Abstract
Wnt signalling is essential for regulation of embryonic development and adult tissue homeostasis1,2,3, and aberrant Wnt signalling is frequently associated with cancers4. Wnt signalling requires palmitoleoylation on a hairpin 2 motif by the endoplasmic reticulum-resident membrane-bound O-acyltransferase Porcupine5,6,7 (PORCN). This modification is indispensable for Wnt binding to its receptor Frizzled, which triggers signalling8,9. Here we report four cryo-electron microscopy structures of human PORCN: the complex with the palmitoleoyl-coenzyme A (palmitoleoyl-CoA) substrate; the complex with the PORCN inhibitor LGK974, an anti-cancer drug currently in clinical trials10; the complex with LGK974 and WNT3A hairpin 2 (WNT3Ap); and the complex with a synthetic palmitoleoylated WNT3Ap analogue. The structures reveal that hairpin 2 of WNT3A, which is well conserved in all Wnt ligands, inserts into PORCN from the lumenal side, and the palmitoleoyl-CoA accesses the enzyme from the cytosolic side. The catalytic histidine triggers the transfer of the unsaturated palmitoleoyl group to the target serine on the Wnt hairpin 2, facilitated by the proximity of the two substrates. The inhibitor-bound structure shows that LGK974 occupies the palmitoleoyl-CoA binding site to prevent the reaction. Thus, this work provides a mechanism for Wnt acylation and advances the development of PORCN inhibitors for cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The 3D cryo-EM maps have been deposited in the Electron Microscopy Data Bank under the accession numbers EMD-26707, EMD-26708, EMD-26709, EMD-26710 and EMD-26711. Atomic coordinates for the atomic model have been deposited in the Protein Data Bank under the accession numbers 7URA, 7URC, 7URD, 7URE and 7URF. Additional data supporting the findings in this study are provided as source data and supplementary information to this manuscript. Source data are provided with this paper.
Change history
18 July 2022
Fig. 4a was updated to correct the alignment of the stars and lines in panel a which had shifted slightly to the right.
References
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Taipale, J. & Beachy, P. A. The Hedgehog and Wnt signalling pathways in cancer. Nature 411, 349–354 (2001).
MacDonald, B. T., Tamai, K. & He, X. Wnt/β-catenin signaling: components, mechanisms, and diseases. Dev. Cell 17, 9–26 (2009).
Zhan, T., Rindtorff, N. & Boutros, M. Wnt signaling in cancer. Oncogene 36, 1461–1473 (2017).
Willert, K. et al. Wnt proteins are lipid-modified and can act as stem cell growth factors. Nature 423, 448–452 (2003).
Zhai, L., Chaturvedi, D. & Cumberledge, S. Drosophila wnt-1 undergoes a hydrophobic modification and is targeted to lipid rafts, a process that requires porcupine. J. Biol. Chem. 279, 33220–33227 (2004).
Hofmann, K. A superfamily of membrane-bound O-acyltransferases with implications for wnt signaling. Trends Biochem. Sci 25, 111–112 (2000).
Janda, C. Y., Waghray, D., Levin, A. M., Thomas, C. & Garcia, K. C. Structural basis of Wnt recognition by Frizzled. Science 337, 59–64 (2012).
Hirai, H., Matoba, K., Mihara, E., Arimori, T. & Takagi, J. Crystal structure of a mammalian Wnt–frizzled complex. Nat. Struct. Mol. Biol. 26, 372–379 (2019).
Rodon, J. et al. Phase 1 study of single-agent WNT974, a first-in-class Porcupine inhibitor, in patients with advanced solid tumours. Br. J. Cancer 125, 28–37 (2021).
Nusse, R. Wnts and Hedgehogs: lipid-modified proteins and similarities in signaling mechanisms at the cell surface. Development 130, 5297–5305 (2003).
Qi, X. & Li, X. Mechanistic Insights into the generation and transduction of Hedgehog signaling. Trends Biochem. Sci 45, 397–410 (2020).
Qi, X., Schmiege, P., Coutavas, E., Wang, J. & Li, X. Structures of human Patched and its complex with native palmitoylated Sonic Hedgehog. Nature 560, 128–132 (2018).
Qi, X., Schmiege, P., Coutavas, E. & Li, X. Two Patched molecules engage distinct sites on Hedgehog yielding a signaling-competent complex. Science 362, eaas8843 (2018).
Proffitt, K. D. & Virshup, D. M. Precise regulation of porcupine activity is required for physiological Wnt signaling. J. Biol. Chem. 287, 34167–34178 (2012).
Routledge, D. & Scholpp, S. Mechanisms of intercellular Wnt transport. Development 146, dev176073 (2019).
Kadowaki, T., Wilder, E., Klingensmith, J., Zachary, K. & Perrimon, N. The segment polarity gene porcupine encodes a putative multitransmembrane protein involved in Wingless processing. Genes Dev. 10, 3116–3128 (1996).
Resh, M. D. Targeting protein lipidation in disease. Trends Mol. Med. 18, 206–214 (2012).
Qian, H. et al. Structural basis for catalysis and substrate specificity of human ACAT1. Nature 581, 333–338 (2020).
Long, T., Sun, Y., Hassan, A., Qi, X. & Li, X. Structure of nevanimibe-bound tetrameric human ACAT1. Nature 581, 339–343 (2020).
Wang, L. et al. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature 581, 329–332 (2020).
Sui, X. et al. Structure and catalytic mechanism of a human triacylglycerol-synthesis enzyme. Nature 581, 323–328 (2020).
Guan, C. et al. Structural insights into the inhibition mechanism of human sterol O-acyltransferase 1 by a competitive inhibitor. Nat. Commun. 11, 2478 (2020).
Long, T., Liu, Y. & Li, X. Molecular structures of human ACAT2 disclose mechanism for selective inhibition. Structure 29, 1410–1418.e4 (2021).
Jiang, Y., Benz, T. L. & Long, S. B. Substrate and product complexes reveal mechanisms of Hedgehog acylation by HHAT. Science 372, 1215–1219 (2021).
Lee, C. J., Rana, M. S., Bae, C., Li, Y. & Banerjee, A. In vitro reconstitution of Wnt acylation reveals structural determinants of substrate recognition by the acyltransferase human Porcupine. J.Biol. Chem. 294, 231–245 (2019).
Grzeschik, K. H. et al. Deficiency of PORCN, a regulator of Wnt signaling, is associated with focal dermal hypoplasia. Nat. Genet. 39, 833–835 (2007).
Wang, X. et al. Mutations in X-linked PORCN, a putative regulator of Wnt signaling, cause focal dermal hypoplasia. Nat. Genet. 39, 836–838 (2007).
Liu, J. et al. Targeting Wnt-driven cancer through the inhibition of Porcupine by LGK974. Proc. Natl Acad. Sci. USA 110, 20224–20229 (2013).
Jiang, X. et al. Inactivating mutations of RNF43 confer Wnt dependency in pancreatic ductal adenocarcinoma. Proc. Natl Acad. Sci. USA 110, 12649–12654 (2013).
Madan, B. et al. Wnt addiction of genetically defined cancers reversed by PORCN inhibition. Oncogene 35, 2197–2207 (2016).
Asciolla, J. J., Miele, M. M., Hendrickson, R. C. & Resh, M. D. An in vitro fatty acylation assay reveals a mechanism for Wnt recognition by the acyltransferase Porcupine. J. Biol. Chem. 292, 13507–13513 (2017).
Rana, M. S. et al. Fatty acyl recognition and transfer by an integral membrane S-acyltransferase. Science 359, eaao6326 (2018).
Coupland, C. E. et al. Structure, mechanism, and inhibition of Hedgehog acyltransferase. Mol. Cell 81, 5025–5038.e5010 (2021).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Ma, D. et al. Crystal structure of a membrane-bound O-acyltransferase. Nature 562, 286–290 (2018).
Rios-Esteves, J. & Resh, M. D. Stearoyl CoA desaturase is required to produce active, lipid-modified Wnt proteins. Cell Rep. 4, 1072–1081 (2013).
Yu, J. et al. Structural model of human PORCN illuminates disease-associated variants and drug-binding sites. J. Cell Sci. 134, jcs259383 (2021).
Kakugawa, S. et al. Notum deacylates Wnt proteins to suppress signalling activity. Nature 519, 187–192 (2015).
Rios-Esteves, J., Haugen, B. & Resh, M. D. Identification of key residues and regions important for porcupine-mediated Wnt acylation. J. Biol. Chem. 289, 17009–17019 (2014).
Nygaard, R. et al. Structural basis of WLS/Evi-mediated Wnt transport and secretion. Cell 184, 194–206.e114 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Tuladhar, R. et al. Stereoselective fatty acylation is essential for the release of lipidated WNT proteins from the acyltransferase Porcupine (PORCN). J. Biol. Chem. 294, 6273–6282 (2019).
Zhong, Q. et al. Cryo-EM structure of human Wntless in complex with Wnt3a. Nat. Commun. 12, 4541 (2021).
Sun, Y. et al. Molecular basis of cholesterol efflux via ABCG subfamily transporters. Proc. Natl Acad. Sci. USA 118, e2110483118 (2021).
Wang, Q. et al. A combination of human broadly neutralizing antibodies against hepatitis B virus HBsAg with distinct epitopes suppresses escape mutations. Cell Host Microbe 28, 335–349.e336 (2020).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Brooks, B. R. et al. CHARMM: A program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 4, 30 (1983).
Li, H., Robertson, A. D. & Jensen, J. H. Very fast empirical prediction and rationalization of protein pKa values. Proteins 61, 704–721 (2005).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Lomize, M. A., Lomize, A. L., Pogozheva, I. D. & Mosberg, H. I. OPM: orientations of proteins in membranes database. Bioinformatics 22, 623–625 (2006).
Jorgensen, W., Chandrasekhar, J., Madura, J., Impey, R. & Klein, M. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 9 (1983).
Vanommeslaeghe, K. et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 16 (1995).
Acknowledgements
Cryo-EM data were collected at the UT Southwestern Medical Center Cryo-EM Facility (funded in part by the CPRIT Award RP170644). We thank D. Stoddard, L. Friedberg, L. Esparza and Y. Qin for technical support; A. Lemoff at the UTSW Proteomics Core for mass spectrometry analysis; C. Chattergee and Z. Chen for discussion; and E. Debler and P. Schmiege for editing the manuscript. This work was supported by NIH P01 HL020948, P01 HL160487, R01 GM135343 and Welch Foundation (I-1957) (to X.L.). N.E.-M. was funded in part by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany’s Excellence Strategy—EXC 2008-390540038-UniSysCat. X.Q. is a recipient of a DDBrown Fellowship of Life Sciences Research Foundation. B.W. is a Southwestern Medical Foundation Scholar in Biomedical Research. X.L. is a Damon Runyon-Rachleff Innovator supported by the Damon Runyon Cancer Research Foundation (DRR-53S-19) and a Rita C. and William P. Clements Jr Scholar in Biomedical Research at UT Southwestern Medical Center.
Author information
Authors and Affiliations
Contributions
X.L. conceived the project and designed the research with Y.L. and X.Q. Y.L. purified the protein and conducted the functional assays. Y.L., X.Q., T.L. and Y.S. carried out cryo-EM work. X.Q. refined the structures and contributed the HHAT structure. Y.L. and L.D. screened the antibody. N.E.-M. conducted the molecular dynamics simulations. R.W.Z. and B.W. synthesized WNT3Ap, pamWNT3Ap and conducted LC–MS. Y.L., X.Q., B.W. and X.L. analysed the data. X.L. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks K. Christopher Garcia and Stephen Long for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
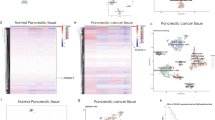
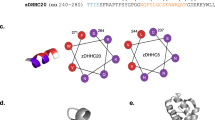
Extended Data Fig. 1 LC-MS analysis of PORCN-mediated WNT3Ap acylation and MS analysis of pamWNT3Ap.
a, The secretion pathway of WNT ligand. In the endoplasmic reticulum (yellow), WNT (purple) is modified by PORCN (orange). Transport of the modified WNT is mediated by WLS (blue). WNTs are secreted to extracellular space via vesicles. After binding to FZD (green), WNTs trigger the signal transduction. Notum (light blue) acts as a deacylase, removing the lipid of WNT to abolish the signal. The structure of palmitoleated WNT3A hairpin 2 is shown. b, Representative LC chromatograms of PORCN-mediated WNT3Ap acylation with (red) and without (blue) LGK974. ESI-MS: m/z calculated for [M+3H]3+ 840.041, found 840.039, and m/z calculated for [palmitoleated WNT3Ap+3H]3+ 918.779, found 918.778. c, Quantitative analysis of the product ratio after the reaction. Data are mean ± s.d. (n = 3 biologically independent experiments). ****P ≤ 0.0001, two-tailed unpaired t-test using GraphPad Prism 8. d, LC analysis of pamWNT3Ap. The samples (panels b and d) were applied to a HPLC column (DiscoveryBIO wide pore C5, 4.6x150mm, 5 μm) using a gradient in which percentage of solvent B increases from 20% to 80% in 6 min at 1.5 mL/min. e, Mass spectrometry analysis of [pamWNT3Ap+3H]3+. ESI-MS: m/z calculated for [M+3H]3+ (C124H196N33O28S5) 918.451, found 918.446
Extended Data Fig. 2 Protein Purification and data processing of palmitoleoyl-CoA-bound and LGK974-bound PORCN.
a, Representative Superose 6 increase 10/300 GL gel-filtration chromatogram of PORCN complex with Fab2C11. The peak fraction is shown on SDS-PAGE with molecular markers. Each protein is indicated. For gel source data, see Supplementary Fig. 1a. b, The data processing of palmitoleoyl-CoA-bound PORCN. The cryo-EM 3D classes as well as the mask used for the refinement are shown. The final cryo-EM map after cryoSPARC refinement was sharpened with a B-factor value of −170 Å2. c, Fourier shell correlation (FSC) curve of palmitoleoyl-CoA-bound PORCN map as a function of resolution using cryoSPARC output. d, The data processing of LGK974-bound PORCN. The cryo-EM 3D classes as well as the mask used for the refinement are shown. The final cryo-EM map after cryoSPARC refinement was sharpened with a B-factor value of −130 Å2. e, Fourier shell correlation (FSC) curve of LGK974-bound PORCN map as a function of resolution using cryoSPARC output.
Extended Data Fig. 3 cryo-EM map of structural elements of palmitoleoyl-CoA-bound and LGK974-bound PORCN.
a, The Fourier shell correlation (FSC) curves calculated between the refined structure model and the half map used for refinement (yellow), the other half map (gray) and the full map (blue) of palmitoleoyl-CoA-bound PORCN. b, Density map colored by local resolution estimation using cryoSPARC. c, The major helices of PORCN. d, MD simulation suggests that palmitoleoyl-CoA binds to PORCN in the curled-up conformation. After 100 ns simulations, the interaction between residues W300, H357 and the curled-down palmitoleoyl-CoA disrupts (the right bottom panel). The palmitoleoyl-CoA in either conformation with the cryo-EM map is shown. e, The Fourier shell correlation (FSC) curves calculated between the refined structure model and the half map used for refinement (yellow), the other half map (gray) and the full map (blue) of palmitoleoylated WNT3A-bound PORCN. f, Density map colored by local resolution estimation using cryoSPARC. g, The major helices of PORCN.
Extended Data Fig. 4 Data processing of WNT3Ap/LGK974-bound PORCN and pamWNT3Ap-bound PORCN.
a, The data processing of WNT3Ap/LGK974-bound PORCN. The cryo-EM 3D classes as well as the mask used for the refinement are shown. The final cryo-EM map after cryoSPARC refinement was sharpened with a B-factor value of −147 Å2. b, Fourier shell correlation (FSC) curve of WNT3Ap/LGK974-bound PORCN map as a function of resolution using cryoSPARC output. c, The data processing of pamWNT3Ap-bound PORCN. The cryo-EM 3D classes as well as the mask used for the refinement are shown. The final cryo-EM map after cryoSPARC refinement was sharpened with a B-factor value of −140 Å2. d, Fourier shell correlation (FSC) curve of pamWNT3Ap-bound PORCN map as a function of resolution using cryoSPARC output.
Extended Data Fig. 5 cryo-EM map of structural elements of WNT3Ap/LGK974-bound PORCN and pamWNT3Ap-bound PORCN.
a, The Fourier shell correlation (FSC) curves calculated between the refined structure model and the half map used for refinement (yellow), the other half map (gray) and the full map (blue) of WNT3Ap/LGK974-bound PORCN. b, Density map colored by local resolution estimation using cryoSPARC. c, The major helices of PORCN, WNT3Ap and LGK974. d, The Fourier shell correlation (FSC) curves calculated between the refined structure model and the half map used for refinement (yellow), the other half map (gray) and the full map (blue) of pamWNT3Ap-bound PORCN. e, Density map colored by local resolution estimation using cryoSPARC. f, The major helices of PORCN and pamWNT3Ap.
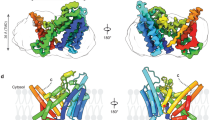
Extended Data Fig. 6 The structure of DHHC20 and the structural comparison of PORCN (gray) with the other MBOAT proteins.
a, The structure of DHHC20. The zinc ions are shown as gray balls. b, The structural comparison with DltB. c, The structural comparison with HHAT. IMP-1575, an HHAT inhibitor, is shown as green sticks. The catalytic H379 of HHAT is shown as sticks. d, The structural comparison with DGAT1. e, The structural comparison with ACAT1. The LGK974 is shown as magenta sticks. Nevanimibe, an ACAT1 inhibitor, is shown as green sticks. The acyl-CoA in the DGAT and ACAT1 is shown as yellow sticks.
Extended Data Fig. 7 Purification of human PORCN variants for activity assays.
Representative Superose 6 increase 10/300 GL gel-filtration chromatogram of PORCN variants in buffer containing 20 mM HEPES, pH 7.5, 150 mM NaCl, and 0.06% Digitonin.
Extended Data Fig. 8 Superimposing the WNT8A-FZD and WNT8A-WLS complexes into the structure of pamWNT3Ap-bound PORCN and the model of WNT3A-bound PORCN.
a, Superimposing the hairpin 2 of the structure of WNT8A-FZD complex into PORCN-pamWNT3Ap complex. The clash between WNT8A ligand and PORCN is indicated by a dashed circle. b, Superimposing the hairpin 2 of the structure of WNT8A-WLS complex into PORCN-pamWNT3Ap complex. The clash between WNT8A ligand and PORCN is indicated by a dashed circle. PAM, palmitoleic acid (red sticks). c, The predicted structure of WNT3A-bound PORCN. Three hairpins of WNT3A have been indicated. d, Structural comparison of the predicted PORCN structure to the cryo-EM structure (PDB:7URA).
Extended Data Fig. 9 The structure of HHAT-SHH-N complex.
a, Representative Superose 6 increase 10/300 GL gel-filtration chromatogram of HHAT-SHH-N complex with Fab3H02. The peak fraction is shown on SDS-PAGE with molecular markers. Each protein is indicated. For gel source data, see Supplementary Fig. 1b. b, Fourier shell correlation (FSC) curve as a function of resolution using RELION-3 output. c, Cryo-EM map of the complex after 3D refinement reveals the mean body of SHH-N protein at the threshold level of 0.003 but not 0.01.
Supplementary information
Supplementary Figure 1
This file contains the uncropped gels used in Extended Data Figs. 2a and 9a.
Supplementary Video 1
Molecular dynamics simulations of PORCN with palmitoleoyl-CoA in curled-up conformation in 100 ns time scale. The PORCN is colored in cyan, and the palmitoleoyl-CoA is colored in orange. Residues Trp300 and His357 are shown in sticks.
Supplementary Video 2
Molecular dynamics simulations of PORCN with palmitoleoyl-CoA in curled-down conformation in 100 ns time scale. The PORCN is colored in cyan, and the palmitoleoyl-CoA is colored in gray. Residues Trp300 and His357 are shown in sticks.
Supplementary Video 3
Molecular dynamics simulations of WNT3A-PORCN complex in 100 ns time scale. The PORCN is colored in cyan and the WNT3A is colored in blue. The disulfide bonds are shown in sticks.
Supplementary Video 4
Molecular dynamics simulations of PORCN in 100 ns time scale. The PORCN TM11 is colored in red.
Rights and permissions
About this article
Cite this article
Liu, Y., Qi, X., Donnelly, L. et al. Mechanisms and inhibition of Porcupine-mediated Wnt acylation. Nature 607, 816–822 (2022). https://doi.org/10.1038/s41586-022-04952-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04952-2
This article is cited by
-
Protein lipidation in cancer: mechanisms, dysregulation and emerging drug targets
Nature Reviews Cancer (2024)
-
The structure of phosphatidylinositol remodeling MBOAT7 reveals its catalytic mechanism and enables inhibitor identification
Nature Communications (2023)
-
Mechanism of action for small-molecule inhibitors of triacylglycerol synthesis
Nature Communications (2023)
-
Cryo-EM structures of Myomaker reveal a molecular basis for myoblast fusion
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.