Abstract
Haematopoietic stem cells (HSCs) arise in the embryo from the arterial endothelium through a process known as the endothelial-to-haematopoietic transition (EHT)1,2,3,4. This process generates hundreds of blood progenitors, of which a fraction go on to become definitive HSCs. It is generally thought that most adult blood is derived from those HSCs, but to what extent other progenitors contribute to adult haematopoiesis is not known. Here we use in situ barcoding and classical fate mapping to assess the developmental and clonal origins of adult blood in mice. Our analysis uncovers an early wave of progenitor specification—independent of traditional HSCs—that begins soon after EHT. These embryonic multipotent progenitors (eMPPs) predominantly drive haematopoiesis in the young adult, have a decreasing yet lifelong contribution over time and are the predominant source of lymphoid output. Putative eMPPs are specified within intra-arterial haematopoietic clusters and represent one fate of the earliest haematopoietic progenitors. Altogether, our results reveal functional heterogeneity during the definitive wave that leads to distinct sources of adult blood.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The Gene Expression Omnibus accession number is GSE180357 ('Single-cell transcriptomic interrogation of haematopoietic progenitors within the E11.5 AGM of the mouse'), which contains the scRNA-seq data. Source data are provided with this paper.
Code availability
Custom code for transposon alignment and analysis has been previously published20.
References
Bertrand, J. Y. et al. Haematopoietic stem cells derive directly from aortic endothelium during development. Nature 464, 108–111 (2010).
Boisset, J. C. et al. In vivo imaging of haematopoietic cells emerging from the mouse aortic endothelium. Nature 464, 116–120 (2010).
Jaffredo, T., Gautier, R., Eichmann, A. & Dieterlen-Lievre, F. Intraaortic hemopoietic cells are derived from endothelial cells during ontogeny. Development 125, 4575–4583 (1998).
Yokomizo, T. & Dzierzak, E. Three-dimensional cartography of hematopoietic clusters in the vasculature of whole mouse embryos. Development 137, 3651–3661(2010).
Dzierzak, E. & Speck, N. A. Of lineage and legacy: the development of mammalian hematopoietic stem cells. Nat. Immunol. 9, 129–136 (2008).
Medvinsky, A., Rybtsov, S. & Taoudi, S. Embryonic origin of the adult hematopoietic system: advances and questions. Development 138, 1017–1031 (2011).
Boisset, J. C. et al. Progressive maturation toward hematopoietic stem cells in the mouse embryo aorta. Blood 125, 465–469 (2015).
Chen, M. J., Yokomizo, T., Zeigler, B. M., Dzierzak, E. & Speck, N. A. Runx1 is required for the endothelial to haematopoietic cell transition but not thereafter. Nature 457, 887–891 (2009).
Kumaravelu, P. et al. Quantitative developmental anatomy of definitive haematopoietic stem cells/long-term repopulating units (HSC/RUs): role of the aorta-gonad-mesonephros (AGM) region and the yolk sac in colonisation of the mouse embryonic liver. Development 129, 4891–4899 (2002).
Rybtsov, S., Ivanovs, A., Zhao, S. & Medvinsky, A. Concealed expansion of immature precursors underpins acute burst of adult HSC activity in foetal liver. Development 143, 1284–1289 (2016).
Robin, C. et al. Human placenta is a potent hematopoietic niche containing hematopoietic stem and progenitor cells throughout development. Cell Stem Cell 5, 385–395 (2009).
Zhou, F. et al. Tracing haematopoietic stem cell formation at single-cell resolution. Nature 533, 487–492 (2016).
Dzierzak, E. & Bigas, A. Blood development: hematopoietic stem cell dependence and independence. Cell Stem Cell 22, 639–651 (2018).
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017).
Ganuza, M. et al. Lifelong haematopoiesis is established by hundreds of precursors throughout mammalian ontogeny. Nat. Cell Biol. 19, 1153–1163 (2017).
Henninger, J. et al. Clonal fate mapping quantifies the number of haematopoietic stem cells that arise during development. Nat. Cell Biol. 19, 17–27 (2017).
Batsivari, A. et al. Understanding hematopoietic stem cell development through functional correlation of their proliferative status with the intra-aortic cluster architecture. Stem Cell Rep. 8, 1549–1562 (2017).
Zhu, Q. et al. Developmental trajectory of prehematopoietic stem cell formation from endothelium. Blood 136, 845–856 (2020).
Baron, C. S. et al. Single-cell transcriptomics reveal the dynamic of haematopoietic stem cell production in the aorta. Nat. Commun. 9, 2517 (2018).
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014).
Pietras, E. M. et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell 17, 35–46 (2015).
Bowling, S. et al. An engineered CRISPR–Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1410–1422 (2020).
Christensen, J. L. & Weissman, I. L. Flk-2 is a marker in hematopoietic stem cell differentiation: a simple method to isolate long-term stem cells. Proc. Natl Acad. Sci. USA 98, 14541–14546 (2001).
Boyer, S. W., Schroeder, A. V., Smith-Berdan, S. & Forsberg, E. C. All hematopoietic cells develop from hematopoietic stem cells through Flk2/Flt3-positive progenitor cells. Cell Stem Cell 9, 64–73 (2011).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Beaudin, A. E. et al. A transient developmental hematopoietic stem cell gives rise to innate-like B and T cells. Cell Stem Cell 19, 768–783 (2016).
Christodoulou, C. et al. Live-animal imaging of native haematopoietic stem and progenitor cells. Nature 578, 278–283 (2020).
Ganuza, M. et al. Murine hematopoietic stem cell activity is derived from pre-circulation embryos but not yolk sacs. Nat. Commun. 9, 5405 (2018).
Rodriguez-Fraticelli, A. E. et al. Clonal analysis of lineage fate in native haematopoiesis. Nature 553, 212–216 (2018).
Forsberg, E. C. et al. Differential expression of novel potential regulators in hematopoietic stem cells. PLoS Genet. 1, e28 (2005).
Linton, P. J. & Dorshkind, K. Age-related changes in lymphocyte development and function. Nat. Immunol. 5, 133–139 (2004).
Cazzola, A. et al. Prenatal origin of pediatric leukemia: lessons from hematopoietic development. Front. Cell Dev. Biol. 8, 618164 (2020).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Dzierzak, E. & de Bruijn, M. Isolation and analysis of hematopoietic stem cells from mouse embryos. Methods Mol. Med. 63, 1–14 (2002).
Yokomizo, T. et al. Whole-mount three-dimensional imaging of internally localized immunostained cells within mouse embryos. Nat. Protoc. 7, 421–431 (2012).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Stern, D. L. Tagmentation-based mapping (TagMap) of mobile dna genomic insertion sites. Preprint at bioRxiv https://doi.org/10.1101/037762 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Lummertz da Rocha, E. et al. Reconstruction of complex single-cell trajectories using CellRouter. Nat. Commun. 9, 892 (2018).
Acknowledgements
We are grateful to members of the F.D.C. laboratory; M. Chen for scientific advice; the Harvard Stem Cell Core for technical support; and R. Mathieu and M. Paktinat for assistance and guidance with FACS. We also thank the laboratory of S. Hormoz and S. Liu for help with 10X encapsulation and sample processing. E.L.d.R. thanks CAPES (Coordination for the Improvement of Higher Education Personnel), Brazil. This work was supported by the National Institutes of Health grants HL128850-01A1 and P01HL13147, by the Evans MDS Foundation and by a Crazy 8 award from the Alex Lemonade Foundation to F.D.C. F.D.C. was a Leukemia and Lymphoma Society and a Howard Hughes Medical Institute Scholar. C.W. was funded by the Jane Coffin Childs Memorial Fund.
Author information
Authors and Affiliations
Contributions
S.H.P., C.C. and F.D.C. designed the study, analysed the data and wrote the manuscript. S.H.P. and C.C. performed and analysed the Flt3-CreER and transposon experiments, with help from Q.Y. (Flt3-CreER labelling experiments and LPS challenge), E.L.d.R. (single-cell sequencing analysis), S.B. (CARLIN model and analysis), L.L. (sequencing analysis), B.J.P.-M. (Flt3-CreER mouse generation) and F.G.O. (CARLIN model and analysis). C.W. performed the statistical analysis of the transposon data. G.Q.D. reviewed the manuscript. F.D.C. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Anna Bigas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of transposon barcoding within the embryo.
A: Diagram of several key processes involved in mouse blood development. B: Model of how the M2/SB/Tn system can be used to infer developmental lineage relationships. C: Relative Sleeping Beauty (SB) transposase mRNA levels at various time points following IV Dox at E9.5, demonstrating strong induction and return to baseline SB mRNA levels around 24–36 h. RNA was taken from VE-Cadherin+ cells sorted from the AGM. Values represent individual embryos and mean (n = 4 embryos). D: Protein levels of SB following induction (whole embryo lysates used for protein extraction; one representative embryo shown of n = 3). For gel source data, see Supplementary Fig. 1. E: The fraction of transposed cells was estimated by ddPCR, using a probe that spans the site of excision of the donor locus. Overall transposition plateaus after 36 h, with an effective transposition window of 12–36 h. Values were normalized to sorted DsRed+ granulocytes. Values represent individual embryos/mice and mean + S.D. (n = 3–8 embryos). F: Percentage of DsRed positive cells within the E12.5, E14.5, and adult bone marrow LSK compartment following intravenous (IV) Dox administration at E9.5. Labelling is consistent across LSK compartments and mature lineages, with a slight reduction in monocyte and B cell labelling. Values represent individual embryos/mice and mean (n = 4–6 mice). G: Transposition efficiency (as measured by DsRed percentage within the granulocyte compartment) at each induction time point. Values represent individual mice and mean + S.D. (n = 3–5 mice). H: In utero induction labels various cell types, including endothelial cells and putative HSPCs emerging within intra-arterial clusters (white arrows). A representative haematopoietic cluster within the umbilical artery of an iE7.5-induced embryo imaged at E10.5 is shown (whole-mount preparation, imaged with confocal microscopy; n = 2). Scale lengths of 50 μm (left) and 10 μm (right). I: Representation of clone sizes of HSC- (grey) or eMPP-derived (blue) Gr and B cells at 3 months and 1 year of age (from iE7.5, 9.5, and 13.5). Each set of plots (Gr and B) is drawn from one representative mouse. J: Comparison of overall contribution to mature lineages by HSCs and eMPPs. Output was normalized to the total HSPC output per lineage, and then averaged to generate a single value per mouse and per compartment. p-values derived from unpaired two-tailed t-tests with Welch correction (p = 0.0015 for iE6.5, p = 0.023 for iE7.5, p = 0.042 for iE8.5, and p = 0.020 for iE13.5; * < 0.05; ** < 0.005). Values represent individual mice and mean + S.D. (n = 3–5 mice).
Extended Data Fig. 2 Development and validation of a new insertion site capture method (TRACE).
A: Experimental flow chart representing the design and development of TRACE (transposase-assisted capture of transposable elements). In brief, custom transposomes were generated by combining annealed adaptors with purified Tn5 transposase. Through a two-step, nested, suppression PCR following tagmentation with the custom transposomes, sequencing libraries can be efficiently generated. B: Tn5 activity assay demonstrating an additive increase in fragmentation. Suppression PCR enables selective amplification of Sleeping Beauty Tn integrations (representative image of one reaction shown, n = 4). For gel source data, see Supplementary Fig. 1. C: Experimental design of detection limit of TRACE, wherein DNA from three HEK293T clones are added at varying dilutions (approximating 5, 10, 25, 50, 100, 1,000, 2,500 cells/clone in a final pool of ~10,000 cells) to a polyclonal DNA sample of integration sites. TRACE enables a semi-linear readout of clone size and has a reliable frequency detection limit of at least 25 cells in 10,000. Values represent individual replicates with a line drawn through their means. D: Cumulative frequency plots (left two graphs) of granulocyte and HSC barcode read-count fractions across the various time points. Each line represents one representative mouse. Only the top 200 barcodes are depicted. The fraction of total reads captured using various read frequency thresholds (right two graphs) is shown for each population, using the same mice as depicted in the cumulative read frequency plots. A threshold of 0.0005 was selected for subsequent transposon analysis as > 90% of reads were captured for each population at each time point. Values represent individual mice. E: Concordance of tags and reads between independently generated libraries from split samples of MPPs and granulocytes or whole-genome-amplified DNA from HSCs, at E9.5 or E13.5. Tags that fell below the read threshold were assigned a frequency of 1x10−4. The per cent of shared reads is depicted on each plot.
Extended Data Fig. 3 Statistical dropout model for quantifying eMPP clones.
A: Schematic for assessing experimental rate of dropout. DNA from DsRed+ adult HSCs from an iE9.5 mouse were extracted and whole-genome-amplified. Limiting dilutions of DNA were used as input for downstream library generation by TRACE. Tags retrieved with each DNA amount were compared against a reference set of high-confidence tags (defined as shared tags recovered from two independently generated libraries with 100 ng starting DNA). B: Retrieval rate (represented as both individual tags and total read counts) of high-confidence tags at limiting DNA input amounts. C: Distribution of HSC barcode reads across all MPP-containing clones from iE9.5. The mode at zero represents putative eMPPs. D: Generative model of barcode recovery and sequencing. E: Convergence of the log likelihood estimate for increasing quantity of \({X}_{i}^{{\rm{pre}}-{\rm{PCR}}}\) samples. Each dot represents an independent sample set for the given sample size. Results reflect the lognormal model applied to iE9.5 data with rmin = 0.7, pe = 0.35, μ = −0.1, σ = 3.6. F: Log likelihood of each MCMC chain, normalized to the max value. Each dot represents an independent chain. The lower values at the beginning represent the burn in period. G: Autocorrelation functions of the pe parameter in each MCMC chain. H: Comparison of the posterior moments of pe between two pools of MCMC restarts (1–25 vs. 25–50). Each dot represents a different combination of conditional parameters rmin, fcapture and model family (lognormal vs geometric). I: Posterior distribution of pe across a range of conditional parameter values, model families and time points. Dotted lines represent the 2.5th and 97.5th percentiles. J: Posterior predictive checks for four models, distinguished by two different values of rmin and two different parametric families of in vivo clone size distribution. For each model, the modal parameter estimates for iE9.5 were used to generate simulated read-count distributions (top). Quantile-quantile plots (bottom) compare the simulated read-count distribution to the empirical distribution from iE9.5. K: Difference of Bayesian evidence for two different values of rmin and two different parametric families of in vivo clone size distribution. Each bar represents a different combination of the remaining parameters (i.e. fcapture and either rmin or the in vivo clone size distribution). L: Difference of Bayesian evidence between models with and without eMPPs respectively. Each bar represents a different conditional value of fcapture. All models shown have rmin = 0.7 and a lognormal distribution for clone sizes in vivo.
Extended Data Fig. 4 Validation of eMPP clones using an alternative barcoding model, CARLIN.
A: Schematic of experiment, in which iCas9;cCarlin mice induced at E10.5 and sacrificed at 1 year of age for bone marrow analysis. Populations (LT-HSC, MPP, MkP, Gr, Mo, B) were isolated by FACS from bone marrow, processed, and analysed for edited Carlin alleles. B: Editing frequencies within granulocytes (n = 3), with an average of approximately 7%. Mean + S.E.M. of the three samples are depicted. C: Contribution of HSCs and eMPPs to mature lineages, represented as the number of UMIs shared/total UMIs in that population. Values represent individual mice; bar plots depict mean + S.D. (n = 3). D: Representative plot of one mouse demonstrating the Carlin alleles across the sampled populations (showing only those alleles found in either HSCs, eMPPs, or both). Each row represents a unique allele, and each column represents the population sampled. The colour of the allele reflects its frequency (in terms of log UMI count fraction) within that population (n = 3 adult mice). E: Lineage bias represented as the log of myeloid (Gr, Mo, and MkP) output / B output between HSCs and eMPPs. Output was first normalized to the total HSPC output per lineage. p-value derived from two-sided paired t-test (p = 0.0035). Values represent individual mice and mean + S.E.M.
Extended Data Fig. 5 Generation and characterization of the Flt3EGFP-CreERT2 mouse model.
A: Schematic of the Flt3 genetic locus targeting approach using a BAC targeting vector. The ATG start site of the first exon was targeted with an EGFP-CreERT2 resulting in a Flt3EGFP-CreERT2 transgenic mouse. B: Bone marrow FACS analysis of adult Flt3EGFP-CreERT2 for Flt3 and EGFP expression demonstrates that although EGFP correlates with endogenous Flt3 protein expression, it also tracks with a small population in the CD34+ and CD34- BM fraction that expresses Flt3 mRNA but undetectable protein levels. C: Representative bone marrow flow analysis of adult LSK populations in a Flt3EGFP-CreERT2 Rosa26loxP-stop-loxP-tdTomato mouse induced at 2 months of age and analysed 48 h later. Per cent Tomato labelling was assessed within the MPP3/4, ST-HSCs and LT-HSCs compartments (top panels, without Tam; bottom panels, with Tam) (n = 6). D: Scheme of experiment designed to test reconstitution potency of Tomato+ bone marrow cells within the adult. Forty-eight hours after induction of a 2-month-old Flt3EGFP-CreERT2 Rosa26loxP-stop-loxP-tdTomato mouse, whole bone marrow was isolated and 500,000 cells were transplanted into sub-lethally irradiated NOD SCID mice. Per cent Tomato chimerism of the CD45.2 fraction for granulocytes, B cells, and T cells was assessed monthly for 4 months post-transplantation. Each line represents an individual recipient mouse (n = 4). E: Scheme of experiment designed to test long-term reconstitution ability of E13.5 cKit+ cells fetal liver cells from Flt3EGFP-CreERT2 Rosa26loxP-stop-loxP-tdTomato embryos induced at E12.5. A total of 10,000 cKit+ fetal liver cells were transplanted into lethally irradiated CD45.1 recipients, along with 100,000 helper CD45.1 bone marrow cells. Per cent Tomato chimerism of the CD45.2 fraction for granulocytes, B cells, and T cells was assessed monthly for 10 months post-transplantation. Each blue line represents an individual recipient mouse that was transplanted with 10,000 cKit+ cells (n = 4).
Extended Data Fig. 6 Additional characterization of iE14.5, iE10.5 and iE9.5 Flt3-CreER mice.
A: Experimental design of embryonic lineage tracing using the EYFP reporter, with induction at E14.5. B: Representative E15.5 fetal liver flow analysis of iE14.5 Flt3EGFP-CreERT2 Rosa26LSL -EYFP embryos (n = 7). C: Experimental design of embryonic lineage tracing using the Tomato reporter, with induction at E10.5. D: Representative E15.5 fetal liver flow analysis of E10.5-induced Flt3EGFP-CreERT2 Rosa26LSL-tdTomato embryos examining labelling within stem cell and progenitor compartments (n = 8). E: Quantitative analysis of Tomato chimerism in bone marrow populations over time, following induction at E10.5. Multiple embryos or mice were grouped together for each time point. Total mice analysed: E15.5 fetal liver (n = 8), 9–12 months (n = 5), and > 12 months (n = 4). Significance assessed through one-way ANOVA with Holm–Sidak-adjusted multiple comparisons for LT-HSC, MPP3/4, and Gr (p = 0.0489 at 9–12 months; * < 0.05). Box plots drawn from 25th to 75th percentiles, centred about the median, with whiskers representing the min and max points. F: Experimental design of embryonic lineage tracing with the Flt3EGFP-CreERT2 Rosa26loxP-stop-loxP-tdTomato mouse model at iE9.5. G: Quantitative analysis of the per cent Tomato labelling in MPP3/4, ST-HSC and LT-HSCs fractions of the E15.5 fetal liver. Values represent individual embryos and mean + S.D. (n = 7). H: Quantitative analysis of percentage Tomato label in bone marrow (LSK) and peripheral blood populations at 1 month and 6–7 months of age in iE9.5-induced mice, demonstrating lymphoid-biased Tomato labelling and minimal MPP labelling over time. Values represent individual embryos and mean + S.D. (n = 5 for each time point).
Extended Data Fig. 7 Characterization of Tomato+ progenitors after an LPS challenge.
A: Experimental design of LPS challenge. Mice were induced in utero at iE14.5 and traced as adults. Peripheral blood was collected regularly, assessing for Tomato chimerism within granulocytes, B cells, and T cells. Two LPS challenges (IP administered) were given—one at 11 weeks of age and the other at 23 weeks of age. The control group received PBS delivered IP. B: Fraction of Tomato+ labelling relative to pre-LPS/PBS levels (at 10 weeks of age) for granulocytes, B cells, and T cells. Asterisks represent time points at which the difference between the PBS and LPS conditions were significant (p < 0.05; assessed using two-tailed, unpaired t-tests with Welch correction). Modified box plots are drawn from min to max, with a line at the mean. C: Frequencies of MPP3/4s, ST-HSCs, and LT-HSCs (as a fraction of the total LSK compartment) in the PBS and LPS conditions. Significance assessed using two-tailed, unpaired t-tests with Welch correction. Values represent individual mice and mean + S.D. D: Fraction of Tomato+ labelling relative to MPP3/4 (normalized per mouse). Values represent individual mice and mean + S.D. Significance assessed using two-tailed, unpaired t-tests with Welch correction. For all experiments, n = 3 (PBS arm) and n = 5 (LPS arm). The same cohort of mice were followed throughout.
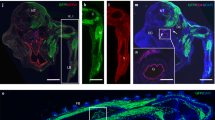
Extended Data Fig. 8 Flt3+ labelling within the E10.5 yolk sac and fetal liver.
A: Flow analysis of the AGM at E9.5, E10.5, and E11.5 and yolk sac (YS) at E9.5 and E10.5 (representative embryo; inductions performed one day earlier; n = 3–5). Flow plots demonstrate GFP and Tomato expression of all live cells. Tomato+ cells are present within the AGM and YS beginning at E10.5. B: Whole-mount confocal merged images of the E10.5 fetal liver bud, AGM, and umbilical artery in an embryo induced at E9.5 (representative image; n = 3). Top image: cKit and Tomato staining. Tomato+ (red) cells are detected within the fetal liver bud, in addition to other cKit+ (white) cells. Scale bar: 50 µm. Middle and bottom images: CD31, CD45, and Tomato staining. Arrows highlight Tomato+ CD45+ cells, and arrowheads highlight Tomato+ cells with diminished CD45+ labelling. Scale bars represent 50 µm (middle) and 20 µm (bottom). Sorting schemes can be found in Supplementary Fig. 4. C: Expression of haematopoietic and endothelial markers (CD45, CD31, CD41, and cKit) within the Tomato+ compartment within the E10.5 yolk sac (YS). Tomato+ cells are in black and superimposed on all Ter119- cells. E10.5 plots represent a concatenation of 5 embryos independently analysed. D: Fraction of Il7r+ cells within the Tomato+ cKit+ CD45+ Lin- compartment at the E10.5 AGM (n = 4, concatenated) and E11.5 AGM (n = 3, concatenated).
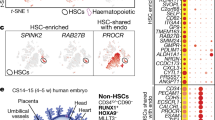
Extended Data Fig. 9 Transcriptional characterization of Flt3+ cells.
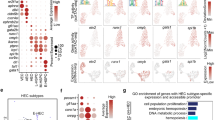
A: Sorting scheme for the scRNA-seq experiment. Three populations were independently sorted and isolated from three, pooled E11.5 AGMs or fetal livers (AGM pre-HSCs and haemogenic endothelial cells; AGM progenitors; and fetal liver progenitors). B: The top five differentially expressed genes per cluster. C: Source composition of each cluster. D: Normalized expression pattern of key differentiated and pre-HSC genes, visualized in the UMAP space. E: Adult LT-HSC, ST-HSC, and MPP4 gene module scores per cell, derived from the top 50 differentially expressed genes within single-cell adult dataset from ref. 29. Scale reflects normalized scores with a cut-off of 0.
Extended Data Fig. 10 Characterization of putative eMPP cellular origins.
A: Gating scheme for MDS-GFP analysis: pre-HSCs were defined as Lin- CD45+ cKit+ ESAM+ VE-Cadherin+, and progenitors were defined as Lin- CD45+ cKit+ Il7r- Flt3+. GFP positivity was measured in these two compartments and is quantified in the bar plot; mean + S.D. depicted (n = 4 embryos). A representative histogram from one of the four embryos is shown. B: Transcriptomic similarity of Prog cells to defined E11 populations from Baron et al19 using a random forest classifier. (Non-HE/HE: Non-haemogenic endothelium/haemogenic endothelium; YS HSPCs: yolk sac haematopoietic stem and progenitor cells; T1 pre-HSCs; T2 pre-HSCs). Flt3 and Cd48 expression depicted below. C: UMAP plot of ex vivo derived cultures from pre-circulation yolk sac and embryo proper tissues (from Ganuza et al28). D: Normalized expression patterns of several putative eMPP genes identified from our single-cell analysis, including Sox4, Hlf, and Hoxa9. A dot plot of key genes shows preferential enrichment within cells derived from the embryo proper. E: Analysis of CD41+ cKit+ cells in the E9.5 yolk sac of MDS-GFP embryos reveals no GFP positivity (representative embryo shown; n = 4).
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Supplementary References, Supplementary Tables 1–3 and Supplementary Figs. 1–4.
Source data
Rights and permissions
About this article
Cite this article
Patel, S.H., Christodoulou, C., Weinreb, C. et al. Lifelong multilineage contribution by embryonic-born blood progenitors. Nature 606, 747–753 (2022). https://doi.org/10.1038/s41586-022-04804-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04804-z
This article is cited by
-
Cis inhibition of NOTCH1 through JAGGED1 sustains embryonic hematopoietic stem cell fate
Nature Communications (2024)
-
Deciphering cell states and genealogies of human haematopoiesis
Nature (2024)
-
Cross-species transcriptomics reveals bifurcation point during the arterial-to-hemogenic transition
Communications Biology (2023)
-
Platelet and myeloid lineage biases of transplanted single perinatal mouse hematopoietic stem cells
Cell Research (2023)
-
Haematopoietic stem and progenitor cell heterogeneity is inherited from the embryonic endothelium
Nature Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.