Abstract
Central oscillators are primordial neural circuits that generate and control rhythmic movements1,2. Mechanistic understanding of these circuits requires genetic identification of the oscillator neurons and their synaptic connections to enable targeted electrophysiological recording and causal manipulation during behaviours. However, such targeting remains a challenge with mammalian systems. Here we delimit the oscillator circuit that drives rhythmic whisking—a motor action that is central to foraging and active sensing in rodents3,4. We found that the whisking oscillator consists of parvalbumin-expressing inhibitory neurons located in the vibrissa intermediate reticular nucleus (vIRtPV) in the brainstem. vIRtPV neurons receive descending excitatory inputs and form recurrent inhibitory connections among themselves. Silencing vIRtPV neurons eliminated rhythmic whisking and resulted in sustained vibrissae protraction. In vivo recording of opto-tagged vIRtPV neurons in awake mice showed that these cells spike tonically when animals are at rest, and transition to rhythmic bursting at the onset of whisking, suggesting that rhythm generation is probably the result of network dynamics, as opposed to intrinsic cellular properties. Notably, ablating inhibitory synaptic inputs to vIRtPV neurons quenched their rhythmic bursting, impaired the tonic-to-bursting transition and abolished regular whisking. Thus, the whisking oscillator is an all-inhibitory network and recurrent synaptic inhibition has a key role in its rhythmogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data are available from the corresponding authors on reasonable request. Source data are provided with this paper.
Code availability
Custom-made scripts used in this Article are available at GitHub (https://github.com/wanglab-neuro/the-whisking-oscillator-circuit).
References
Marder, E. & Bucher, D. Central pattern generators and the control of rhythmic movements. Curr. Biol. 11, R986–R996 (2001).
Marder, E. & Calabrese, R. L. Principles of rhythmic motor pattern generation. Physiol. Rev. 76, 687–717 (1996).
Vincent, S. B. The Functions of the Vibrissae in the Behavior of the White Rat (Univ. Chicago, 1912).
Welker, W. Analysis of sniffing of the albino rat 1. Behaviour 22, 223–244 (1964).
Moore, J. D. et al. Hierarchy of orofacial rhythms revealed through whisking and breathing. Nature 497, 205–210 (2013).
Moore, J. D., Deschenes, M., Kurnikova, A. & Kleinfeld, D. Activation and measurement of free whisking in the lightly anesthetized rodent. Nat. Protoc. 9, 1792–1802 (2014).
Deschenes, M. et al. Inhibition, not excitation, drives rhythmic whisking. Neuron 90, 374–387 (2016).
Smith, J. C., Ellenberger, H. H., Ballanyi, K., Richter, D. W. & Feldman, J. L. Pre-Botzinger complex: a brainstem region that may generate respiratory rhythm in mammals. Science 254, 726–729 (1991).
Kleinfeld, D., Deschenes, M., Wang, F. & Moore, J. D. More than a rhythm of life: breathing as a binder of orofacial sensation. Nat. Neurosci. 17, 647–651 (2014).
Takatoh, J. et al. Constructing an adult orofacial premotor atlas in Allen mouse CCF. eLife 10, e67291 (2021).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145, dev165753 (2018).
Bellavance, M. A. et al. Parallel inhibitory and excitatory trigemino-facial feedback circuitry for reflexive vibrissa movement. Neuron 95, 673–682 (2017).
Wang, P. et al. Intersectional Cre driver lines generated using split-intein mediated split-Cre reconstitution. Sci. Rep. 2, 497 (2012).
Kato, S. et al. A lentiviral strategy for highly efficient retrograde gene transfer by pseudotyping with fusion envelope glycoprotein. Hum. Gene Ther. 22, 197–206 (2011).
Link, E. et al. Tetanus toxin action: inhibition of neurotransmitter release linked to synaptobrevin proteolysis. Biochem. Biophys. Res. Commun. 189, 1017–1023 (1992).
Schiavo, G. et al. Tetanus and botulinum-B neurotoxins block neurotransmitter release by proteolytic cleavage of synaptobrevin. Nature 359, 832–835 (1992).
Zhang, Y. et al. Identifying local and descending inputs for primary sensory neurons. J. Clin. Invest. 125, 3782–3794 (2015).
Kleinfeld, D., Moore, J. D., Wang, F. & Deschenes, M. The brainstem oscillator for whisking and the case for breathing as the master clock for orofacial motor actions. Cold Spring Harb. Symp. Quant. Biol. 79, 29–39 (2014).
Govorunova, E. G., Sineshchekov, O. A., Janz, R., Liu, X. & Spudich, J. L. Natural light-gated anion channels: a family of microbial rhodopsins for advanced optogenetics. Science 349, 647–650 (2015).
Fenno, L. E. et al. Targeting cells with single vectors using multiple-feature Boolean logic. Nat. Methods 11, 763–772 (2014).
Fenno, L. E. et al. Comprehensive dual- and triple-feature intersectional single-vector delivery of diverse functional payloads to cells of behaving mammals. Neuron 107, 836–853 (2020).
Marshel, J. H. et al. Cortical layer-specific critical dynamics triggering perception. Science 365, eaaw5202 (2019).
Kleinfeld, D. & Sompolinsky, H. Associative neural network model for the generation of temporal patterns. Theory and application to central pattern generators. Biophys. J. 54, 1039–1051 (1988).
Kim, E. J., Jacobs, M. W., Ito-Cole, T. & Callaway, E. M. Improved monosynaptic neural circuit tracing using engineered rabies virus glycoproteins. Cell Rep. 15, 692–699 (2016).
Miyamichi, K. et al. Dissecting local circuits: parvalbumin interneurons underlie broad feedback control of olfactory bulb output. Neuron 80, 1232–1245 (2013).
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007).
Wang, Q. et al. The Allen Mouse Brain Common Coordinate Framework: a 3D reference atlas. Cell 181, 936–953 (2020).
Takatoh, J. et al. New modules are added to vibrissal premotor circuitry with the emergence of exploratory whisking. Neuron 77, 346–360 (2013).
Sreenivasan, V., Karmakar, K., Rijli, F. M. & Petersen, C. C. Parallel pathways from motor and somatosensory cortex for controlling whisker movements in mice. Eur. J. Neurosci. 41, 354–367 (2015).
Beier, K. T. et al. Circuit architecture of VTA dopamine neurons revealed by systematic input-output mapping. Cell 162, 622–634 (2015).
Fortin, G., Jungbluth, S., Lumsden, A. & Champagnat, J. Segmental specification of GABAergic inhibition during development of hindbrain neural networks. Nat. Neurosci. 2, 873–877 (1999).
Friesen, W. O., Poon, M. & Stent, G. S. An oscillatory neuronal circuit generating a locomotory rhythm. Proc. Natl Acad. Sci. USA 73, 3734–3738 (1976).
Grillner, S. Biological pattern generation: the cellular and computational logic of networks in motion. Neuron 52, 751–766 (2006).
Marder, E. & Bucher, D. Understanding circuit dynamics using the stomatogastric nervous system of lobsters and crabs. Annu. Rev. Physiol. 69, 291–316 (2007).
Golomb, D. et al. Theory of hierarchically-organized neuronal oscillator dynamics that mediate rodent rhythmic whisking. Neuron (in the press).
Gross, G. G. et al. An E3-ligase-based method for ablating inhibitory synapses. Nat. Methods 13, 673–678 (2016).
Lin, J. Y., Knutsen, P. M., Muller, A., Kleinfeld, D. & Tsien, R. Y. ReaChR: a red-shifted variant of channelrhodopsin enables deep transcranial optogenetic excitation. Nat. Neurosci. 16, 1499–1508 (2013).
Kiehn, O. Decoding the organization of spinal circuits that control locomotion. Nat. Rev. Neurosci. 17, 224–238 (2016).
Song, J. et al. Multiple rhythm-generating circuits act in tandem with pacemaker properties to control the start and speed of locomotion. Neuron 105, 1048–1061 (2020).
Janczewski, W. A., Tashima, A., Hsu, P., Cui, Y. & Feldman, J. L. Role of inhibition in respiratory pattern generation. J. Neurosci. 33, 5454–5465 (2013).
Del Negro, C. A., Funk, G. D. & Feldman, J. L. Breathing matters. Nat. Rev. Neurosci. 19, 351–367 (2018).
Ausborn, J., Snyder, A. C., Shevtsova, N. A., Rybak, I. A. & Rubin, J. E. State-dependent rhythmogenesis and frequency control in a half-center locomotor CPG. J. Neurophysiol. 119, 96–117 (2018).
Berg, R. W. & Kleinfeld, D. Rhythmic whisking by rat: retraction as well as protraction of the vibrissae is under active muscular control. J. Neurophysiol. 89, 104–117 (2003).
Tervo, D. G. et al. A designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016).
Mattis, J. et al. Principles for applying optogenetic tools derived from direct comparative analysis of microbial opsins. Nat. Methods 9, 159–172 (2011).
Stanek, E. 4th, Rodriguez, E., Zhao, S., Han, B. X. & Wang, F. Supratrigeminal bilaterally projecting neurons maintain basal tone and enable bilateral phasic activation of jaw-closing muscles. J. Neurosci. 36, 7663–7675 (2016).
Rodriguez, E. et al. A craniofacial-specific monosynaptic circuit enables heightened affective pain. Nat. Neurosci. 20, 1734–1743 (2017).
Sakurai, K. et al. Capturing and manipulating activated neuronal ensembles with cane delineates a hypothalamic social-fear circuit. Neuron 92, 739–753 (2016).
Petreanu, L., Mao, T., Sternson, S. M. & Svoboda, K. The subcellular organization of neocortical excitatory connections. Nature 457, 1142–1145 (2009).
Lopes, G. et al. Bonsai: an event-based framework for processing and controlling data streams. Front. Neuroinform. 9, 7 (2015).
Nicovich, P. R. et al. Multimodal cell type correspondence by intersectional mFISH in intact tissues. Preprint at bioRxiv https://doi.org/10.1101/525451 (2019).
Park, Y. G. et al. Protection of tissue physicochemical properties using polyfunctional crosslinkers. Nat. Biotechnol. https://doi.org/10.1038/nbt.4281 (2018).
Claudi, F. et al. Visualizing anatomically registered data with brainrender. eLife 10, e65751 (2021).
Pachitariu, M., Steinmetz, N., Kadir, S., Carandini, M. & Kenneth, D. H. Kilosort: realtime spike-sorting for extracellular electrophysiology with hundreds of channels. Preprint at bioRxiv https://doi.org/10.1101/061481 (2016).
Jun, J. J. et al. Real-time spike sorting platform for high-density extracellular probes with ground-truth validation and drift correction. Preprint at bioRxiv https://doi.org/10.1101/101030 (2017).
Kvitsiani, D. et al. Distinct behavioural and network correlates of two interneuron types in prefrontal cortex. Nature 498, 363–366 (2013).
Berens, P. CircStat: a MATLAB toolbox for circular statistics. J. Stat. Softw. 31, 1–21 (2009).
Clack, N. G. et al. Automated tracking of whiskers in videos of head fixed rodents. PLoS Comput. Biol. 8, e1002591 (2012).
Aljadeff, J., Lansdell, B. J., Fairhall, A. L. & Kleinfeld, D. Analysis of neuronal spike trains, deconstructed. Neuron 91, 221–259 (2016).
Acknowledgements
We thank the members of the Wang laboratory and the aABC U19 BRAIN Circuit Team for discussions and suggestions; B.-X. Han and S. Choi for mouse colony maintenance; and staff at the Boston Children’s Hospital Viral Core for the production of AAV2retro-CAG-Cre. This work is supported by NIH grants U19 NS107466 and R01 NS077986.
Author information
Authors and Affiliations
Contributions
J.T., V.P. and F.W. conceptualized the project, designed experiments and wrote the paper with input from all of the authors. J.T. and V.P. performed the majority of the experiments with help from J.L., P.M.T. and S.L.; L.C. performed slice electrophysiology with crucial support from Z.H.; J.T., V.P., J.L. and P.M.T. analysed data. S.Z. and A.H. produced several viral vectors. D.G. and D.K. contributed to behaviour and electrophysiology data analysis and theoretical issues, and provided help with the intra-aural illumination opto-tagging. F.W. supervised all of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Jan Marino Ramirez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Anatomical and functional characterization of vIRtvGlut2 neurons.
a, Representative image of vibrissa premotor neurons (green) in vIRt traced by a three-step monosynaptic rabies virus tracing method shown in Fig. 1. vGlut2 (red) mRNA were visualized by HCR RNA-FISH. b, Molecular identity of premotor vIRt neurons. 67.0 ± 1.0% (n = 4) and 22.8 ± 1.3% (n = 3) of premotor vIRt neurons are vGat+ and vGlut2+, respectively. 69.6 ± 3.4% (n = 4) of vGat+ premotor vIRt neurons are PV+. c, Strategy for labelling vGlut2+ premotor vIRt neurons (vIRtvGlut2). Retrograde lentivirus carrying FlpO (RG-LV-FlpO) and AAV carrying Cre- and Flp-dependent-EYFP (AAV2/8-Con/Fon-EYFP) are injected into the vFMN and vIRt of the vGlut2-Cre animal, respectively. c, Projection pattern of vIRtvGlut2 neurons in the vFMN. d’,d’’,d’’’, High-magnification views of the boxed areas in d. The dorsolateral (d’), lateral (d’’), and ventrolateral (d’’’) facial motor subnuclei that contain motoneurons innervating the nasolabialis/maxillolabialis, nasolabialis profundus, and intrinsic muscles, respectively. e, A schematic of vIRtvGlut2 silencing experiment. f, Experimental setup for vibrissa tracking. One second running triggers 1 s continuous 561 nm laser stimulation. g, Representative image of NpHR3.3-EYFP expression in vIRt. h, Example vibrissa trace during laser-OFF and -ON period. i, Quantification of whisking amplitude (Laser-OFF, 7.9 ± 1.2° vs. Laser-ON, 7.9 ± 1.3°, n = 6, p = 0.8438, Wilcoxon signed-rank test). j, Quantification of whisking midpoint (Laser-OFF, 53.5 ± 3.1° vs. Laser-ON, 55.5 ± 3.5°, n = 6, p = 0.1562, Wilcoxon signed-rank test). k, Power spectrum analysis of whisking frequency in Laser-OFF and -ON period. Shaded areas are mean ± s.e.m. Data are mean ± s.e.m. Brain sections were counterstained with Neurotrace Blue (d, g).
Extended Data Fig. 2 Optogenetic silencing of vIRtPV neurons also impaired whisking.
a, A schematic of vIRt PV GtACR2 silencing experiment. b, Representative image of GtACR2 expression and the optic fibre track. c, Optic fibre tip placements over vIRt from all AAV-Flex-GtACR2-GFP injected animals. d, Experimental setup for vibrissa tracking. One second running triggers 1 s continuous 473 nm laser stimulation. e, Example vibrissa trace during laser-OFF and -ON period. f, Quantification of whisking amplitude (Laser-OFF, 15.6 ± 0.7° vs. Laser-ON, 10.2 ± 1.1°, n = 5, p = 0.0312, Wilcoxon signed-rank test). g, Power spectrum analysis of whisking frequency in Laser-OFF and -ON period. Data are mean ± s.e.m. * P ≤ 0.05. Brain sections were counterstained with Neurotrace Blue (b).
Extended Data Fig. 3 Antidromic opto-tagging of a ChRmine-expressing vIRtPV neuron via light stimulation through the ear.
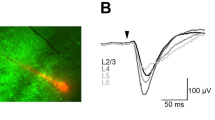
a, Overlapped average waveform, before (black) and during (red) stimulation periods. b, Raster plot of spike times aligned to stimulation onset for 40 light pulses. c, Example single channel recording trace showing antidromic spikes from the opto-tagged unit during a light pulse. d, Firing rate of that unit averaged over 40 light stimulations epochs. e, Unit activity during transition from resting to whisking state. vibrissa angle traces for the ipsilateral C2 vibrissa. Bottom: Spike raster plot for this opto-tagged vIRtPV-ChRmine neuron. f, Phase tuning. Average spike rate across whisking phases for this opto-tagged vIRtPV-ChRmine neuron, in polar (top) and cartesian coordinates (bottom).
Extended Data Fig. 4 Slow oscillation vIRt units and additional analysis of transition from tonic to rhythmic firing of vIRt units.
a, b, Two vIRt units with “slow” rhythmic activity patterns. Top: breathing trace. Middle: Vibrissa angle and midpoint traces. Bottom: raster plot of spiking events (protraction phases shown in beige). c, Top: vibrissa trace and raster plot for a retraction unit. Bottom: Time-frequency spectrum of that spiking activity. The transition to rhythmic bursting appears as a high power frequency band, corresponding to whisking frequency. d, Same as c, for a “slow” oscillation unit. Here the transition to whisking shows a low frequency band, similar to vibrissa midpoint variations or breathing rhythm. e, Left: Time-frequency spiking spectrum for a retraction unit, averaged over all whisking bouts. Right: Inter-spike interval histograms for that retraction unit. Red, overall ISI. Blue, ISI during long whisking bouts. The ISI distribution becomes bimodal, with a strong peak at short interval corresponding to bursts. f, same as e, for a “slow” oscillation unit.
Extended Data Fig. 5 Elevated extracellular potassium concentrations induce bursting but not rhythmic activity in vIRtPV neurons.
a,b, Cell-attached recording of a vIRtPV unit before (a) and after (b) bath application of K 9 mM. c, Representative histological verification of targeted recording. One of the tdTomato-expressing vIRtPV in slice is filled with green Alexa 488 dye from the recording pipette. d, Example of burst, post increase of extracellular potassium concentration. e, f, Inter-spike interval distribution, pre and post increase of extracellular potassium concentration, for one cell (e, pre: top, post: bottom) and all cells (f, ISI shown in log scale. Dash line: 40 Hz). g, Percentage of ISIs shorter than 25 ms (i.e., above 40 Hz), pre and post event, for all cells (average shown in black). h, Inter-burst frequency, for each cell, pre and post. No bursting frequency band is observed for any cell.
Extended Data Fig. 6 Pre-vIRtPV neurons in the brainstem and motor cortex, and neurotransmitter characterization of pre-vIRtPV neurons.
a, Representative image of ΔG-GFP labelled pre-vIRtPV neurons in the brainstem (continued from Fig. 3c). Scale bars, 200 µm. b, Representative image of ΔG-GFP labelled pre-vIRtPV neurons in the cortex. Scale bar, 500 µm. c, Zoomed image of the boxed area in b, d, Representative 3D reconstructed image of labelled pre-vIRtPV neurons in the cortex (magenta). Shaded areas denote the primary motor (MOp) and secondary motor cortices (MOs). e, Neurotransmitter phenotype of pre-vIRtPV neurons determined by fluorescent in situ hybridization or HCR RNA-FISH.
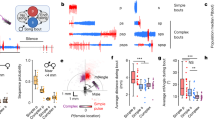
Extended Data Fig. 7 Comparison of the distributions of vIRtPV presynaptic neurons and vibrissal premotor neurons.
a, Three-dimensionally reconstructed vIRtPV presynaptic neurons (vIRt-pre, cyan, n = 3) and vibrissal premotor neurons (vibrissa-pre, magenta, n = 4) in the Allen Mouse Brain CCF in coronal planes. b, Density analysis of presynaptic neurons (vIRt-pre, cyan, n = 3) and vibrissal premotor neurons (vibrissa-pre, magenta, n = 4) in coronal and sagittal planes. DN, dentate nucleus; IP, interposed nucleus.
Extended Data Fig. 8 Additional results from vIRtPV-ChRmine-GFE3 mice.
a, Average whisking activity of a vIRtPV-ChRmine-GFE3 mouse, showing the mean of the pixel intensity difference between each consecutive video frame. Blue to red colour scale: low to high activity. In contrast to the ipsilateral (GFE3) side, contralateral whisking is highly active across the whole whisking range. b, Increased firing, but no rhythmic activity at whisking initiation for a opto-tagged vIRtPV-ChRmine-GFE3 unit (whisking initiation time determined from contralateral C2 vibrissa). c, Whisking phase tuning of all opto-tagged vIRtPV-ChRmine-GFE3 single units (conventions as in Figs. 2 and 4).
Extended Data Fig. 9 Schematic model of the vIRtPV circuit that generates rhythmic whisking in normal and experimental conditions.
a, In Resting state, vIRtPV neurons show unsynchronized tonic activity. b, In Normal whisking condition, tonic excitatory inputs to vFMN protractor motoneurons protract vibrissae. Concurrently, tonic excitatory inputs to vIRtPV neurons induce recurrent inhibition within vIRtPV and which in turn switches vIRtPV from tonic firing to synchronized rhythmic bursting mode. The rhythmic signal from vIRtPV periodically silences vFMN protractor motoneurons and leads to rhythmic whisking. The rhythmic inhibitory signal from the inspiratory rhythm generator preBötC resets the activity of vIRtPV. The expiratory oscillator activates vFMN retractor motoneurons. c, In vIRtPV-TeLC condition, outputs from vIRtPV are abolished. Lack of inhibition from vIRtPV results in strong continuous activation of vFMN protractor motoneurons and strong protraction of vibrissae. Because of the strong tonic activity of protractor intrinsic muscles, extrinsic retractor muscles play a minor role in vibrissa movement. d, In vIRtPV-GFE3 condition, tonic excitation induces strong unsynchronized tonic inhibitory outputs from vIRtPV to vFMN protractor motoneurons, which results in a less protracted midpoint compared with vIRtPV-TeLC’s. Under this condition, the contribution of expiratory oscillator-extrinsic retractor muscles becomes pronounced. A group of inhibitory neurons in the left top corner of vIRt indicates PV–/vGat+ vIRt neurons. Dotted lines denote putative connections.
Supplementary information
Supplementary Video 1
TeLC-mediated silencing of vIRtPV neurons. High-speed videography showing whisking deficits after TeLC silencing of vIRtPV neurons (left). The control side (right) shows no behavioural deficit. The video is slowed five times from the original speed.
Supplementary Video 2
vIRtPV neurons show rhythmic bursting in the whisking retraction phase. High-speed videography showing a mouse whisking while running on a wheel. The whisking phase polar plot is overlaid in the bottom left corner, with the current whisking phase shown as a green bar. Spikes from a simultaneously recorded opto-tagged vIRtPV retraction unit are indicated by an audio cue and red dot on the whisking phase polar plot.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Takatoh, J., Prevosto, V., Thompson, P.M. et al. The whisking oscillator circuit. Nature 609, 560–568 (2022). https://doi.org/10.1038/s41586-022-05144-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05144-8
This article is cited by
-
Latent neural population dynamics underlying breathing, opioid-induced respiratory depression and gasping
Nature Neuroscience (2024)
-
Cerebellar state estimation enables resilient coupling across behavioural domains
Scientific Reports (2024)
-
Thalamocortical loops as temporal demodulators across senses
Communications Biology (2023)
-
Neural mechanisms for the localization of unexpected external motion
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.