Abstract
Although bradykinesia, tremor and rigidity are the hallmark motor defects in patients with Parkinson’s disease (PD), patients also experience motor learning impairments and non-motor symptoms such as depression1. The neural circuit basis for these different symptoms of PD are not well understood. Although current treatments are effective for locomotion deficits in PD2,3, therapeutic strategies targeting motor learning deficits and non-motor symptoms are lacking4,5,6. Here we found that distinct parafascicular (PF) thalamic subpopulations project to caudate putamen (CPu), subthalamic nucleus (STN) and nucleus accumbens (NAc). Whereas PF→CPu and PF→STN circuits are critical for locomotion and motor learning, respectively, inhibition of the PF→NAc circuit induced a depression-like state. Whereas chemogenetically manipulating CPu-projecting PF neurons led to a long-term restoration of locomotion, optogenetic long-term potentiation (LTP) at PF→STN synapses restored motor learning behaviour in an acute mouse model of PD. Furthermore, activation of NAc-projecting PF neurons rescued depression-like phenotypes. Further, we identified nicotinic acetylcholine receptors capable of modulating PF circuits to rescue different PD phenotypes. Thus, targeting PF thalamic circuits may be an effective strategy for treating motor and non-motor deficits in PD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Reagents are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Albin, R. L., Young, A. B. & Penney, J. B. The functional anatomy of basal ganglia disorders. Trends Neurosci. 12, 366–375 (1989).
The Parkinson Study Group. Levodopa and the progression of Parkinson’s disease. N. Engl. J. Med. 351, 2498–2508 (2004).
Hamani, C., Saint-Cyr, J. A., Fraser, J., Kaplitt, M. & Lozano, A. M. The subthalamic nucleus in the context of movement disorders. Brain 127, 4–20 (2004).
Smiley-Oyen, A. L., Worringham, C. J. & Cross, C. L. Motor learning processes in a movement-scaling task in olivopontocerebellar atrophy and Parkinson’s disease. Exp. Brain Res. 152, 453–465 (2003).
Marinelli, L., Quartarone, A., Hallett, M., Frazzitta, G. & Ghilardi, M. F. The many facets of motor learning and their relevance for Parkinson’s disease. Clin. Neurophysiol. 128, 1127–1141 (2017).
Poewe, W. Non-motor symptoms in Parkinson’s disease. Eur. J. Neurol. 15, 14–20 (2008).
Saalmann, Y. B. Intralaminar and medial thalamic influence on cortical synchrony, information transmission, and cognition. Front. Syst. Neurosci. 8, 83 (2014).
Smith, Y., Raju, D. V., Pare, J. F. & Sidibe, M. The thalamostriatal sytem: a highly specific network of the basal ganglia circuitry. Trends Neurosci. 27, 520–527 (2004).
Brown, H. D., Baker, P. M. & Ragozzino, M. E. The parafascicular thalamic nucleus concomitantly influences behavioral flexibility and dorsomedial striatal acetylcholine output in rats. J. Neurosci. 30, 14390–14398 (2010).
Diaz-Hernandez, E. et al. The thalamostriatal projections contribute to the initiation and execution of a sequence of movements. Neuron 100, 739–752 (2018).
Jouve, L., Salin, P., Melon, C. & Kerkerian-Le Goff, L. Deep brain stimulation of the center median-parafascicular complex of the thalamus has efficient anti-parkinsonian action associated with widespread cellular responses in the basal ganglia network in a rat model of Parkinson’s disease. J. Neurosci. 30, 9919–9928 (2010).
Berendse, H. W. & Groenewegen, H. J. Organization of the thalamostriatal projections in the rat, with special emphasis on the ventral striatum. J. Comp. Neurol. 299, 187–228 (1990).
Heshmati, M. & Russo, S. J. Anhedonia and the brain reward circuitry in depression. Curr. Behav. Neurosci. Rep. 2, 146–153 (2015).
Smith, Y. & Parent, A. Differential connections of caudate nucleus and putamen in the squirrel monkey (Saimiri sciureus). Neuroscience 18, 347–371 (1986).
Kita, T., Shigematsu, N. & Kita, H. Intralaminar and tectal projections to the subthalamus in the rat. Eur. J. Neurosci. 44, 2899–2908 (2016).
Mouroux, M., Hassani, O. K. & Feger, J. Electrophysiological study of the excitatory parafascicular projection to the subthalamic nucleus and evidence for ipsi- and contralateral controls. Neuroscience 67, 399–407 (1995).
Wickersham, I. R., Sullivan, H. A. & Seung, H. S. Axonal and subcellular labelling using modified rabies viral vectors. Nat. Commun. 4, 2332–2332 (2013).
Chatterjee, S. et al. Nontoxic, double-deletion-mutant rabies viral vectors for retrograde targeting of projection neurons. Nat. Neurosci. 21, 638–646 (2018).
Thompson, K. J. et al. DREADD agonist 21 is an effective agonist for muscarinic-based DREADDs in vitro and in vivo. ACS Pharmacol. Transl. Sci. 1, 61–72 (2018).
Parker, P. R. L., Lalive, A. L. & Kreitzer, A. C. Pathway-specific remodeling of thalamostriatal synapses in parkinsonian mice. Neuron 89, 734–740 (2016).
Luh, L. M., Das, I. & Bertolotti, A. qMotor, a set of rules for sensitive, robust and quantitative measurements of motor performances in mice. Nat. Protoc. 12, 1451–1457 (2017).
Voorn, P., Vanderschuren, L. J., Groenewegen, H. J., Robbins, T. W. & Pennartz, C. M. Putting a spin on the dorsal-ventral divide of the striatum. Trends Neurosci. 27, 468–474 (2004).
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007).
Barroso-Chinea, P. et al. Expression of the mRNAs encoding for the vesicular glutamate transporters 1 and 2 in the rat thalamus. J. Comp. Neurol. 5, 703–715 (2007).
Parolari, L., Schneeberger, M., Heintz, N. & Friedman, J. M. Functional analysis of distinct populations of subthalamic nucleus neurons on Parkinson’s disease and OCD-like behaviors in mice. Preprint at bioRxiv https://doi.org/10.1038/s41380-021-01162-6 (2020).
Hontanilla, B., Parent, A. & Gimenez-Amaya, J. M. Parvalbumin and calbindin D-28k in the entopeduncular nucleus, subthalamic nucleus, and substantia nigra of the rat as revealed by double-immunohistochemical methods. Synapse 4, 359–367 (1997).
Levesque, J. C. & Parent, A. GABAergic interneurons in human subthalamic nucleus. Mov. Disord. 5, 574–584 (2005).
Ferguson, B. R. & Gao, W. J. PV interneurons: Critical regulators of E/I balances for prefrontal cortex-dependent behavior and psychiatric disorders. Front. Neural Circuits 12, 37 (2018).
Smith, Y. & Parent, A. Neurons of the subthalamic nucleus in primates display glutamate but not GABA immunoreactivity. Brain Res. 453, 353–356 (1988).
Emmi, A., Antonini, A., Macchi, V., Porzionato, A. & De Caro, R. Anatomy and connectivity of the subthalamic nucleus in humans and non-human primates. Front. Neuroanat. 14, 13 (2020).
Zhang, Y. et al. MeCP2 in cholinergic interneurons of nucleus accumbens regulates fear learning. eLife 9, e55342 (2020).
Deumens, R., Blokland, A. & Prickaerts, J. Modeling Parkinson’s disease in rats: an evaluation of 6-OHDA lesions of the nigrostriatal pathway. Exp. Neurol. 175, 303–317 (2002).
Carvalho, M. M. et al. Behavioral characterization of the 6-hydroxydopamine model of Parkinson’s disease and pharmacological rescuing of non-motor deficits. Mol. Neurodegener. 8, 14 (2013).
Nabavi, S. et al. Engineering a memory with LTD and LTP. Nature 511, 348–352 (2014).
Roy, D. S. et al. Memory retrieval by activating engram cells in mouse models of early Alzheimer’s disease. Nature 531, 508–512 (2016).
Gong, X. et al. An ultra-sensitive step-function opsin for minimally invasive optogenetic stimulation in mice and macaques. Neuron 107, 38–51 (2020).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Feduccia, A. A., Chatterjee, S. & Bartlett, S. E. Neuronal nicotinic acetylcholine receptors: neuroplastic changes underlying alcohol and nicotine addictions. Front. Mol. Neurosci. 5, 83 (2012).
Pettorossi, V. E. & Grassi, S. Different contributions of platelet-activating factor and nitric oxide in long-term potentiation of the rat medial vestibular nuclei. Acta Otolaryngol. Suppl. 545, 160–165 (2001).
Pitcher, G. M., Beggs, S., Woo, R. S., Mei, L. & Salter, M. W. ErbB4 is a suppressor of long-term potentiation in the adult hippocampus. Neuroreport 19, 139–143 (2008).
Derrick, B. E. & Martinez, J. L. Opioid receptor activation is one factor underlying the frequency dependence of mossy fiber LTP induction. J. Neurosci. 14, 4359–4367 (1994).
Hajos, M. et al. The selective α7 nicotinic acetylcholine receptor agonist PNU-282987 [N-[(3R)-1-Azabicyclo[2.2.2]oct-3-yl]-4-chlorobenzamide hydrochloride] enhances GABAergic synaptic activity in brain slices and restores auditory gating deficits in anesthetized rats. J. Pharmacol. Exp. Ther. 312, 1213–1222 (2005).
Roy, D. S. et al. Anterior thalamic dysfunction underlies cognitive deficits in a subset of neuropsychiatric disease models. Neuron 109, 2590–2603 (2021).
Tanimura, A., Du, Y., Kondapalli, J., Wokosin, D. L. & Surmeier, D. J. Cholinergic interneurons amplify thalamostriatal excitation of striatal indirect pathway neurons in Parkinson’s disease models. Neuron 101, 444–458 (2019).
McIntosh, J. M. et al. Analogs of alpha-conotoxin MII are selective for α6-containing nicotinic acetylcholine receptors. Mol. Pharmacol. 65, 944–952 (2004).
Beatty, J. A., Sylwestrak, E. L. & Cox, C. L. Two distinct populations of projection neurons in the rat lateral parafascicular thalamic nucleus and their cholinergic responsiveness. Neuroscience 162, 155–173 (2009).
Mandelbaum, G. et al. Distinct cortical–thalamic–striatal circuits through the parafascicular nucleus. Neuron 102, 636–652 (2019).
Klawonn, A. M. & Malenka, R. C. Nucleus accumbens modulation in reward and aversion. Cold Spring Harb. Symp. Quant. Biol. 83, 119–129 (2018).
Nelson, A. B. & Kreitzer, A. C. Reassessing models of basal ganglia function and dysfunction. Annu. Rev. Neurosci. 37, 117–135 (2014).
Watson, G. D. R. et al. Thalamic projections to the subthalamic nucleus contribute to movement initiation and rescue of parkinsonian symptoms. Sci. Adv. 7, eabe9192 (2021).
Kondoh, K. et al. A specific area of olfactory cortex involved in stress hormone responses to predator odours. Nature 532, 103–106 (2016).
Liu, K. et al. Lhx6-positive GABA-releasing neurons of the zona incerta promote sleep. Nature 548, 582–587 (2017).
Makara, J. K. et al. Involvement of nitric oxide in depolarization-induced suppression of inhibition in hippocampal pyramidal cells during activation of cholinergic receptors. J. Neurosci. 27, 10211–10222 (2007).
Pitcher, G. M. et al. Schizophrenia susceptibility pathway neuregulin 1-ErbB4 suppresses Src upregulation of NMDA receptors. Nat. Med. 17, 470–478 (2011).
Matsuyama, S. & Matsumoto, A. Epibatidine induces long-term potentiation (LTP) via activation of α4β2 nicotinic acetylcholine receptors (nAChRs) in vivo in the intact mouse dentate gyrus: both α7 and α4β2 nAChRs essential to nicotinic LTP. J. Pharmacol. Sci. 93, 180–187 (2003).
Fu, W. & Jhamandas, J. H. Beta-amyloid peptide activates non-alpha7 nicotinic acetylcholine receptors in rat basal forebrain neurons. J. Neurophysiol. 90, 3130–3136 (2003).
Sala, M. et al. CC4, a dimer of cytisine, is a selective partial agonist at α4β2/α6β2 nAChR with improved selectivity for tobacco smoking cessation. Br. J. Pharmacol. 168, 835–849 (2013).
Acknowledgements
We thank X. Gong and J. Ting for generating the Cre-dependent SOUL construct; C. Wang for viral packaging; M. Fleishman, B. Clear, J. Kim, and H. Zaniewski for technical assistance; Z. Yang and W. Chen for data analysis; and all members of the Feng laboratory for their support. The J. Douglas Tan Fellowship supported Y. Zhang. The Warren Alpert Distinguished Scholar Award and NIH 1K99NS125121-01 supported D.S.R. This work was supported by the Stanley Center for Psychiatric Research at the Broad Institute of MIT and Harvard, Hock E. Tan and K. Lisa Yang Center for Autism Research at MIT, James and Patricia Poitras Center for Psychiatric Disorders Research at MIT, and NIH BRAIN Initiative (U01MH114819) (to G.F.).
Author information
Authors and Affiliations
Contributions
Y. Zhang, D.S.R., and G.F. contributed to study design. Y. Zhang, D.S.R., T.A., C.S., M.E.S. and K.M.S. contributed to data collection and analysis. Y. Zhang, D.S.R., Y.H. and N.E.L. conducted surgeries and histological analyses. Y. Zhu and X.-M.L. conducted all in vivo calcium imaging experiments. Y.C., J.D. and Z.L. conducted all macaque experiments. H.A.S. and I.R.W. provided all rabies viruses. K.B.F. and E.M.C. provided the anterograde HSV viruses. Y. Zhang, D.S.R., and G.F. wrote the paper. All authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Tianyi Mao and the other, anonymous, reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
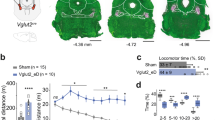
Extended Data Fig. 1 PF projections to CPu, NAc, and STN, and PFNAc neurons have distinct electrophysiological properties as compared to PFCPu and PFSTN populations.
a-b, Anterograde tracing from mouse PF, ChR2-eYFP virus injection site (a), and PF terminals in CPu, NAc, and STN (b). DAPI staining (blue). c, CTB488 injection site in CPu (left) along with corresponding upstream PF labeling (lower magnification image relative to Fig. 1a, right). d, Upstream labeling in PF with 1 (CTB555) vs. 3 (CTB488) injections in CPu. 80.5% of CPu-projecting PF neurons (i.e., CTB488) are labeled by a single injection of CTB555 in CPu (n = 3 mice). e-f, Injection sites of CTB555 in STN (left) (e) and CTB633 in NAc (left) (f) along with corresponding upstream PF labeling images (lower magnification relative to Fig. 1a, both right panels). g-i, Representative high magnification images from Fig. 1a, showing low overlap between CPu- and NAc-projecting PF neurons (g), CPu- and STN-projecting PF neurons (h), and STN- and NAc-projecting PF neurons (i). j, Percentage of CTB488+ only (CPu only), CTB555+ only (STN only), and CTB633+ only (NAc only) in PF. Dashed lines indicate chance level (20.49% for CPu only, 15.7% for STN only, 8.28% for NAc only) calculated using NEUN staining (n = 4 mice per group). k, Percentage of NEUN+ PF neurons projecting to the three different targets (n = 4 mice per group). l-p, Electrophysiological properties of PFCPu, PFNAc, and PFSTN neurons, which were labeled using retrograde RV injected into CPu, NAc, and STN respectively. Input resistance (Rin) (l), membrane time constant (tau) (m), membrane capacitance (Cm) (n), action potential amplitude (o), and action potential afterhyperpolarization (AHP) amplitude (p) (For Rin, tau, and Cm, PFCPu: n = 20 neurons (7, 7, 6), PFSTN: n = 17 neurons (6, 5, 6), PFNAc: n = 19 neurons (6, 7, 6) from 3 mice each. For amplitude and AHP, PFCPu: n = 18 neurons (5, 5, 8), PFSTN: n = 16 neurons (6, 5, 5), PFNAc: n = 19 neurons (6, 7, 6) from 3 mice each). q, ChR2-eYFP virus was injected in PF and ex vivo recordings were performed at PF cell bodies to validate virus expression (left), light responses in ChR2-labeled PF neurons were evoked by a 473 nm pulse train, shown in both current and voltage clamp modes (right). r, Reliability of ChR2 was compared across PF populations by recording from retrogradely-labeled CTB+ neurons that also expressed ChR2 (n = 12 neurons (4, 4, 4) from 3 mice for each group). s, Onset latency of evoked EPSCs recorded in postsynaptic neurons receiving PF input (PF→CPu: n = 19 neurons (7, 6, 6), PF→STN: n = 26 neurons (10, 9, 7), PF→NAc: n = 20 neurons (7, 7, 6) from 3 mice each). Data are presented as mean ± SEM; *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant. One-sample t test (j), one-way ANOVA followed by Bonferroni post-hoc test (k-p, s), and two-way ANOVA with repeated measures followed by Bonferroni post-hoc test (r). CPu only P = 0.033, STN only P = 0.028, NAc only P = 0.006 (j), F = 8.88, P = 0.007, CPu vs. STN t = 1.4, STN vs. NAc t = 2.74, CPu vs. NAc t = 4.14 (k), F = 27.71, P < 0.0001, PFCPu vs. PFNAc t = 6.57, PFSTN vs. PFNAc t = 6.28 (l), F = 17.48, P < 0.0001, PFCPu vs. PFNAc t = 5.17, PFSTN vs. PFNAc t = 5.04 (m), F = 11.76, P < 0.0001, PFCPu vs. PFNAc t = 3.37, PFSTN vs. PFNAc t = 4.69 (n), F = 7.19, P = 0.0018, PFCPu vs. PFNAc t = 3.78 (o), F = 6.11, P = 0.0042, PFCPu vs. PFNAc t = 3.26, PFSTN vs. PFNAc t = 2.65 (p), Amplitude: F = 0.03, DFn = 2, DFd = 297, P = 0.97, spikes/stimulus: F = 0.68, DFn = 2, DFd = 297, P = 0.51 (r), F = 0.11, P = 0.89 (s)
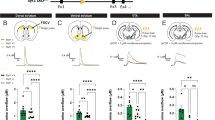
Extended Data Fig. 2 PF circuit electrophysiology, PF→STN terminal inhibition during motor learning, and cell type-specific tracing for the PF→CPu circuit.
a-c, Representative traces and quantification of evoked EPSCs in the presence of tetrodotoxin (TTX), 4-aminopyridine (4AP), and 6-cyano-7-nitroquinovaline-2,3-dione (CNQX) for PF→CPu (a), PF→STN (b), and PF→NAc (c) circuits (PF→CPu: n = 4 neurons (2, 1, 1), PF→STN: n = 3 neurons (1, 1, 1), PF→NAc: n = 3 neurons (1, 1, 1) from 3 mice each). Norm. peak EPSC amplitude plotted. d, Representative traces and quantification of paired-pulse ratio (also referred to as short-term plasticity) recordings in PF circuits (PF→CPu: n = 20 neurons (7, 6, 7), PF→STN: n = 23 neurons (8, 7, 8), PF→NAc: n = 20 neurons (7, 6, 7) from 3 mice each). e, C21-induced, reversible neuronal inhibition of a PF neuron expressing hM4Di ex vivo, using a step current injection protocol. f, C21-induced inhibition of PFCPu or PFSTN neurons during rotarod tests (n = 14 mCh, n = 19 PFCPu, n = 18 PFSTN mice). g, Experimental protocol for AMPA/NMDA ratio recordings. CaMKII-ChR2-eYFP virus was injected in PF, 3 weeks later animals were trained on the rotarod paradigm, and 1 hr after the end of training ex vivo recordings were performed. h, CaMKII-eArch3.0-eYFP virus was injected in PF and fibers were implanted above STN. eArch-eYFP virus expression in PF (left), and light-induced neuronal inhibition ex vivo (right). i, PF→STN terminal inhibition followed by cFos staining in STN using home cage or rotarod mice validated effective in vivo terminal inhibition (n = 4 mice per group). cFos was stained using a 633 secondary antibody (pseudocolored red). j, PF→STN terminal inhibition during rotarod. eYFP control mice received a CaMKII-eYFP virus in PF in place of the eArch virus. Norm. latency plotted relative to day 1 (n = 10 eYFP, n = 9 eArch mice). k-m, Monosynaptic retrograde RV tracing from D1+ or D2+ MSNs in CPu. Images show starter cells (k), FISH co-staining of GFP with D1 or D2 (l), and corresponding PF labeling (m) (n = 6 D1-Cre, n = 5 D2-Cre mice). n, Representative traces (left) and current-frequency curves (right) of ex vivo recordings from D2- (putative D1+) or D2+ MSNs in CPu (D1: n = 14 neurons (5, 4, 5), D2: n = 12 neurons (4, 5, 3) from 3 mice each). o, Anterior to posterior (AP) distribution of PV neurons in STN. Cre-dependent mCh virus was injected into STN of PV-Cre mice. Data are presented as mean ± SEM; *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant. One-way ANOVA followed by Bonferroni post-hoc test (a-c, f), two-way ANOVA with repeated measures followed by Bonferroni post-hoc test (d, n), unpaired t test (i, m), and two-tailed paired t test (j). F = 24.64, P < 0.0001, Baseline vs. TTX t = 4.56, TTX vs. TTX + 4AP t = 7.23, TTX + 4AP vs. TTX + 4AP + CNQX t = 6.99 (a), F = 9.59, P = 0.005, TTX vs. TTX + 4AP t = 4.71, TTX + 4AP vs. TTX + 4AP + CNQX t = 4.57 (b), F = 468.7, P < 0.0001, Baseline vs. TTX t = 22.74, TTX vs. TTX + 4AP t = 30.83, TTX + 4AP vs. TTX + 4AP + CNQX t = 28.97 (c), F = 6.41, DFn = 2, DFd = 120, P = 0.003 (d), Day 1: F = 11.30, P < 0.0001, mCh vs. PFCPu t = 3.25, mCh vs. PFSTN t = 1.00, Day 2: F = 22.58, P < 0.0001, mCh vs. PFCPu t = 1.70, mCh vs. PFSTN t = 4.38 (f), mCh P = 0.012, eArch P = 0.55 (i), eYFP P = 0.0032, eArch P = 0.24 (j), P = 0.61 (m), F = 28.31, DFn = 1, DFd = 240, P < 0.0001 (n)
Extended Data Fig. 3 Cell type-specific tracing for the PF→STN circuit, and PV+ STN neurons are critical for motor learning, are excitatory in nature, and lack local STN connectivity.
a-c, Monosynaptic retrograde RV tracing from VGLUT2+ or PV+ STN neurons. Images show starter cells in STN (a), FISH co-staining of GFP with VGLUT2 or PV (b), and corresponding PF labeling (c, lower magnification image relative to Fig. 2h) (n = 7 mice per group). d-e, Expression of a Cre-dependent hM4Di-mCh virus in STN (d), FISH co-staining of mCh with VGLUT2 or PV (e) in Vglut2-Cre or PV-Cre mice, related to experiments in Fig. 2j, k. f, Inhibition of STN VGLUT2- neurons during rotarod using a Cre-Off virus (DO-NpHR-eYFP) injected in STN of Vglut2-Cre mice. eYFP mice received a control Cre-Off virus (DO-eYFP) in place of the NpHR virus (n = 7 mice per group). FISH co-staining of VGLUT2 with eYFP (used a red fluorophore to visualize the eYFP probe). g, Inhibition of STN PV+ neurons that receive PF inputs during rotarod using an anterograde AAV expressing Cre injected in PF, and a Cre-On/Flp-On virus (COn/FOn-NpHR3.3-eYFP) injected in STN of PV-Flp mice. eYFP mice received a control Cre-On/Flp-On virus (COn/FOn-eYFP) in place of the NpHR3.3 virus (n = 7 mice per group). FISH co-staining of PV with eYFP (used a red fluorophore to visualize the eYFP probe). h-i, FISH staining of GFP-labeled VGLUT2+ neurons and endogenous VGLUT2 expression (h) or GFP-labeled PV+ neurons and endogenous PV expression (i) showed a high degree of overlap for each of these mouse lines. GFP labeling was achieved by injecting a Cre-dependent AAV in STN of Vglut2-Cre or PV-Cre mice. j-k, Half width (j) and spontaneous firing (k) measured in ex vivo recordings from STN VGLUT2+ and PV+ neurons. To visualize VGLUT2+ and PV+ neurons, a Cre-dependent mCh virus was injected in STN of Vglut2-Cre or PV-Cre mice (Half width, VGLUT2+: n = 18 neurons (6, 5, 7), PV+: n = 12 neurons (4, 4, 4) from 3 mice each; Spontaneous firing, VGLUT2+: n = 9 neurons (3, 3, 3), PV+: n = 9 neurons (3, 3, 3) from 3 mice each). l, Input resistance (Rin), membrane time constant (tau), membrane capacitance (Cm), and action potential amplitude of STN VGLUT2+ and PV+ neurons. To visualize VGLUT2+ and PV+ neurons, a Cre-dependent mCh virus was injected in STN of Vglut2-Cre or PV-Cre mice (Rin, tau, and Cm, VGLUT2+: n = 21 neurons (8, 8, 5), PV+: n = 13 neurons (5, 5, 3) from 3 mice each; Amplitude, VGLUT2+: n = 18 neurons (6, 5, 7), PV+: n = 12 neurons (4, 4, 4) from 3 mice each). m, VGLUT2, PV FISH staining in wild type STN sections. Plot (right) shows the proportion of VGLUT2+ or VGLUT3+ neurons among the total number of PV+ STN neurons (n = 5 VGLUT2+, n = 4 VGLUT3+ mice). n, Representative traces of evoked IPSCs (n = 10 neurons (3, 4, 3) from 3 mice) and evoked EPSCs (n = 10 neurons (2, 4, 4) from 3 mice) recorded in STN neurons during activation of ChR2-eYFP-expressing PV+ STN neurons. PV-Cre mice were injected with a Cre-dependent ChR2-eYFP virus in STN. Data are presented as mean ± SEM; *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant. Two-tailed unpaired t test (c, j-m) and two-tailed paired t test (f, g). P < 0.0001 (c), Cre-Off eYFP P = 0.001, Cre-Off NpHR P = 0.12 (f), eYFP P = 0.019, NpHR3.3 P = 0.85 (g), P = 0.0012 (j), P = 0.62 (k), Rin P = 0.14, Tau P = 0.11, Cm P = 0.26, Amplitude P = 0.51 (l), P = 0.0001 (m)
Extended Data Fig. 4 PV+ STN neurons send excitatory projections to GP and SNr, and PFNAc manipulations in motor, positive valence, and negative valence behaviors.
a, Monosynaptic HSV anterograde tracing from PV+ neurons in STN, using PV-Cre mice. Images show HSV helper cells (red) in STN overlapping with HSV-GFP-expressing cells (green) (i.e., starter cells), lack of labeling in known input brain regions (motor cortex, M1/M2 or PF thalamus) indicating that this virus does not travel retrogradely, and anterograde labeling in GP. b-c, Cre-dependent ChR2-eYFP virus injection (inj) site of PV-Cre mice (b), and representative images showing eYFP+ axonal terminals in GP and SNr (c). d-e, Retrogradely-labeled neurons (green) from GP (d) or SNr (e) colocalized with PV+ neurons in STN. Retrograde labeling was achieved by injecting AAVretro-GFP in GP or SNr. PV neurons were labeled by injecting a Cre-dependent mCh virus in STN of PV-Cre mice. f-g, Expression of Cre-dependent ChR2-mCh virus in STN of Vglut2-Cre or PV-Cre mice (f), FISH co-staining of mCh with VGLUT2 or PV (g). DAPI staining (blue). h, Representative traces, onset latency, and amplitude of evoked EPSCs recorded in GP neurons during terminal activation of STN VGLUT2+ or PV+ neurons (STNVGLUT2→GP: n = 12 neurons (4, 5, 3), STNPV→GP: n = 12 neurons (4, 4, 4) from 3 mice each). i, Representative traces of evoked EPSCs in the presence of picrotoxin (PTX) and 6-cyano-7-nitroquinovaline-2,3-dione (CNQX) for the STNVGLUT2→GP and STNPV→GP circuits. j, Representative traces of evoked EPSCs recorded in SNr neurons during activation of STN VGLUT2+ or PV+ neuronal terminals (STNVGLUT2→SNr: n = 22 neurons (6, 8, 8), STNPV→SNr: n = 18 neurons (6, 6, 6) from 3 mice each). k, GAD1 or GAD2 FISH with PV immunostaining in STN sections of cynomolgus macaques. l, C21-induced inhibition of PFNAc neurons during rotarod tests. mCh control mice data from Extended Data Fig. 2f (n = 14 mCh, n = 16 PFNAc mice). m, Real-time place preference behavior. NAc-projecting PF neurons were labeled by injecting RVdGL-Cre in NAc and Cre-dependent ChR2-eYFP in PF. Control group received the same injections except with Cre-dependent eYFP in PF. Blue light activation was performed at 20 Hz, only in the right side of the chamber (10 mW at patch cord) (n = 10 mice per group). n-o, Cocaine-induced conditioned place preference behavior, heat maps (n), quantification (o). NAc-projecting PF neurons were labeled by injecting RVdGL-Cre in NAc and Cre-dependent hM4Di-mCh in PF (referred to as hM4Di or PFNAc groups). Control group received the same injections except with Cre-dependent mCh in PF (mCh group). Baseline preference in the 3-chamber arena referred to as pre-exposure. After cocaine-induced training was completed (left side for cocaine, right side for saline), recall tests demonstrated conditioned place preference. C21-induced inhibition of PFNAc neurons was performed during the training phase (n = 12 mice per group). p, Contextual fear conditioning behavior. Mice were prepared similar to panel n. C21-induced inhibition was performed by injecting C21 40 min before the training day, followed by a long-term memory (LTM) test 24 hr later (n = 9 mice per group). q, Fiber photometry recordings from PFCPu, PFSTN, or PFNAc neurons by injecting a retrograde AAV expressing Cre in CPu, STN, or NAc, and Cre-dependent GCaMP6s in PF, followed by fibers placed above PF (left). Representative images (middle). Baseline in vivo GCaMP expression level (arbitrary units or a.u.) (right). Data are presented as mean ± SEM; *P < 0.05. NS, not significant. Two-tailed unpaired t test (h, l, m, p), two-tailed paired t test (d), and one-way ANOVA followed by Bonferroni post-hoc test (q). Onset latency P = 0.56, amplitude P = 0.012 (h), Day 1 P = 0.69, Day 2 P = 0.20 (l), P = 0.01 (m), Pre-exposure: mCh P = 0.46, PFNAc P = 0.34, Recall: mCh P = 0.02, PFNAc P = 0.21 (o), Training P = 0.89, LTM test P = 0.38 (p), F = 2.17, P = 0.16 (q)
Extended Data Fig. 5 In vivo fiber photometry analyses for open field and rotarod recordings.
a-f, Area under the curve (AUC) per sec for open field aligned to the onset of immobility (0 to 4 s compared to baseline −4 to 0 s) (a), 405 and 470 nm traces for each PF population for open field recordings aligned to the onset of immobility (b), averaged fluorescence change aligned to the end of immobility (c), AUC per sec for open field aligned to the end of immobility (−4 to 0 s compared to baseline 0 to 4 s) (d), 405 and 470 nm traces for each PF population for open field recordings aligned to the end of immobility (e), correlation between fluorescence change and mouse speed in the open field (r, correlation coefficient) (n = 5 mice per group) (f). g-j, AUC per sec for rotarod aligned to the onset of acceleration (0 to 4 s compared to baseline −2 to 0 s) (g), 405 and 470 nm traces for each PF population for rotarod recordings aligned to the onset of acceleration (h), AUC per sec for rotarod aligned to the onset of acceleration (0 to 4 s compared to baseline −2 to 0 s) for each individual session (i), 405 and 470 nm traces for each PF population for rotarod recordings aligned to the onset of acceleration showing individual session data (n = 5 mice per group) (j). Data are presented as mean ± SEM; *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant. One-way ANOVA followed by Bonferroni post-hoc test (a, d, g, i) and correlation analysis (f). F = 8.83, P = 0.004, PFCPu vs. PFSTN t = 3.42, PFCPu vs. PFNAc t = 3.83 (a), F = 0.168, P = 0.84, PFCPu vs. PFSTN t = 0.19, PFCPu vs. PFNAc t = 0.38 (d), F = 24.49, P < 0.0001, PFCPu vs. PFSTN t = 6.07, PFSTN vs. PFNAc t = 6.05 (g), 1st: F = 5.75, P = 0.02, PFCPu vs. PFSTN t = 3.02, PFSTN vs. PFNAc t = 2.85, 2nd: F = 4.816, P = 0.03, PFCPu vs. PFSTN t = 2.54, PFSTN vs. PFNAc t = 2.82, 3rd: F = 17.28, P = 0.0003, PFCPu vs. PFSTN t = 5.13, PFSTN vs. PFNAc t = 5.05 (i)
Extended Data Fig. 6 In vivo fiber photometry analyses for the end of the acceleration epoch in rotarod and tail suspension recordings.
a-c, Averaged fluorescence change and 405/470 nm traces for each PF population for rotarod recordings aligned to the end of acceleration (a), AUC per sec for rotarod aligned to the end of acceleration (−4 to 0 s compared to baseline 0 to 2 s) (b), correlation between fluorescence change and mouse speed (computed based on the speed of the rod) in the rotarod test (r, correlation coefficient) (n = 5 mice per group) (c). d-f, AUC per sec for tail suspension aligned to the onset of struggling (0 to 4 s compared to baseline −2 to 0 s) (d), 405 and 470 nm traces for each PF population for tail suspension recordings aligned to the onset of struggling (e), averaged fluorescence change and 405/470 nm traces aligned to the end of struggling, AUC per sec for tail suspension aligned to the end of struggling (−4 to 0 s compared to baseline 0 to 2 s) (n = 5 mice per group) (f). g-i, Averaged fluorescence change and AUC per sec for PFCPu hM4Di mice during open field (g), PFSTN hM4Di mice during rotarod (h), and PFNAc hM4Di mice during tail suspension (n = 5 mice per group) (i). PFCPu hM4Di, PFSTN hM4Di, and PFNAc hM4Di mice received a retrograde AAV expressing Cre in CPu, STN, or NAc, Cre-dependent GCaMP6s and hM4Di viruses in PF, followed by fibers placed above PF. Data are presented as mean ± SEM; *P < 0.05, **P < 0.01. NS, not significant. One-way ANOVA followed by Bonferroni post-hoc test (b, d, f) and two-tailed paired t test (g-i). F = 3.95, P = 0.048, PFCPu vs. PFSTN t = 2.68, PFSTN vs. PFNAc t = 2.07 (b), F = 9.73, P = 0.003, PFCPu vs. PFNAc t = 3.25, PFSTN vs. PFNAc t = 4.21 (d), F = 7.8, P = 0.0068, PFCPu vs. PFNAc t = 3.563, PFSTN vs. PFNAc t = 3.258 (f), P = 0.87 (g), P = 0.74 (h), P = 0.8 (i)
Extended Data Fig. 7 Prolonged chemogenetic inhibition of PFCPu neurons rescued locomotion in PD model mice up to 10 days later.
a, TH staining in CPu of saline control (left) or 6-OHDA-injected PD model (right) mice. b, Open field test using saline control and PD model mice 30 days after 6-OHDA injections (n = 11 saline, n = 8 PD model mice). c, Representative traces and evoked EPSC amplitudes recorded in CPu neurons receiving PF input from control and PD model mice (Control: n = 15 neurons (5, 5, 5), PD: n = 16 neurons (5, 6, 5) from 3 mice each). d, PFCPu inhibition in PD model mice by injecting a retrograde RVdGL-Cre in CPu, Cre-dependent hM4Di-mCh in PF, and 6-OHDA in SNc. e-j, Open arena behavior protocol using two different manipulation strategies starting from 14 days after 6-OHDA injections (e). For single manipulation experiments, total distance during a 20-min open arena test was measured prior to the C21 injection (Baseline), or 40 min and 3 days after one C21 injection (Test) (Baseline: n = 20 WTmCh, n = 16 PDmCh, n = 20 PDhM4Di mice; 40 min and 3 days: n = 14 WTmCh, n = 12 PDmCh, n = 14 PDhM4Di mice) (f). Saline was injected 40 min before the baseline session. For prolonged manipulation experiments, total distance, number of movements, and moving time (g-j) during tests were measured 7 days and 10 days after the 3-day prolonged C21 protocol (Day 7 and Day 10: n = 20 WTmCh, n = 16 PDmCh, n = 20 PDhM4Di mice). In panels h and i, baseline data plotted from panel f. WTmCh mice were injected with a retrograde RVdGL-Cre injected in CPu, Cre-dependent mCh in PF, and saline in SNc. Day 7 data related to Fig. 4b (j). Data are presented as mean ± SEM; **P < 0.01, ***P < 0.001. NS, not significant. Two-tailed unpaired t test (b, c), and one-way ANOVA followed by Bonferroni post-hoc test (f, g). P = 0.033 (b), P = 0.004 (c), Baseline: F = 17.77, P < 0.0001, WTmCh vs. PDmCh t = 5.26, PDmCh vs. PDhM4Di t = 0.006, 40 min after C21: F = 10.7, P = 0.0002, WTmCh vs. PDmCh t = 4.35, PDmCh vs. PDhM4Di t = 3.67, 3 days after C21: F = 15.35, P < 0.0001, WTmCh vs. PDmCh t = 5.44, PDmCh vs. PDhM4Di t = 2.06 (f), Day 7: F = 9.46, P = 0.0003, WTmCh vs. PDmCh t = 4.05, PDmCh vs. PDhM4Di t = 3.57, Day 10: F = 10.72, P = 0.0001, WTmCh vs. PDmCh t = 4.37, PDmCh vs. PDhM4Di t = 3.69 (g)
Extended Data Fig. 8 Strengthening the PF→STN circuit restores motor learning in PD model mice through PV+ STN neurons, and PD model mice show depression-like phenotypes.
a, AMPA/NMDA ratio recordings of PF→STN circuit from WT and PD model mice in home cage and rotarod conditions (WT data from Fig. 2e, PD data from Fig. 4d). b, cFos activation of STN neurons using home cage and rotarod PD model mice (n = 4 mice per group). c, PF→STN circuit manipulation in PD model mice by injecting CaMKII-oChIEF-mCh virus in PF, optic fibers above STN, and 6-OHDA injections in SNc (left). Expression of oChIEF-mCh in PF (right). d, Optical protocols including 20 Hz- and 100 Hz-only protocols. 20 Hz blue light was delivered during trials whereas 100 Hz blue light was delivered between trials. e, Activation of the PF→STN circuit in PD model mice using the 20 Hz or 100 Hz protocol during rotarod behavior (20 Hz: n = 9, 100 Hz: n = 7 mice). f, STN cFos activation in PD model mice (n = 4 mice per group) following PF→STN circuit activation with the 20 + 100 Hz protocol during rotarod. Control PD model mice (PDmCh) was prepared similar to the PDoChIEF mice except that a mCh virus was injected in PF. g-h, PF→STN circuit strengthening with VGLUT2 or PV inhibition in STN during rotarod, using PD model mice. CaMKII-oChIEF-mCh virus was injected in PF, Cre-dependent hM4Di virus in STN, 6-OHDA in SNc, and fibers targeted STN of Vglut2-Cre or PV-Cre mice (g). Rotarod behavior (n = 8 Vglut2-Cre, n = 7 PV-Cre mice) (h). i-j, Sucrose preference (n = 8 mice per group) (i) and total immobility time in forced swim (n = 9 WT, n = 8 PD + PFCPu mCh, n = 10 PD + PFCPu hM4Di mice) and tail suspension (n = 9 WT, n = 8 PD + PFCPu mCh, n = 8 PD + PFCPu hM4Di mice) tests in PD (j). To rule out the possibility that decreased locomotion of PD model mice resulted in their performance in these assays, a retrograde RVdGL-Cre was injected in CPu and Cre-dependent hM4Di-mCh was injected in PF of PD model mice (PD + PFCPu hM4Di group). For WT and PD + PFCPu mCh groups, Cre-dependent mCh was injected in PF in place of the hM4Di virus. C21 was injected 40 min before tests to rescue locomotion deficits in PD model mice by manipulating the PF→CPu circuit. k-m, Monosynaptic retrograde RV tracing from D1+ or D2+ NAc neurons. Images show starter cells in NAc (k), FISH co-staining of GFP with D1 or D2 (l), and corresponding PF labeling (n = 3 mice per group) (m). n, Representative traces (left) and current-frequency curves (right) of ex vivo recordings from D2- (putative D1+) or D2+ MSNs in NAc using D2-eGFP mice (D1: n = 20 neurons (6, 7, 7), D2: n = 16 neurons (6, 4, 6) from 3 mice each). o, Representative images of PF sections stained with cFos from mice expressing SOUL in PF neurons, including a no light group (SOUL - light) and a 5 min light activated group (SOUL + light) in the home cage. Data are presented as mean ± SEM; *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant. One-way ANOVA followed by Bonferroni post-hoc test (a), two-tailed unpaired t test (b, f, m), two-tailed paired t test (e, h), and two-way ANOVA with repeated measures followed by Bonferroni post-hoc test (n). F = 5.81, P = 0.0018, Home cage WT vs. home cage PD t = 4.04, Rotarod WT vs. Home cage PD t = 1.15 (a), P = 0.84 (b), 20 Hz P = 0.87, 100 Hz P = 0.72 (e), P = 0.012 (f), Vglut2-Cre P = 0.005, PV-Cre P = 0.59 (h), F = 4.28, P = 0.028, WT vs. PD + PFCPu mCh t = 2.71, PD + PFCPu mCh vs. PD + PFCPu hM4Di t = 0.39 (i), Forced swim: F = 8.92, P = 0.0013, WT vs. PD + PFCPu mCh t = 3.95, PD + PFCPu mCh vs. PD + PFCPu hM4Di t = 0.88, Tail suspension: F = 7.18, P = 0.004, WT vs. PD + PFCPu mCh t = 3.21, PD + PFCPu mCh vs. PD + PFCPu hM4Di t = 0.07 (j), P = 0.58 (m), F = 38.34, DFn = 1, DFd = 272, P < 0.0001 (n)
Extended Data Fig. 9 Testing molecular target candidates in the PF→STN and PF→CPu circuits.
a, GUCD1 (guanylyl cyclase gene, the NO receptor), ERBB4, and OPRM1 (μ-opioid receptor gene) FISH staining with PV in STN sections. 65.6% of all PV+ neurons were GUCD1+, 58.5% of all PV+ neurons were ERBB4+, 57.2% of all PV+ neurons were OPRM1+(n = 3 mice per group). b, AMPA/NMDA ratio recordings of the PF→STN circuit, specifically PV+ STN neurons, before and after optical high frequency stimulation (HFS) in PD model mice (n = 8 neurons (2, 3, 3) from 3 mice). oChIEF-mCh virus was injected in PF, Cre-dependent eYFP virus in STN, and 6-OHDA in SNc of PV-Cre mice. c-d, With DAMGO (μ-opioid receptor agonist), SNP (NO receptor agonist), AG1478 (ERBB4 receptor antagonist) (c) or PNU282987 (α7 agonist) (d) bath application, AMPA/NMDA ratio recordings of the PF→STNPV circuit before and after optical HFS in PD model mice (DAMGO: n = 7 neurons (2, 2, 3), SNP: n = 7 neurons (3, 2, 2), AG1478: n = 7 neurons (2, 2, 3), PNU282987: n = 8 neurons (2, 3, 3) from 3 mice each). oChIEF-mCh virus was injected in PF, Cre-dependent eYFP virus in STN, and 6-OHDA in SNc of PV-Cre mice. e, Local bilateral infusion of PNU282987, α-Ctx MII, or epibatidine into STN of PD model mice prior to rotarod behavior assays (n = 8 mice per group). f, α7 nAChRs FISH staining with PV in STN sections from mCh and KD mice, α7 in PV+ cells is decreased by 84% in KD mice as compared to mCh controls (n = 3 mCh, n = 4 α7 KD mice). g, Activating α7 nAChRs without (left) or with (right) α7 KD from PV+ STN neurons during rotarod, using PD model mice (n = 10 mCh, n = 8 α7 KD mice). PDα7 KD received a Cre-dependent Cas9 and gRNA viruses (for knockdown of α7 nAChRs) and cannula implants above STN, and 6-OHDA in SNc of PV-Cre mice. h, α6 nAChRs FISH staining with D1 or D2 in mouse CPu sections and with PFCPu neurons in mouse PF sections. PFCPu neurons were labeled by injecting AAVretro-GFP in CPu (n = 3 D1+, n = 3 D2+, n = 4 PFCPu+ mice). i, Representative cannula implant targeting CPu for local infusion experiments (left), local bilateral infusion of PBS, PNU282987, or epibatidine into CPu of PD model mice and infusion of PBS into CPu of WT mice prior to open field behavior (n = 10 WTPBS, 9 PDPBS, 9 PDPNU, 9 PDEpibat mice) (right). WTPBS and PDPBS data from Fig. 5e. j, α6 nAChRs FISH staining with PFCPu neurons in PF sections from mCh and KD mice, α6 in PFCPu+ cells is decreased by 78% in KD mice as compared to mCh controls (n = 4 mice per group). k, Total distance mice travelled in the open field after KD of α6 in PFCPu neurons of PD model mice (n = 8 mice per group). PDα6 KD received a retrograde RVdGL-Cre in CPu, Cre-dependent Cas9 and gRNA viruses (for knockdown of α6 nAChRs) in PFCPu neurons, and 6-OHDA in SNc. Data are presented as mean ± SEM; *P < 0.05, ***P < 0.001. NS, not significant. One-way ANOVA followed by Bonferroni post-hoc test (a, h, i), two-tailed paired t test (b-e, g), and two-tailed unpaired t test (f, j, k). F = 0.45, P = 0.13 (a), P = 0.60 (b), DAMGO P = 0.48, SNP P = 0.82, AG1478 P = 0.12 (c), P = 0.011 (d), PNU282987 P = 0.034, α-Ctx MII P = 0.34, Epibatidine P = 0.66 (e), P < 0.0001 (f), mCh + PNU282987 P = 0.04, α7 KD + PNU282987 P = 0.37 (g), F = 456.9, P < 0.0001, D1+ vs. PFCPu+ t = 25.49, D2+ vs. PFCPu+ t = 25.6 (h), F = 5.16, P = 0.0049, WTPBS vs. PDPBS t = 3.45, WTPBS vs. PDPNU t = 3.09, WTPBS vs. PDEpibat t = 2.86 (i), P < 0.0001 (j), P = 0.02 (k)
Extended Data Fig. 10 Testing molecular target candidates in the PF→NAc circuit, and rescue of motor phenotypes in the same mice using PFCPu cell body and PF→STN circuit manipulations in vivo.
a, Before and after bath application of PNU282987 (α7 agonist), CC4 (α6 agonist), and UB165 (α3 agonist), evoked EPSCs were recorded from D1 MSNs in NAc of PD model mice. oChIEF-mCh virus was injected in PF and 6-OHDA in SNc of D2-eGFP mice (PNU282987: n = 6 neurons (1, 2, 1, 1, 1), CC4: n = 7 neurons (1, 2, 1, 1, 2), UB165: n = 8 neurons (2, 2, 1, 2, 1) from 5 mice each). b, Representative cannula implant targeting NAc for local infusion experiments (left), local bilateral infusion of PBS, α-Ctx MII, or PNU282987 into NAc of PD model mice and infusion of PBS into NAc of WT mice prior to depression-like behavior tests (n = 9 mice per group) (right). WTPBS and PDPBS data from Fig. 5h. c, β2 nAChRs FISH staining with D1 in NAc sections from mCh and KD mice, β2 in D1+ cells is decreased by 80% in KD mice as compared to mCh controls (n = 3 mCh, n = 4 β2 KD mice). d, Activating β2 nAChRs without (PDmCh+Epibat) or with (PDβ2 KD+Epibat) β2 KD from D1+ NAc neurons during depression-like behaviors, using PD model mice (n = 9 mice per group). Dashed line indicates WT level from Fig. 5h. PDβ2 KD received a Cre-dependent Cas9 and gRNA viruses (for knockdown of β2 nAChRs), cannula implants in NAc, and 6-OHDA in SNc of D1-Cre mice. e, A retrograde RVdGL-Cre was injected in CPu, Cre-dependent hM4Di-mCitrine (for PFCPu labeling), and CaMKII-oChIEF-mCh (for STN terminal manipulations) in PF, optic fibers targeting STN, and 6-OHDA injected in SNc. f, Total distance in the open field was measured 40 min after C21 injections. PD model mice that received 6-OHDA injections in SNc without any subsequent PF manipulation was the baseline group (labeled PD). PD model mice with PFCPu manipulations are referred to as PD Rescue (n = 11 PD, n = 8 PD Rescue mice). g, Rotarod tests were performed after the open field paradigm. For rotarod, PF→STN circuit strengthening using our optical LTP protocol (20 + 100 Hz) was performed (n = 11 PD, n = 8 PD Rescue mice). Data are presented as mean ± SEM; *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant. Two-tailed paired t test (a, g), one-way ANOVA followed by Bonferroni post-hoc test (b), and two-tailed unpaired t test (c, d, f). PNU282987 P = 0.14, CC4 P = 0.17, UB165 P = 0.10 (a), Sucrose preference: F = 8.71, P = 0.0002, WTPBS vs. PDPBS t = 4.57, WTPBS vs. PDα-Ctx MII t = 4.14, WTPBS vs. PDPNU282987 t = 3.57, Forced swim: F = 5.59, P = 0.0034, WTPBS vs. PDPBS t = 3.16, WTPBS vs. PDα-Ctx MII t = 3.32, WTPBS vs. PDPNU282987 t = 3.52, Tail suspension: F = 5.32, P = 0.0044, WTPBS vs. PDPBS t = 3.44, WTPBS vs. PDα-Ctx MII t = 3.28, WTPBS vs. PDPNU282987 t = 2.99 (b), P < 0.0001 (c), Sucrose preference P = 0.022, Forced swim P = 0.012, Tail suspension P = 0.029 (d), P = 0.0017 (f), PD P = 0.78, PD Rescue P = 0.011 (g)
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, Y., Roy, D.S., Zhu, Y. et al. Targeting thalamic circuits rescues motor and mood deficits in PD mice. Nature 607, 321–329 (2022). https://doi.org/10.1038/s41586-022-04806-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04806-x
This article is cited by
-
Long-term labeling and imaging of synaptically connected neuronal networks in vivo using double-deletion-mutant rabies viruses
Nature Neuroscience (2024)
-
Characterization of graded 6-Hydroxydopamine unilateral lesion in medial forebrain bundle of mice
Scientific Reports (2024)
-
Protopanaxadiols Eliminate Behavioral Impairments and Mitochondrial Dysfunction in Parkinson’s Disease Mice Model
Neurochemical Research (2024)
-
Neuronal and synaptic adaptations underlying the benefits of deep brain stimulation for Parkinson's disease
Translational Neurodegeneration (2023)
-
Neural modulation with photothermally active nanomaterials
Nature Reviews Bioengineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.