
Quintana, E. et al. Efficient tumour formation by single human melanoma cells. Nature 456, 593–598 (2008).
Stemmler, M. P., Eccles, R. L., Brabletz, S. & Brabletz, T. Non-redundant functions of EMT transcription factors. Nat. Cell Biol. 21, 102–112 (2019).
Bosenberg, M. et al. Characterization of melanocyte-specific inducible Cre recombinase transgenic mice. Genesis 44, 262–267 (2006).
Krimpenfort, P., Quon, K. C., Mooi, W. J., Loonstra, A. & Berns, A. Loss of p16Ink4a confers susceptibility to metastatic melanoma in mice. Nature 413, 83–86 (2001).
Dankort, D. et al. BrafV600E cooperates with Pten loss to induce metastatic melanoma. Nat. Genet. 41, 544–552 (2009).
Maria Bosisio, F. et al. Functional heterogeneity of lymphocytic patterns in primary melanoma dissected through single-cell multiplexing. eLife https://doi.org/10.7554/eLife.53008 (2020).
Susaki, E. A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Yates, A. D. et al. Ensembl 2020. Nucleic Acids Res. 48, D682–D688 (2020).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Gans, J. D. & Wolinsky, M. Improved assay-dependent searching of nucleic acid sequence databases. Nucleic Acids Res. 36, e74 (2008).
Rodriguez, J. M. et al. APPRIS 2017: principal isoforms for multiple gene sets. Nucleic Acids Res. 46, D213–D217 (2018).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. Cell detection with star-convex polygons. In Proc. Medical Image Computing and Computer Assisted Intervention—MICCAI 2018 (eds Frangi, A. et al.) Vol. 11071, 265–273 (Springer, 2018).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Rousseeuw, P. J. Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Oren, Y. et al. Cycling cancer persister cells arise from lineages with distinct programs. Nature 596, 576–582 (2021).
Guzmán, C., Bagga, M., Kaur, A., Westermarck, J. & Abankwa, D. ColonyArea: an ImageJ plugin to automatically quantify colony formation in clonogenic assays. PLoS ONE 9, e92444 (2014).
Acknowledgements
We thank G. Ghanem for providing us with the MM lines; H. Clevers for providing us with the Confetti-reporter mouse strain; and O. Van Goethem and V. Benne for their assistance with the mouse experiments; the members of the VIB Technology Watch members, M. Bontinck and Y.-C. Wang, for facilitating collaborations with Resolve Biosciences and BGI research, respectively; and Y. Oren and the laboratories of A. Regev and J. Brugge for providing the watermelon vector. P.K. received financial support from an Marie Curie Individual Fellowship (H2020‐MSCA‐IF‐2018, 841092), FWO (1210520N) and Stichting tegen Kanker (Foundation against Cancer, 2021-028). J.P. received financial support from a Marie Curie Individual Fellowship (H2020‐MSCA‐IF‐2019, 896897). A.N. received postdoctoral from the KU Leuven (PDMT1/21/035). C.P. received PhD research fellowships from FWO (11M3822N) and of the Boehringer Ingelheim Fonds. J.W. received a postdoctoral research fellowship from Stichting Tegen Kanker (Foundation against Cancer; 2019-100). O.M.-B. was supported by the 12T1217N project by the FWO at the program under the Marie Skłodowska-Curie grant agreement no. 665501. D.P. and N.V.D. received PhD fellowships from the VIB PhD international programme and FWO-SB 1S79619N, respectively. F.R. acknowledges support from the Alexander von Humboldt Foundation. The computational resources and services used in this work were provided by the VSC (Flemish Supercomputer Centre), funded by the Research Foundation–Flanders (FWO) and the Flemish Government, department EWI. B.D.S. acknowledges funding from the Royal Society E. P. Abraham Research Professorship (RP\R1\180165), EPSRC (EP/P034616/1) and Wellcome Trust (219478/Z/19/Z). This work was supported by the FWO (G0C530N and G070622N), Stichting Tegen Kanker (FAF-F/2018/1265), Neftkens foundation, Melanoma Research Alliance (MRA, EIA#623591), KU Leuven (C1 grant) and the Belgian Excellence of Science (EOS) programme to J.-C.M.
Author information
Authors and Affiliations
Contributions
P.K. designed and conducted experiments, and acquired, analysed and interpreted the data. P.K., I.B. and B.D.S. acquired, analysed and interpreted data for the mathematical modelling. P.K. generated the mouse models and performed all the in vivo experiments assisted by C.P.; P.K. performed all of the single-cell mouse experiments on the NRAS background, and G. Bervoets and E.L. performed all of the single-cell human biopsies. N.V.R. and E.L. performed the single-cell experiments in the BRAF background. F.R., J.P. and Y.H. analysed the single-cell data from mouse and human lesions and data from TCGA. A.N. conducted in vitro experiments of melanoma–endothelial cell interplay. M.D., D.P. and J.W. conducted and analysed in vitro experiments to assess PRRX1 function in melanoma. P.K. and S.M. conducted spatial transcriptomics experiments, and analyses were performed by F.R. and R.B.; P.K., F.R., J.H.K., H.L. and X.Z. performed the stereo-seq experiments and downstream analyses. O.M.-B. and J.L. provided support for the FACS sorting/analysis. B.P. provided computational support for molecular cartography experiments. O.B., Y.V.H., F.B. and J.v.d.O. provided human samples and pathology support. A.R. provided the Prrx1::creER-IRES2-EGFP allele mice. G. Bergers, Y.S., D.L. and C.B. helped with the interpretation of all data and concepts underlying the perivascular niche and mode of tumour growth. All of the authors read and edited the manuscript. P.K., F.R. and J.-C.M. conceptualized and designed the research study and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Thomas Hofer, Samantha Morris, Frederic de Sauvage and Amaya Viros for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cellular composition and discriminative regulons of NRAS-driven mouse melanoma.
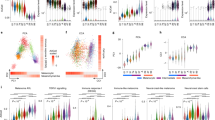
a, UMAP illustrating cell type diversity in NRASQ61K/°;Ink4a−/− lesions (43k cells). b, Activity (AUCell score) of a melanoma specific gene set5 in NRASQ61K/°;Ink4a−/− lesions. c, Violin plot depicting mean Copy Number Variation (CNV) scores for the different cell types, including the malignant compartment, identified in NRASQ61K/°;Ink4a−/− lesions. d, Relative proportions of the melanoma transcriptional cell states highlighting a strong representation across all 5 NRASQ61K/°;Ink4a−/− tumours, illustrated as stacked bar (left panel) and as UMAP (right panel). e, Proliferative (PRO or melanocytic) and invasive (INV or mesenchymal-like) gene sets34 activity (AUCell score) in all mouse malignant cells (umap plots, left panel) or per transcriptional state (violin plots, right panel). f, Heatmap depicts changes in regulon activities for seurat clusters described in NRASQ61K/°;Ink4a−/− lesions g, Heatmap depicting average expression of selected Neural Crest Stem Cell (NCSC) and pre-migratory (pre-EMT) Neural Crest markers.
Extended Data Fig. 2 Single-cell transcriptomic landscape in BRAF-mutant mouse and human melanoma lesions.
a, UMAP visualization of >2.600 malignant cells analysed by scRNA-seq and integrated across 2 different BRAFV600E;Ptenl/l lesions. The Seurat clusters were annotated using signature-based annotation (left panel). Mitf expression and Proliferative (PRO or Melanocytic) and invasive (INV or mesenchymal-like) gene sets35 activity (AUCell score) across all malignant cells (right panels). b, Stacked bar chart illustrating the distribution of malignant cell states (binary cut-offs) extracted from NRASQ61K/°;Ink4a−/− and projected onto scRNA-seq data of drug naive human melanoma lesions. Each graph is categorized based on the main genetic driver mutations identified. WT indicates lesions that do not carry BRAF neither NRAS mutations; N/A indicates lesions for which genetic information is lacking. c, Stacked bar chart illustrating the distribution of malignant cell states (binary cut-offs) extracted from NRASQ61K/°;Ink4a−/− lesions and projected onto a publicly available scRNA-seq dataset (malignant treatment-naïve cells) of human melanoma biopsies11. The patient IDs and main genetic driver mutations are indicated. WT indicates lesions that do not carry BRAF neither NRAS mutations; N/A indicates lesions for which genetic information is lacking; met. stands for metastatic and prim. for primary lesions.
Extended Data Fig. 3 Characterization of the Confetti melanoma mouse model.
a, H&E staining of melanoma model Tyr::NRASQ61K/°;Ink4a−/−;Tyr::CreERT2/°;Rosa26-LSL-Confetti+/+. Representative image from n = 3 independent tumours. b, Confocal images of labelled melanoma subclones acquired by confocal imaging upon TAM administration and immunostained for melanocytic lineage markers MITF, gp100 and MLANA (magenta) exhibiting heterogeneous expression. Images from n = 4 independent tumours. c, Representative confocal images showing Confetti labelling efficiency in melanoma tumours 3 days after the administration of different doses of TAM in order to achieve the optimal dose to perform clonal analyses. Images from n = 4 independent experiments. d, Representative confocal images showing Confetti labelling of melanoma cells 5 days after TAM administration (0.5 mg). Images from n = 6 independent tumours. e, Violin plot depicting the proportion of labelled melanoma cells expressing each Confetti fluorophore 5 days after TAM administration (0.5 mg). Six melanoma tumours (two sectional areas of each) were analysed. Black line represents median and grey lines the 25th to 75th percentile. f, Representative confocal image at low magnification illustrating the clone size heterogeneity in tumours that have expanded 10-fold from the initial size after TAM administration (0.5 mg). Arrowheads show several single and/or oligoclones that exhibit minor expansion over time. Images from n = 8 independent tumours
Extended Data Fig. 4 Clone size distributions are consistent with the hierarchical model.
a, Cluster-cluster distance correlation indicating a high likelihood that clusters of a given colour are found near clusters of the same colour for distances below 100 μm. This threshold, indicated by a dashed line, was then used in the association of clones. Solid line and shaded area correspond to mean and ±SEM for n = 5 biological repeats. b–e, Cumulative distribution (CDF) of clone sizes for YFP+ and RPF+ cells in samples 2-5 (for number of clones per sample see Supplementary Table 5) Sample 1 is presented in Fig. 2f. The dashed black lines correspond to the bi-exponential fits of the data, as predicted by the hierarchical model, for each dataset we show the R-squared (R2) and standard error of the fit (S) of the theoretical CDFs to the data. f, Sensitivity of the stem cell expansion rate, cycling rate and stem cell fraction to different choices of the stem cell symmetric division probability, r. The vertical dotted line indicates the value r = 0.75 used here to estimating the model parameters (see Supplementary Table 5). g–k, Comparison of the empirical CDFs of clone sizes for each of the n = 5 biological repeats and the corresponding distributions and SD obtained from performing 10.000 stochastic simulations of the two-compartment stem-progenitor cell model using the parameters in Supplementary Table 5. The p-values from two-sample Kolmogorov-Smirnov tests comparing the empirical and numerical distribution of clone sizes are shown. In f and i, the RFP channel of sample 3 is not shown as it was considered to be below the threshold of clonality. For Extended Data Fig. 5 see Supplementary Note in Supplementary Information)
Extended Data Fig. 5 Hierarchical melanoma growth is independent of the adaptive immune system.
a, Schematic representation of the generated melanoma mouse model transplanted in immunocompromised mice (Foxn1nu). Schematic made with Biorender.com. b, Tumour growth kinetics (mm3) of individual tumours in immunodeficient mice (yellow), and corrected mean obtained from fitting an exponential growth to each individual sample (black curve). Dashed lines indicate tumour duplication times. Dotted red curve refers to the mean growth kinetics extracted from Fig. 2b. c, Confocal images showing Confetti labelling efficiency in melanoma tumours 3 days after the administration of different doses of TAM in order to achieve the optimal dose to perform clonal analyses. Representative images from n = 3 independent tumours. d, Confocal image showing Confetti labelling of melanoma cells 5 days after TAM administration (0.5 mg) in immunodeficient background. Representative image from n = 5 independent tumours. e, Violin plot depicting the proportion of labelled melanoma cells expressing each Confetti fluorophore 5 days after TAM administration (0.5 mg). Six melanoma tumours (two sectional area of each) were analysed. Black line represents median and grey lines the 25th to 75th percentile. f, Confocal images of melanoma tumours in immunodeficient background exhibiting Confetti labelling 5 days after initial induction (1.5-fold tumour increase) and at relative growth of 10-fold upon single low dose intraperitoneal Tamoxifen administration (0.5 mg). Representative images from n = 3 independent tumours
Extended Data Fig. 6 Spatially mapping melanoma heterogeneity.
a, Spatially resolved quantification of stress-like (hypoxia) state (prediction score) by label transfer of scRNA-seq data onto Visium spots (sample 1,2,3). b, Scatter plot shows a positive correlation (Spearman) between the stress-like (hypoxia) activity score and the distance to the nearest BV (Fisher’s exact test, two-tailed p value). c, Spatially resolved quantification of pre-EMT NC stem-like state (prediction score) by label transfer of scRNA-seq data onto Visium spots (sample 2,3). d, Violin plots and box plots showed the number of Stereo-seq detected genes (left) and Molecular Identifier (MID) (right) per bin50 (25 µm diameter). The middle horizontal line of the boxplot represents the median. e, Gene count visualization of Stereo-seq data. f, ROI coloured by percentage of mitochondrial gene count (bin50). g, Scatter plot shows correlation (Spearman) of stress-like (hypoxia) activity (AUCell score) and distance to closest blood vessel per bin based on Stereo-seq bin50 data (Fisher’s exact test, two-tailed p value). h, UMAP of all segmented cells (170k) from molecular cartography (see Fig. 3f), coloured by Seurat cluster and melanoma (melanocytic state) gene signature activity (AUCell score). i, UMAP of malignant subset (109k) from molecular cartography, coloured by Seurat cluster and melanoma (melanocytic state) gene signature activity (AUCell score). j, Expression of pre-EMT NC stem-like cell markers assessed by scRNA-seq. k, Expression of pre-EMT NC stem-like cell markers assessed by Molecular Cartography. l, Expression of stress-like (hypoxia) cell markers as assessed by scRNA-seq (upper panels) and Molecular Cartography (lower panels). m, Horizontal histogram showing median distance (black line) to the nearest blood Vessel (BV) for all cells annotated as stress-like (hypoxia) or expressing Sox10 and one of the indicated pre-EMT NC stem-like markers. Exact p values are indicated. n, Representative confocal image showing mRNA expression of the indicated genes, as detected by RNAscope, in blood vessel (BV) free and containing areas. Yellow arrowheads indicate the presence of cells positive for both Sox10 (red) and Notch3 (green). Endothelial Cells (Dll4+) are highlighted in orange. RBCs, Red Blood Cells. o, Quantification of m (n = 6; p < 0.0001). Boxes extend from the 25th to 75th percentile. The middle line represents the median. Whiskers represent min to max values. In panels m and o significance was assessed by an unpaired nonparametric, two-tailed Mann-Whitney Test (approximate p value, p < 0.0001)
Extended Data Fig. 7 Mapping the melanoma perivascular niche in human primary melanomas.
a, Confocal image of immunostaining of human primary melanoma lesions for CD31, SOX10, VCAN (upper panel, 3 representative lesions from 5 tumours) or CD31, SOX10, NES (lower panel, 3 representative lesions from 7 tumours). The main driver mutations are indicated. b, Spatial distribution of Confetti clones in respect to vessels. Panel of xy images of a clone (indicated with the red arrowhead) in close proximity to the vessel at indicated z-stack positions (z1-z24). V corresponds to the vessel. c, Violin plot depicting 2D quantification of subclones categorized in two groups based on their size and their spatial distance (μm) from the BV (visualized by AQP1 staining, n = 6; p < 0.0001). Dashed line represents median and grey lines the 25th to 75th percentile. Significance was by an unpaired nonparametric, two-tailed Mann-Whitney Test. d. Cumulative distribution of clone sizes for clones directly in contact with the vasculature (distance < 20 m, solid lines) and clones detached from it (distance ≥ 20 m, dashed lines) for RFP (black circles) and YFP (grey squares) channels, obtained from short-term clonal labelling of (left) 4 days and (right) 10 days post TAM, where the vasculature was segmented after staining with CD31 staining in n = 2 biological repeats for each time-point (shown separately)
Extended Data Fig. 8 Melanoma-Endothelial cell heterotypic interaction promotes growth and induction of the pre-EMT NC stem-like phenotype.
a, Schematic representation of Fig. 3i. b, Kaplan-Meier survival curves from log-rank (Mantel-Cox) test of mice described in a (p = 0.00288, n is indicated in the graph). c, Schematic representation of in vitro assays using the Watermelon system. d, Representative FACS profiles from Fig. 3j and experiment described in c. Percentages of low mcherry (mNeon-positive) populations. e, NRASQ61K/°;Ink4a−/− cells cultured either alone or in the presence of bEnd.3 cells for 48h and processed for scRNA-seq. Left, UMAP by sample identity; middle, expression of the endothelial cell marker Pecam1; and right, pan-melanoma marker Sox10. f, Heatmap showing the overall transcriptional reprogramming effect observed following exposure of NRASQ61K/°;Ink4a−/− melanoma cells to bEnd.3 endothelial cells. g, Percentage of melanoma cells positive for the melanocytic and Neural Crest-like signatures (binary AUCell score). h, Melanospheres of NRASQ61K/°;Ink4a−/− cells grown in the presence of GFP-labelled bEnd.3 ECs for 10 days and the percentage of Nestin-positive cells was correlated with the amount of bEnd.3/GFP-positive cells. Linear regression test was used for statistical significance (n = 19). i, Predicted cell-to-cell interactions based on known ligand-receptor pairs between tumour microenvironment populations (ligands) and the pre-EMT NC stem-like malignant cells (receptors) in scRNA seq data from mouse tumours as inferred by NicheNet. j, Dotplot showing gene expression intensities of selected receptors over the different Seurat clusters. k, Circos plot illustrating Dll4-Notch3 predicted interaction between cell types in scRNA seq data from mouse tumours and inferred by CellChat. The edge width is proportional to the prediction score. Note the predicted interaction of pre-EMT NC stem-like state with ECs. l, UMAP highlighting the specific expression of the ligand Dll4 in ECs and the receptor Notch3 in subpopulation of malignant cells. m, Left panel shows venn diagram highlighting the overlap of the pre-EMT NC stem-like cell state (cluster 4) upregulated genes with NOTCH signalling signature (Reactome). Right panel shows venn diagram highlighting the significant overlap of the pre-EMT NC stem-like cell state (cluster 4) upregulated genes with a Notch3 activation signature31. Hypergeometric distribution test was used for statistical analysis. P values are indicated in each Venn diagram. n, Rbpj expression in the malignant cluster of scRNA data from NRASQ61K/°;Ink4a−/− lesions projected either as UMAP (left panel) or as violin plot (right panel) for each malignant transcriptional state. Schematic in a and c made with Biorender.com
Extended Data Fig. 9 PRRX1 is a master regulator of the mesenchymal-like transcriptional program.
a, Violin plots depict conserved Prrx1 regulon activity scores (AUCell) across mouse Seurat clusters. b, Violin plots depict conserved PRRX1 regulon activity scores (AUCell) across human Seurat clusters. c, PRRX1 expression in n = 375 TCGA_SKCM patients stratified as invasive (n = 123), immune-related (n = 116) and proliferative (n = 136) phenotypes based on their gene expression profile described previously35. Box plots summarize the values per phenotypic group and show the median. Boxes extend from the 25th to 75th percentiles, whiskers and outliers were plotted according to the Tukey method. Significance was assessed by an unpaired nonparametric, two-tailed Mann-Whitney Test (approximate p value, p < 0.0001). d, Heatmap generated using the TCGA database illustrating the gene expression profile of patients with PRRX1 high and low expression (20th percentile cut-off). Genes from the conserved PRRX1 regulon are co-expressed together with PRRX1. e, Left panel, PRRX1 expression (226695_at) in invasive (n = 71) and proliferative (n = 81) melanoma cell lines as previously described36. Error bars represent ±SEM. Significance was assessed by an unpaired nonparametric, two-tailed Mann-Whitney Test (approximate P value, p < 0.0001). Right panel, qPCR analysis for PRRX1 in short term melanoma cells lines categorized as proliferative (n = 11) and invasive (n = 7) based on their gene expression profile). Error bars represent ±SEM. Significance was assessed by unpaired nonparametric two-tailed Student’s t-test. (p = 0.1509). f, PRRX1 expression (upper panel) and signature activity (lower panel) in short term proliferative/melanocytic melanoma cell lines (MM057, MM074 and MM087) transfected with SOX10 siRNA or non-targeting control. Baseline refers to the non-transfected melanoma cell lines. g, Left panel, Matrigel-invasion assay upon silencing of PRRX1 in MM099 cells. Right panel, quantification (n = 3; p = 0.0011). h, Relative mRNA expression of PRRX1 upon siPRRX1 (n = 3; p = 0.0003). In g and h significance was assessed by unpaired two-tailed Student’s t-test
Extended Data Fig. 10 Reprogramming of Prrx1+ melanoma cells during metastatic spreading.
a, Expression of melanocytic lineage markers in malignant FACS-sorted tdTomato+ cells isolated from a primary melanoma lesion of Met-track mice, 2 days (early labelled) post-TAM. b, Expression of Prrx1 and well-established melanoma mesenchymal-like markers in cells described in a. c, Expression of pre-EMT NC stem-like cell markers and activity (AUCell score) of the pre-EMT NC stem-like cell signature in cells described in a. d, Confocal image of a lymph node metastasis 4 weeks post-TAM. Cells positive for both GFP and tdTomato reporters (subcapsular region) as well as tdTomato+ cells expressing lower to undetectable levels of GFP. Black regions correspond to the pigmented melanoma cells. BF, Bright Field. Representative images from 5 different tumours. e, Box plots showing the percentage of tdTomato+/GFPhigh versus and tdTomato+/GFPlow cells in lymph nodes (n = 4 mice). Boxes extend from the 25th to 75th percentile. The middle line represents the median. Whiskers represent min to max values. f, Confocal image of single tdTomato+ (and GFPlow) cells in the liver of a Met-Track mouse 4 weeks post-TAM. Representative image from 2 different tumours. g, Violin plots of Mitf expression and pigmentation genes in FACS-sorted tdTomato+ fraction isolated from lung metastases 2 days (early labelled) and 30-days (late labelled) post-TAM administration
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, Supplementary Note and Supplementary References to support the minimal model of growth. Supplementary Fig. 1 illustrates the gating strategy to analyse the mCherrylow population after co-culture of melanoma and endothelial cells. Supplementary Fig. 2 illustrates the gating strategy to isolate tdTomato+ PRRX1 cells from primary and metastatic sites from Met-track mice after tamoxifen induction. The Supplementary Note describes the theoretical framework used to investigate the clonal dynamics and the experimental observations for the predicted minimal mathematical model of melanoma growth.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Karras, P., Bordeu, I., Pozniak, J. et al. A cellular hierarchy in melanoma uncouples growth and metastasis. Nature 610, 190–198 (2022). https://doi.org/10.1038/s41586-022-05242-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05242-7
This article is cited by
-
Dormancy of cutaneous melanoma
Cancer Cell International (2024)
-
CellCharter reveals spatial cell niches associated with tissue remodeling and cell plasticity
Nature Genetics (2024)
-
Molecular patterns of resistance to immune checkpoint blockade in melanoma
Nature Communications (2024)
-
ZEB1 controls a lineage-specific transcriptional program essential for melanoma cell state transitions
Oncogene (2024)
-
Mapping the single cell spatial immune landscapes of the melanoma microenvironment
Clinical & Experimental Metastasis (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.