Abstract
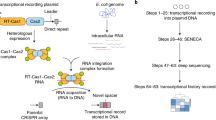
Biological processes depend on the differential expression of genes over time, but methods to make physical recordings of these processes are limited. Here we report a molecular system for making time-ordered recordings of transcriptional events into living genomes. We do this through engineered RNA barcodes, based on prokaryotic retrons1, that are reverse transcribed into DNA and integrated into the genome using the CRISPR–Cas system2. The unidirectional integration of barcodes by CRISPR integrases enables reconstruction of transcriptional event timing based on a physical record through simple, logical rules rather than relying on pretrained classifiers or post hoc inferential methods. For disambiguation in the field, we will refer to this system as a Retro-Cascorder.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the article and its Supplementary Information or will be made available from the authors on request. Sequencing data associated with this study are available in the NCBI Sequence Read Archive (PRJNA838025).
Code availability
Custom code to process and analyse data from this study is available on GitHub (https://github.com/Shipman-Lab/Spacer-Seq).
References
Simon, A. J., Ellington, A. D. & Finkelstein, I. J. Retrons and their applications in genome engineering. Nucleic Acids Res. 47, 11007–11019 (2019).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Church, G. M., Gao, Y. & Kosuri, S. Next-generation digital information storage in DNA. Science 337, 1628–1628 (2012).
Shipman, S. L., Nivala, J., Macklis, J. D. & Church, G. M. CRISPR–Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature 547, 345–349 (2017).
Yim, S. S. et al. Robust direct digital-to-biological data storage in living cells. Nat. Chem. Biol. 17, 246–253 (2021).
Ceze, L., Nivala, J. & Strauss, K. Molecular digital data storage using DNA. Nat. Rev. Genet. 20, 456–466 (2019).
Roquet, N., Soleimany, A. P., Ferris, A. C., Aaronson, S. & Lu, T. K. Synthetic recombinase-based state machines in living cells. Science 353, aad8559 (2016).
Sheth, R. U., Yim, S. S., Wu, F. L. & Wang, H. H. Multiplex recording of cellular events over time on CRISPR biological tape. Science 358, 1457–1461 (2017).
Schmidt, F., Cherepkova, M. Y. & Platt, R. J. Transcriptional recording by CRISPR spacer acquisition from RNA. Nature 562, 380–385 (2018).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Perli, S. D., Cui, C. H. & Lu, T. K. Continuous genetic recording with self-targeting CRISPR–Cas in human cells. Science 353, aag0511 (2016).
Park, J. et al. Recording of elapsed time and temporal information about biological events using Cas9. Cell 184, 1047–1063 (2021).
Shipman, S. L., Nivala, J., Macklis, J. D. & Church, G. M. Molecular recordings by directed CRISPR spacer acquisition. Science 353, aaf1175 (2016).
Simon, A. J., Morrow, B. R. & Ellington, A. D. Retroelement-based genome editing and evolution. ACS Synth. Biol. 7, 2600–2611 (2018).
Sharon, E. et al. Functional genetic variants revealed by massively parallel precise genome editing. Cell 175, 544–557 (2018).
Schubert, M. G. et al. High-throughput functional variant screens via in vivo production of single-stranded DNA. Proc. Natl Acad. Sci. USA 118, e2018181118 (2021).
Lopez, S. C., Crawford, K. D., Lear, S. K., Bhattarai-Kline, S. & Shipman, S. L. Precise genome editing across kingdoms of life using retron-derived DNA. Nat. Chem. Biol. 18, 199–206 (2022).
Farzadfard, F. & Lu, T. K. Genomically encoded analog memory with precise in vivo DNA writing in living cell populations. Science 346, 1256272 (2014).
Yosef, I., Goren, M. G. & Qimron, U. Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res. 40, 5569–5576 (2012).
Nuñez, J. K. et al. Cas1–Cas2 complex formation mediates spacer acquisition during CRISPR–Cas adaptive immunity. Nat. Struct. Mol. Biol. 21, 528–534 (2014).
Wang, J. et al. Structural and mechanistic basis of PAM-dependent spacer acquisition in CRISPR–Cas systems. Cell 163, 840–853 (2015).
Millman, A. et al. Bacterial retrons function in anti-phage defense. Cell 183, 1551–1561 (2020).
Bobonis, J. et al. Bacterial retrons encode tripartite toxin/antitoxin systems. Preprint at bioRxiv https://doi.org/10.1101/2020.06.22.160168 (2020).
Lampson, B. C. et al. Reverse transcriptase in a clinical strain of Escherichia coli: production of branched RNA-linked msDNA. Science 243, 1033–1038 (1989).
Silas, S. et al. Direct CRISPR spacer acquisition from RNA by a natural reverse transcriptase–Cas1 fusion protein. Science 351, aad4234 (2016).
Bonnet, J., Subsoontorn, P. & Endy, D. Rewritable digital data storage in live cells via engineered control of recombination directionality. Proc. Natl Acad. Sci. USA 109, 8884–8889 (2012).
Kim, S. et al. Selective loading and processing of prespacers for precise CRISPR adaptation. Nature 579, 141–145 (2020).
Ramachandran, A., Summerville, L., Learn, B. A., DeBell, L. & Bailey, S. Processing and integration of functionally oriented prespacers in the Escherichia coli CRISPR system depends on bacterial host exonucleases. J. Biol. Chem. 295, 3403–3414 (2020).
Chapman, K. B. & Boeke, J. D. Isolation and characterization of the gene encoding yeast debranching enzyme. Cell 65, 483–492 (1991).
Lim, D. Structure and biosynthesis of unbranched multicopy single-stranded DNA by reverse transcriptase in a clinical Eschehchia coli isolate. Mol. Microbiol. 6, 3531–3542 (1992).
Jung, H., Liang, J., Jung, Y. & Lim, D. Characterization of cell death in Escherichia coli mediated by XseA, a large subunit of exonuclease VII. J. Microbiol. 53, 820–828 (2015).
Han, E. S. et al. RecJ exonuclease: substrates, products and interaction with SSB. Nucleic Acids Res. 34, 1084–1091 (2006).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Grubbs, F. E. Procedures for detecting outlying observations in samples. Technometrics 11, 1–21 (1969).
Stefansky, W. Rejecting outliers in factorial designs. Technometrics 14, 469–479 (1972).
Hayflick, L. & Moorhead, P. S. The serial cultivation of human diploid cell strains. Exp. Cell. Res. 25, 585–621 (1961).
Yang, L. et al. Permanent genetic memory with >1-byte capacity. Nat. Methods 11, 1261–1266 (2014).
Yehl, K. & Lu, T. Scaling computation and memory in living cells. Curr. Opin. Biomed. Eng. 4, 143–151 (2017).
Mosberg, J. A., Gregg, C. J., Lajoie, M. J., Wang, H. H. & Church, G. M. Improving lambda Red genome engineering in Escherichia coli via rational removal of endogenous nucleases. PLoS ONE 7, e44638 (2012).
Moore, S. D. In Strain Engineering: Methods and Protocols (ed. Williams, J. A.) 155–169 (Humana Press, 2011).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Rogers, J. K. et al. Synthetic biosensors for precise gene control and real-time monitoring of metabolites. Nucleic Acids Res. 43, 7648–7660 (2015).
Acknowledgements
Work was supported by funding from the Simons Foundation Autism Research Initiative (SFARI) Bridge to Independence Award Program, the Pew Biomedical Scholars Program, the NIH/NIGMS (1DP2GM140917-01) and the UCSF Program for Breakthrough Biomedical Research. S.L.S. is a Chan Zuckerberg Biohub investigator and acknowledges additional funding support from the L.K. Whittier Foundation. S.K.L. was supported by an NSF Graduate Research Fellowship (2034836). S.C.L. was supported by a Berkeley Fellowship for Graduate Study. We thank K. Claiborn for editorial assistance.
Author information
Authors and Affiliations
Contributions
S.L.S. conceived the study with J.N. and G.M.C. contributing. S.B.-K. and S.L.S. designed experiments and analysed all data. Contributions to data collection were made by S.K.L. (Extended Data Fig. 4), C.B.F. (Extended Data Fig. 5) and S.L.S. (Figs. 1c,e and 3b–e). S.B.-K. collected all other data. E.R.L., M.G.S. and S.C.L. performed preliminary experiments not included in the figures. S.B.-K. performed the computational modelling of recordings. S.B.-K. wrote the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
S.L.S., G.M.C., M.G.S. and J.N. are named inventors on a patent application assigned to Harvard College, “Method of recording multiplexed biological information into a CRISPR array using a retron” (US20200115706A1).
Peer review
Peer review information
Nature thanks Channabasavaiah Gurumurthy and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Accompaniment to Figure 1.
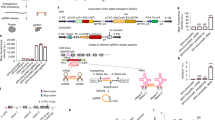
a, Hypothetical Eco1 wild-type ncRNA-linked RT-DNA structure. b, Hypothetical Eco1 v32 ncRNA-linked RT-DNA structure and hypothetical duplexed RT-DNA prespacer structure. Nucleotides that are altered from wild-type Eco1 are shown in orange. c, Hypothetical Eco1 v35 ncRNA-linked RT-DNA structure and hypothetical duplexed RT-DNA prespacer structure. Nucleotides that are altered from wild-type Eco1 are shown in green.
Extended Data Fig. 2 Accompaniment to Figure 2.
Hypothetical barcoded Eco1 v35 ncRNA-linked RT-DNA structure and hypothetical duplexed RT-DNA prespacer structure. Bases used to barcode retrons are shown in red.
Extended Data Fig. 3 Accompaniment to Figure 3.
a, Hypothetical wild-type Eco4 ncRNA-linked RT-DNA structure. ExoVII-dependent RT-DNA cleavage site is shown as a red slash. b, Eco4-derived spacer sequences and orientations. Bases are coloured to match Figure 3f. c, Proportion of Eco4-derived spacers in each orientation. Open circles are individual biological replicates.
Extended Data Fig. 4 Change in YFP fluorescence when expressed using inducible promoters.
The y-axis shows fluorescence (in arbitrary units) normalized to culture density (OD600).
Extended Data Fig. 5 Growth curves (upper plot) and max growth rates (lower plot) of E. coli with different combinations of retron recording components and inducers.
In growth curve plots the solid line is the mean OD600 of three biological replicates, with dotted lines showing the standard deviation. In maximum growth rate plots, each symbol is a single biological replicate. Bars show the mean and standard deviation. Statistically significant differences in maximum growth rate, as calculated by Tukey’s multiple comparison’s test, are highlighted. a, Growth kinetics of E. coli with different combinations of retron recording plasmids, all without inducers. b, Growth kinetics of E. coli with recording plasmid pSBK.079, with and without inducers. c, Growth kinetics of E. coli with signal plasmid pSBK.134, with and without inducers. Only one biological replicate is present in condition 'pSBK.134 + aTc' (pink). d, Growth kinetics of E. coli with signal plasmid pSBK.136, with and without inducers. e, Growth kinetics of E. coli with signal plasmid pSBK.134 and recording plasmid pSBK.079, with and without inducers. f, Growth kinetics of E. coli with signal plasmid pSBK.136 and recording plasmid pSBK.079, with and without inducers.
Extended Data Fig. 6 Accompaniment to Figure 4.
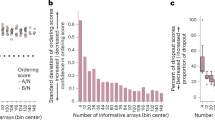
a, Ordering rules for pSBK.134 A-before-B replicates. The scores for each rule, and the composite score, are shown for each individual replicate. X-containing boxes indicate that no informative arrays, for that particular rule, were present in that replicate. b, As in panel a, ordering rules for pSBK.134 B-before-A replicates. c, As in panel a, ordering rules for pSBK.136 A-before-B replicates. d, As in panel a, ordering rules for pSBK.136 B-before-A replicates.
Extended Data Fig. 7 Long-term stability of retron-derived recordings in CRISPR arrays.
a, Ordering rules for 24+24-h, A-before-B recordings during post-recording multiday culture. Individual and composite scores are shown for samples taken on days 0, 2, 5, and 9 of culture. Each open circle represents the score, for that rule, from a single biological replicate. A total of 3 biological replicates are shown here. b, Changes in ordering rule scores over time in biological replicate 1. c, Changes in ordering rule scores over time in biological replicate 2. d, Changes in ordering rule scores over time in biological replicate 3.
Supplementary information
Supplementary Information
This file contains Supplementary Fig. 1 and Supplementary Tables 1–4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhattarai-Kline, S., Lear, S.K., Fishman, C.B. et al. Recording gene expression order in DNA by CRISPR addition of retron barcodes. Nature 608, 217–225 (2022). https://doi.org/10.1038/s41586-022-04994-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04994-6
This article is cited by
-
Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases
Genome Medicine (2024)
-
Genome expansion by a CRISPR trimmer-integrase
Nature (2023)
-
Histones direct site-specific CRISPR spacer acquisition in model archaeon
Nature Microbiology (2023)
-
Temporally resolved transcriptional recording in E. coli DNA using a Retro-Cascorder
Nature Protocols (2023)
-
Genomic Mosaicism of the Brain: Origin, Impact, and Utility
Neuroscience Bulletin (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.