Abstract
Synonymous mutations in protein-coding genes do not alter protein sequences and are thus generally presumed to be neutral or nearly neutral1,2,3,4,5. Here, to experimentally verify this presumption, we constructed 8,341 yeast mutants each carrying a synonymous, nonsynonymous or nonsense mutation in one of 21 endogenous genes with diverse functions and expression levels and measured their fitness relative to the wild type in a rich medium. Three-quarters of synonymous mutations resulted in a significant reduction in fitness, and the distribution of fitness effects was overall similar—albeit nonidentical—between synonymous and nonsynonymous mutations. Both synonymous and nonsynonymous mutations frequently disturbed the level of mRNA expression of the mutated gene, and the extent of the disturbance partially predicted the fitness effect. Investigations in additional environments revealed greater across-environment fitness variations for nonsynonymous mutants than for synonymous mutants despite their similar fitness distributions in each environment, suggesting that a smaller proportion of nonsynonymous mutants than synonymous mutants are always non-deleterious in a changing environment to permit fixation, potentially explaining the common observation of substantially lower nonsynonymous than synonymous substitution rates. The strong non-neutrality of most synonymous mutations, if it holds true for other genes and in other organisms, would require re-examination of numerous biological conclusions about mutation, selection, effective population size, divergence time and disease mechanisms that rely on the assumption that synoymous mutations are neutral.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing data generated in this study have been deposited into NCBI with the Bioproject ID PRJNA750109. Public data used include gene function annotations in the Saccharomyces Genome Database (https://www.yeastgenome.org/) and genomic coding sequences of S. paradoxus, C. glabrata, and S. castellii and genomic sequences of S. mikatae and S. uvarum from the NCBI genome assembly database (https://www.ncbi.nlm.nih.gov/assembly/). Source data are provided with this paper.
Code availability
Custom code is available at https://github.com/song88180/Mutational-Fitness-Effects and https://doi.org/10.5281/zenodo.5908478.
References
Kimura, M. Genetic variability maintained in a finite population due to mutational production of neutral and nearly neutral isoalleles. Genet Res 11, 247–269 (1968).
King, J. L. & Jukes, T. H. Non-Darwinian evolution. Science 164, 788–798 (1969).
Nei, M. & Kumar, S. Molecular Evolution and Phylogenetics (Oxford Univ. Press, 2000).
Li, W.-H. Molecular Evolution (Sinauer, 1997).
Graur, D., Sater, A. K. & Cooper, T. F. Molecular and Genome Evolution (Sinauer, 2016).
Hershberg, R. & Petrov, D. A. Selection on codon bias. Annu. Rev. Genet. 42, 287–299 (2008).
Chamary, J. V., Parmley, J. L. & Hurst, L. D. Hearing silence: non-neutral evolution at synonymous sites in mammals. Nat. Rev. Genet. 7, 98–108 (2006).
Plotkin, J. B. & Kudla, G. Synonymous but not the same: the causes and consequences of codon bias. Nat. Rev. Genet. 12, 32–42 (2011).
Stergachis, A. B. et al. Exonic transcription factor binding directs codon choice and affects protein evolution. Science 342, 1367–1372 (2013).
Zhou, Z. et al. Codon usage is an important determinant of gene expression levels largely through its effects on transcription. Proc. Natl Acad. Sci. USA 113, E6117–E6125 (2016).
Park, C., Chen, X., Yang, J. R. & Zhang, J. Differential requirements for mRNA folding partially explain why highly expressed proteins evolve slowly. Proc. Natl Acad. Sci. USA 110, E678–E686 (2013).
Presnyak, V. et al. Codon optimality is a major determinant of mRNA stability. Cell 160, 1111–1124 (2015).
Chen, S. et al. Codon-resolution analysis reveals a direct and context-dependent impact of individual synonymous mutations on mRNA level. Mol. Biol. Evol. 34, 2944–2958 (2017).
Kudla, G., Murray, A. W., Tollervey, D. & Plotkin, J. B. Coding-sequence determinants of gene expression in Escherichia coli. Science 324, 255–258 (2009).
Qian, W., Yang, J. R., Pearson, N. M., Maclean, C. & Zhang, J. Balanced codon usage optimizes eukaryotic translational efficiency. PLoS Genet. 8, e1002603 (2012).
Frumkin, I. et al. Codon usage of highly expressed genes affects proteome-wide translation efficiency. Proc. Natl Acad. Sci. USA 115, E4940–E4949 (2018).
Akashi, H. Synonymous codon usage in Drosophila melanogaster: natural selection and translational accuracy. Genetics 136, 927–935 (1994).
Sun, M. & Zhang, J. Preferred synonymous codons are translated more accurately: proteomic evidence, among-species variation, and mechanistic basis. Sci. Adv. (in the press).
Buhr, F. et al. Synonymous codons direct cotranslational folding toward different protein conformations. Mol. Cell. 61, 341–351 (2016).
Walsh, I. M., Bowman, M. A., Soto Santarriaga, I. F., Rodriguez, A. & Clark, P. L. Synonymous codon substitutions perturb cotranslational protein folding in vivo and impair cell fitness. Proc. Natl Acad. Sci. USA 117, 3528–3534 (2020).
Gilissen, C., Hoischen, A., Brunner, H. G. & Veltman, J. A. Disease gene identification strategies for exome sequencing. Eur. J. Hum. Genet. 20, 490–497 (2012).
Agashe, D., Martinez-Gomez, N. C., Drummond, D. A. & Marx, C. J. Good codons, bad transcript: large reductions in gene expression and fitness arising from synonymous mutations in a key enzyme. Mol. Biol. Evol. 30, 549–560 (2013).
Kristofich, J. et al. Synonymous mutations make dramatic contributions to fitness when growth is limited by a weak-link enzyme. PLoS Genet. 14, e1007615 (2018).
Lebeuf-Taylor, E., McCloskey, N., Bailey, S. F., Hinz, A. & Kassen, R. The distribution of fitness effects among synonymous mutations in a gene under directional selection. eLife 8, e45952 (2019).
Lind, P. A., Berg, O. G. & Andersson, D. I. Mutational robustness of ribosomal protein genes. Science 330, 825–827 (2010).
Sharon, E. et al. Functional genetic variants revealed by massively parallel precise genome editing. Cell 175, 544–557 (2018).
She, R. & Jarosz, D. F. Mapping causal variants with single-nucleotide resolution reveals biochemical drivers of phenotypic change. Cell 172, 478–490 (2018).
Qian, W., Ma, D., Xiao, C., Wang, Z. & Zhang, J. The genomic landscape and evolutionary resolution of antagonistic pleiotropy in yeast. Cell Rep. 2, 1399–1410 (2012).
Li, C., Qian, W., Maclean, M. & Zhang, J. The fitness landscape of a tRNA gene. Science 352, 837–840 (2016).
Chen, P. & Zhang, J. Asexual experimental evolution of yeast does not curtail transposable elements. Mol. Biol. Evol. 38, 2831–2842 (2021).
Keren, L. et al. Massively parallel interrogation of the effects of gene expression levels on fitness. Cell 166, 1282–1294 (2016).
Chang, Y. F., Imam, J. S. & Wilkinson, M. F. The nonsense-mediated decay RNA surveillance pathway. Annu. Rev. Biochem. 76, 51–74 (2007).
Monteiro, P. T. et al. YEASTRACT+: a portal for cross-species comparative genomics of transcription regulation in yeasts. Nucleic Acids Res. 48, D642–D649 (2020).
Sharp, P. M. & Li, W. H. The codon adaptation index—a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 15, 1281–1295 (1987).
Radhakrishnan, A. et al. The DEAD-Box protein Dhh1p couples mRNA decay and translation by monitoring codon optimality. Cell 167, 122–132 (2016).
Yang, J. R., Chen, X. & Zhang, J. Codon-by-codon modulation of translational speed and accuracy via mRNA folding. PLoS Biol. 12, e1001910 (2014).
Faure, G., Ogurtsov, A. Y., Shabalina, S. A. & Koonin, E. V. Role of mRNA structure in the control of protein folding. Nucleic Acids Res. 44, 10898–10911 (2016).
Goncalves, P., Valerio, E., Correia, C., de Almeida, J. M. & Sampaio, J. P. Evidence for divergent evolution of growth temperature preference in sympatric Saccharomyces species. PLoS ONE 6, e20739 (2011).
Kimura, M. The Neutral Theory of Molecular Evolution (Cambridge Univ. Press, 1983).
Lewontin, R. C. & Cohen, D. On population growth in a randomly varying environment. Proc. Natl Acad. Sci. USA 62, 1056–1060 (1969).
Gillespie, J. H. Natural selection for within-generation variance in offspring number II. Discrite haploid models. Genetics 81, 403–413 (1975).
Kimura, M. & Ohta, T. The average number of generations until fixation of a mutant gene in a finite population. Genetics 61, 763–771 (1969).
Flynn, J. M. et al. Comprehensive fitness maps of Hsp90 show widespread environmental dependence. eLife 9, e53810 (2020).
Dandage, R. et al. Differential strengths of molecular determinants guide environment specific mutational fates. PLoS Genet. 14, e1007419 (2018).
Chen, P. & Zhang, J. Antagonistic pleiotropy conceals molecular adaptations in changing environments. Nat. Ecol. Evol. 4, 461–469 (2020).
Azizoglu, A., Brent, R. & Rudolf, F. A precisely adjustable, variation-suppressed eukaryotic transcriptional controller to enable genetic discovery. eLife 10, e69549 (2021).
Natsume, T. & Kanemaki, M. T. Conditional degrons for controlling protein expression at the protein level. Annu. Rev. Genet. 51, 83–102 (2017).
Zhang, J. & Yang, J. R. Determinants of the rate of protein sequence evolution. Nat. Rev. Genet. 16, 409–420 (2015).
Wu, Z. et al. Expression level is a major modifier of the fitness landscape of a protein coding gene. Nat. Ecol. Evol. 6, 103–115 (2022).
Sauna, Z. E. & Kimchi-Sarfaty, C. Understanding the contribution of synonymous mutations to human disease. Nat. Rev. Genet. 12, 683–691 (2011).
Lee, T. I. & Young, R. A. Transcriptional regulation and its misregulation in disease. Cell 152, 1237–1251 (2013).
Chou, H. J., Donnard, E., Gustafsson, H. T., Garber, M. & Rando, O. J. Transcriptome-wide analysis of roles for tRNA modifications in translational regulation. Mol. Cell. 68, 978–992 (2017).
Laughery, M. F. et al. New vectors for simple and streamlined CRISPR–Cas9 genome editing in Saccharomyces cerevisiae. Yeast 32, 711–720 (2015).
Warringer, J., Ericson, E., Fernandez, L., Nerman, O. & Blomberg, A. High-resolution yeast phenomics resolves different physiological features in the saline response. Proc. Natl Acad. Sci. USA 100, 15724–15729 (2003).
Honlinger, A. et al. Tom7 modulates the dynamics of the mitochondrial outer membrane translocase and plays a pathway-related role in protein import. EMBO J. 15, 2125–2137 (1996).
Potapov, V. & Ong, J. L. Examining sources of error in PCR by single-molecule sequencing. PLoS ONE 12, e0169774 (2017).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 33, W465–W467 (2005).
Ranwez, V., Douzery, E. J. P., Cambon, C., Chantret, N. & Delsuc, F. MACSE v2: toolkit for the alignment of coding sequences accounting for frameshifts and stop codons. Mol. Biol. Evol. 35, 2582–2584 (2018).
Hofacker, I. L. et al. Fast folding and comparison of RNA secondary structures. Monatsh. Chem. 125, 167–188 (1994).
Zhang, J. & He, X. Significant impact of protein dispensability on the instantaneous rate of protein evolution. Mol. Biol. Evol. 22, 1147–1155 (2005).
Acknowledgements
We thank P. Chen, H. Liu, and H. Xu for technical assistance and W. Qian, X. Wei, J.-R. Yang, and members of the Zhang laboratory for valuable comments. This work was supported by the U.S. National Institutes of Health research grant R35GM139484 to J.Z.
Author information
Authors and Affiliations
Contributions
J.Z. conceived the study. X.S., C.L., and J.Z. designed the study. X.S. performed the experiments. X.S. and S.S. analysed the data. X.S. and J.Z. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Properties of wild-type and mutant strains analyzed.
a, Experimental procedure for testing cellular respiratory functions. Cells from each of the 21 mutant libraries were spread on YPD and YPG plates, followed by colony counting after growth. Respiration is needed for cell growth on YPG but not on YPD. b, Mean ratio of YPD colony number to YPG colony number for each mutant library, based on three replicates per library. Error bars show the standard error of the mean. The negative control is deficient in respiration due to gene deletions (see Methods). c, Maximum growth rates of three reconstituted wild-type strains and BY4742. WT1 was used as the wild-type control in en masse competitions with mutants. The red error bar indicates the standard error of the mean based on 16 replicates each shown by a dot (15 for BY4742). P-values are from two-tailed t-tests. The growth rate is not significantly different among the four strains (P = 0.58, one-factor ANOVA test). d, Ploidy of one T48 population per mutant library assessed by flow cytometry. SYTOX Green fluorescence was analyzed using the BL2 detector that measured the output from the 488-nm laser (blue). In control flow cytometry profiles, the two peaks respectively represent cells in the G1 and G2/M cell-cycle stages (1C and 2C DNA content for haploids while 2C and 4C for diploids)
Extended Data Fig. 2 Mutant fitness quantification.
a, Fractions of synonymous (yellow) and nonsynonymous (blue) mutants among designed but unobserved mutants and those among observed mutants. Nonsense mutants are not considered. Numbers in the bars are numbers of mutants. The distributions of synonymous and nonsynonymous mutants among the unobserved and observed mutant groups are not significantly different (P > 0.05, Fisher’s exact test). b–f, Correlation between every two of the four replicates in estimated mutant fitness under YPD at 30 °C. The correlation between replicate 1 and replicate 2 is presented in Fig. 1c. Each dot is a mutant and the dotted line indicates the diagonal. Pearson’s correlation r and its associated P-value are presented. Among-genotype sum of squares explains 93.8% of the total sum of squares (one-factor ANOVA)
Extended Data Fig. 3 Mutant fitness distribution under YPD at 30 °C.
a, Distribution of the fitness of 169 nonsense mutants. The peak around 0.94 is caused by 26 nonsense mutants of GET1 that all have fitness of about 0.94. b, Cumulative frequency distributions of log10(mutant fitness) of nonsynonymous (blue) and synonymous (yellow) mutants. c, The full figure of Fig. 2c, including low-fitness mutants that are not shown in Fig. 2c. d, The full figure of Fig. 2e, including low-fitness and high-fitness mutants that are not shown in Fig. 2e
Extended Data Fig. 4 Coding mutations influence the mRNA level of the mutated gene.
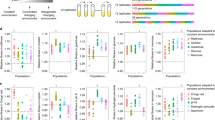
a, Non-significant negative correlation between the mean fitness of synonymous mutants of a gene and the expression level of the gene. Each dot represents a gene. Spearman’s correlation ρ and associated P-value are presented. b–g, Correlation in mutant REL between replicates, which are indicated on the axes of each panel. Each dot is a mutant, and the dotted line indicates the diagonal. Pearson’s correlation r and its associated P-value are presented. Among-genotype sum of squares explains 89.7% of total sum of squares (one-factor ANOVA). h, Cumulative frequency distributions of REL of nonsynonymous and synonymous mutants. i, Relative expression level (REL) distributions of nonsynonymous (blue) and synonymous (yellow) mutants of 20 individual genes shown by box plots. The lower and upper edges of a box represent the first (qu1) and third (qu3) quartiles, respectively, the horizontal line inside the box indicates the median (md), the whiskers extend to the most extreme values inside inner fences, md ± 1.5(qu3-qu1), and the dots show outliers. Nonsynonymous and synonymous distributions of each gene are compared by a two-tailed Wilcoxon rank-sum test, with FDR-adjusted P-values indicated as follows: *, P < 0.05; ⁑, P < 0.01, ⁂, P < 0.001. j, Distribution of REL of nonsense mutants
Extended Data Fig. 5 Mechanisms underlying coding mutations’ fitness effects.
a–b, Box plots showing similar absolute fractional changes in the mRNA level induced by nonsynonymous (a) or synonymous (b) mutations within and outside TF-binding sites. The lower and upper edges of a box represent the first (qu1) and third (qu3) quartiles, respectively, the horizontal line inside the box indicates the median (md), the whiskers extend to the most extreme values inside inner fences, md ± 1.5(qu3-qu1), and the dots show outliers. P-values are from two-tailed Wilcoxon rank-sum test (n = 1191, 4736, 367, and 1411, respectively, for the four bars from left to right). c–d. Positive correlation between rCAI and rescaled fitness among nonsynonymous (c) and synonymous (d) mutants, respectively. e, Fraction of synonymous mutations lowering CAI increases with the expression level of the gene. f, Fraction of synonymous mutations lowering the expression level increases with the expression level of the gene. g, Fraction of nonsynonymous mutations lowering CAI increases with the expression level of the gene. h, Fraction of nonsynonymous mutations lowering the expression level increases with the expression level of the gene. i, Mean rescaled fitness of synonymous mutants declines with the expression level of the gene. j, Mean rescaled fitness of nonsynonymous mutants declines with the expression level of the gene. Because deleting a more highly expressed gene tends to cause a greater fitness reduction60, the finding in panel j means that the mean fitness reduction caused by a nonsynonymous mutation should rise with the expression level of the gene. In e-j, each dot represents a gene. k–l, positive correlation between the relative mRNA folding strength (rMFS) of a nonsynonymous (k) or synonymous (l) mutant and its rescaled fitness when rMFS is below 1. The rMFS of a mutant is its mRNA folding strength (i.e., the absolute value of its minimal folding energy) divided by that of the wild-type. In each panel, the correlation is separately computed for mutants with rMFS < 1 and those with rMFS > 1. In c-l, rank correlations (ρ) and associated P-values are shown
Extended Data Fig. 6 A higher coefficient of variation (CV) of fitness across environments for nonsynonymous than synonymous mutants can create a nonsynonymous to synonymous substitution rate ratio (dN/dS) that is substantially below 1 despite similar fitness effects of synonymous and nonsynonymous mutations in each environment.
a, Mean expected dN/dS from 1000 simulations of a population that experiences multiple different environments. A mutant is purged if its fitness is lower than a preset cutoff such as 0.98 or 0.99 in any environment. Shaded areas represent 95% confidence intervals. a. Results with CV = 0.004 for synonymous mutants. b, Results with CV = 0.005 for synonymous mutants. Note that, under the fitness cutoff of 0.99, dN/dS starts to increase with the number (m) of environments when m is large. Raising m reduces the fraction of synonymous mutations that are always neutral (FANS) as well as the fraction of nonsynonymous mutations that are always neural (FANN). Because the fitness CV is larger for nonsynonymous than synonymous mutants in the simulation, FANN decreases with m more quickly than does FANS when m is small. When m is large, FANN is small, making it possible for FANS to decrease with m more quickly than FANN. As a result, dN/dS might increase with m when m is large
Extended Data Fig. 7 Pairwise correlation between replicates in estimated mutant fitness in each of the three additional environments used.
a–c, Correlation between every two of the three replicates in estimated mutant fitness under SC at 37 °C. Each dot is a mutant and the dotted line indicates the diagonal. Pearson’s correlation r and its associated P-value are presented. Among-genotype sum of squares explains 96.1% of the total sum of squares (one-factor ANOVA). d–f, Correlation between every two of the three replicates in estimated mutant fitness under YPD + 0.375 mM H2O2. Among-genotype sum of squares explains 94.4% of the total sum of squares. g–i, Correlation between every two of the three replicates in estimated mutant fitness under YPE. j, Correlation between replicates 1 and 3 in estimated mutant fitness under YPE after the exclusion of SNF6 mutants. k, Correlation between replicates 2 and 3 in estimated mutant fitness under YPE after exclusion of SNF6 mutants. Panels g-k suggest that the fitness estimates of SNF6 mutants in replicate 3 under YPE are unreliable, so are unused in fitness estimation in YPE. When SNF6 is excluded, among-genotype sum of squares explains 91.0% of the total sum of squares in YPE
Extended Data Fig. 8 Mutant fitness in the three additional environments used.
a–c, Fractions of synonymous (yellow) and nonsynonymous (blue) mutants among designed but unobserved mutants and those among observed mutants in each environment. Nonsense mutants are not considered. Numbers in the bars are numbers of mutants. The distributions of synonymous and nonsynonymous mutants among the unobserved and observed mutant groups are not significantly different in each environment (P > 0.05, Fisher’s exact test). d–f, Cumulative frequency distributions of fitness of nonsynonymous and synonymous mutants in each environment. g–i, Fitness distributions of nonsynonymous and synonymous mutants of 19 individual genes shown by box plots in each environment. The lower and upper edges of a box represent the first (qu1) and third (qu3) quartiles, respectively, the horizontal line inside the box indicates the median (md), the whiskers extend to the most extreme values inside inner fences, md ± 1.5(qu3-qu1), and the dots show outliers. Nonsynonymous and synonymous distributions for each gene are compared by a two-tailed Wilcoxon sum-rank test, with the FDR-adjusted P-value indicated as follows: *, P < 0.05; ⁑, P < 0.01, ⁂, P < 0.001. j–l, Fractions of mutants with fitness significantly below 1 (P < 0.05), significantly above 1, and neither, respectively, in each environment. The error bar shows one standard error. The distributional difference between synonymous and nonsynonymous mutants among the three bins is tested by two-tailed Fisher’s exact test, with the P-value indicated. At FDR = 0.05, 40.7% and 0.7% of nonsynonymous mutations and 34.8% and 0.5% of synonymous mutations are significantly deleterious and beneficial, respectively, in SC+37 °C. These values become 35.5%, 1.7%, 31.9% and 1.6% in YPD+H2O2, and 47.6%, 1.4%, 45.6%, and 1.0% in YPE
Extended Data Fig. 9 Fractions of nonsynonymous (blue) and synonymous (yellow) neutral mutations in one environment (indicated on the X-axis) that become deleterious in any of the other three environments.
The fractions are higher for nonsynonymous than synonymous mutations (P < 0.05, paired t-test). A mutation is considered deleterious if its fitness is significantly lower than 1 (P < 0.05) and neutral if its fitness is not significantly different from 1
Extended Data Fig. 10 A new model explaining the widespread negative correlation between the mRNA level of a gene and its evolutionary rate measured by the nonsynonymous or amino acid substitution rate.
Compared with nonsynonymous mutations in lowly expressed genes, those in highly expressed genes tend to reduce the gene expression level and hence tend to be deleterious. As a result, the evolutionary rate of a gene measured by the nonsynonymous or amino acid substitution rate is negatively correlated with the gene expression level. The height of a symbol represents the quantity considered.
Supplementary information
Supplementary Data 1
Properties of the 21 genes studied.
Supplementary Data 2
Genotype frequencies in each replicate competition.
Supplementary Data 3
Fitness of all mutants in all environments.
Supplementary Data 4
Relative mRNA levels of all mutants in YPD.
Supplementary Data 5
Primers used in the study.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shen, X., Song, S., Li, C. et al. Synonymous mutations in representative yeast genes are mostly strongly non-neutral. Nature 606, 725–731 (2022). https://doi.org/10.1038/s41586-022-04823-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04823-w
This article is cited by
-
Iron-related gene mutations driving global Mycobacterium tuberculosis transmission revealed by whole-genome sequencing
BMC Genomics (2024)
-
Fitness difference between two synonymous mutations of Phytophthora infestans ATP6 gene
BMC Ecology and Evolution (2024)
-
Genetic analysis and molecular basis of G6PD deficiency among malaria patients in Thailand: implications for safe use of 8-aminoquinolines
Malaria Journal (2024)
-
A non-human primate model for human norovirus infection
Nature Microbiology (2024)
-
Codon language embeddings provide strong signals for use in protein engineering
Nature Machine Intelligence (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.