Abstract
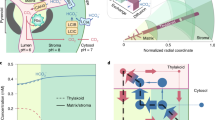
Global photosynthesis consumes ten times more CO2 than net anthropogenic emissions, and microalgae account for nearly half of this consumption1. The high efficiency of algal photosynthesis relies on a mechanism concentrating CO2 (CCM) at the catalytic site of the carboxylating enzyme RuBisCO, which enhances CO2 fixation2. Although many cellular components involved in the transport and sequestration of inorganic carbon have been identified3,4, how microalgae supply energy to concentrate CO2 against a thermodynamic gradient remains unknown4,5,6. Here we show that in the green alga Chlamydomonas reinhardtii, the combined action of cyclic electron flow and O2 photoreduction—which depend on PGRL1 and flavodiiron proteins, respectively—generate a low luminal pH that is essential for CCM function. We suggest that luminal protons are used downstream of thylakoid bestrophin-like transporters, probably for the conversion of bicarbonate to CO2. We further establish that an electron flow from chloroplast to mitochondria contributes to energizing non-thylakoid inorganic carbon transporters, probably by supplying ATP. We propose an integrated view of the network supplying energy to the CCM, and describe how algal cells distribute energy from photosynthesis to power different CCM processes. These results suggest a route for the transfer of a functional algal CCM to plants to improve crop productivity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Genes studied in this Article can be found on https://phytozome-next.jgi.doe.gov/ under the loci Cre12.g531900 (FLVA), Cre16.g691800 (FLVB), Cre07.g340200 (PGRL1), Cre16.g662600 (BST1), Cre16.g663400 (BST2) and Cre16.g663450 (BST3).
References
Field, C. B., Behrenfeld, M. J., Randerson, J. T. & Falkowski, P. Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281, 237–240 (1998).
Mackey, K. R., Morris, J. J., Morel, F. M. & Kranz, S. A. Response of photosynthesis to ocean acidification. Oceanography 28, 74–91 (2015).
Mackinder, L. C. M. et al. A spatial interactome reveals the protein organization of the algal CO2-concentrating mechanism. Cell 171, 133–147.e114 (2017).
Mackinder, L. C. M. The Chlamydomonas CO2-concentrating mechanism and its potential for engineering photosynthesis in plants. New Phytol. 217, 54–61 (2018).
Raven, J. A. Inorganic carbon acquisition by eukaryotic algae: four current questions. Photosynth. Res. 106, 123–134 (2010).
Raven, J. A., Beardall, J. & Giordano, M. Energy costs of carbon dioxide concentrating mechanisms in aquatic organisms. Photosynth. Res. 121, 111–124 (2014).
Maberly, S. C. & Gontero, B. Ecological imperatives for aquatic CO2-concentrating mechanisms. J. Exp. Bot. 68, 3797–3814 (2017).
Savir, Y., Noor, E., Milo, R. & Tlusty, T. Cross-species analysis traces adaptation of Rubisco toward optimality in a low-dimensional landscape. Proc. Natl Acad. Sci. USA 107, 3475–3480 (2010).
Reinfelder, J. R. Carbon concentrating mechanisms in eukaryotic marine phytoplankton. Annu. Rev. Mar. Sci. 3, 291–315 (2011).
Moroney, J. V. et al. The carbonic anhydrase isoforms of Chlamydomonas reinhardtii: intracellular location, expression, and physiological roles. Photosynth. Res. 109, 133–149 (2011).
Duanmu, D., Miller, A. R., Horken, K. M., Weeks, D. P. & Spalding, M. H. Knockdown of limiting-CO2-induced gene HLA3 decreases HCO3− transport and photosynthetic Ci affinity in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 106, 5990–5995 (2009).
Wang, Y. & Spalding, M. H. Acclimation to very low CO2: contribution of limiting CO2 inducible proteins, LCIB and LCIA, to inorganic carbon uptake in Chlamydomonas reinhardtii. Plant Physiol. 166, 2040–2050 (2014).
Yamano, T., Sato, E., Iguchi, H., Fukuda, Y. & Fukuzawa, H. Characterization of cooperative bicarbonate uptake into chloroplast stroma in the green alga Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 112, 7315–7320 (2015).
Mukherjee, A. et al. Thylakoid localized bestrophin-like proteins are essential for the CO2 concentrating mechanism of Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 116, 16915–16920 (2019).
Karlsson, J. et al. A novel α-type carbonic anhydrase associated with the thylakoid membrane in Chlamydomonas reinhardtii is required for growth at ambient CO2. EMBO J. 17, 1208–1216 (1998).
Raven, J. A. CO2-concentrating mechanisms: a direct role for thylakoid lumen acidification? Plant Cell Environ. 20, 147–154 (1997).
Badger, M. R., Kaplan, A. & Berry, J. A. Internal inorganic carbon pool of Chlamydomonas reinhardtii. Plant Physiol. 66, 407–413 (1980).
Allen, J. F. Photosynthesis of ATP—electrons, proton pumps, rotors, and poise. Cell 110, 273–276 (2002).
Allen, J. F. Cyclic, pseudocyclic and noncyclic photophosphorylation: new links in the chain. Trends Plant Sci. 8, 15–19 (2003).
Munekage, Y. et al. PGR5 is involved in cyclic electron flow around photosystem I and is essential for photoprotection in Arabidopsis. Cell 110, 361–371 (2002).
Johnson, X. et al. Proton gradient regulation 5-mediated cyclic electron flow under ATP- or redox-limited conditions: a study of ΔATPase pgr5 and ΔrbcL pgr5 mutants in the green alga Chlamydomonas reinhardtii. Plant Physiol. 165, 438–452 (2014).
DalCorso, G. et al. A complex containing PGRL1 and PGR5 is involved in the switch between linear and cyclic electron flow in Arabidopsis. Cell 132, 273–285 (2008).
Tolleter, D. et al. Control of hydrogen photoproduction by the proton gradient generated by cyclic electron flow in Chlamydomonas reinhardtii. Plant Cell 23, 2619–2630 (2011).
Curien, G. et al. The water to water cycles in microalgae. Plant Cell Physiol. 57, 1354–1363 (2016).
Helman, Y. et al. Genes encoding a-type flavoproteins are essential for photoreduction of O2 in cyanobacteria. Curr. Biol. 13, 230–235 (2003).
Gerotto, C. et al. Flavodiiron proteins act as safety valve for electrons in Physcomitrella patens. Proc. Natl Acad. Sci. USA 113, 12322–12327 (2016).
Shimakawa, G. et al. The Liverwort, Marchantia, drives alternative electron flow using a flavodiiron protein to protect PSI. Plant Physiol. 173, 1636–1647 (2017).
Chaux, F. et al. Flavodiiron proteins promote fast and transient O2 photoreduction in Chlamydomonas. Plant Physiol. 174, 1825–1836 (2017).
Dang, K. V. et al. Combined increases in mitochondrial cooperation and oxygen photoreduction compensate for deficiency in cyclic electron flow in Chlamydomonas reinhardtii. Plant Cell 26, 3036–3050 (2014).
Bailleul, B. et al. Energetic coupling between plastids and mitochondria drives CO2 assimilation in diatoms. Nature 524, 366 (2015).
Wang, Y., Stessman, D. J. & Spalding, M. H. The CO2 concentrating mechanism and photosynthetic carbon assimilation in limiting CO2: how Chlamydomonas works against the gradient. Plant J. 82, 429–448 (2015).
Kono, A. & Spalding, M. H. LCI1, a Chlamydomonas reinhardtii plasma membrane protein, functions in active CO2 uptake under low CO2. Plant J. 102, 1127–1141 (2020).
Bonente, G. et al. Analysis of LhcSR3, a protein essential for feedback de-excitation in the green alga Chlamydomonas reinhardtii. PLoS Biol. 9, e1000577 (2011).
Tian, L. et al. pH dependence, kinetics and light-harvesting regulation of nonphotochemical quenching in Chlamydomonas. Proc. Natl Acad. Sci. USA 116, 8320–8325 (2019).
Sültemeyer, D. F., Klug, K. & Fock, H. P. Effect of dissolved inorganic carbon on oxygen evolution and uptake by Chlamydomonas reinhardtii suspensions adapted to ambient and CO2-enriched air. Photosynth. Res. 12, 25–33 (1987).
Sültemeyer, D., Biehler, K. & Fock, H. P. Evidence for the contribution of pseudocyclic photophosphorylation to the energy requirement of the mechanism for concentrating inorganic carbon in Chlamydomonas. Planta 189, 235–242 (1993).
Lucker, B. & Kramer, D. M. Regulation of cyclic electron flow in Chlamydomonas reinhardtii under fluctuating carbon availability. Photosynthesis Res. 117, 449–459 (2013).
Qu, Z. & Hartzell, H. C. Bestrophin Cl− channels are highly permeable to HCO3−. Am. J. Physiol. Cell Physiol. 294, C1371–C1377 (2008).
Rost, B., Riebesell, U., Burkhardt, S. & Sültemeyer, D. Carbon acquisition of bloom-forming marine phytoplankton. Limnol. Oceanogr. 48, 55–67 (2003).
Basu, S. & Mackey, K. R. M. Phytoplankton as key mediators of the biological carbon pump: their responses to a changing climate. Sustainability 10, 869 (2018).
Atkinson, N. et al. Introducing an algal carbon-concentrating mechanism into higher plants: location and incorporation of key components. Plant Biotechnol. J. 14, 1302–1315 (2016).
Meyer, M. T., McCormick, A. J. & Griffiths, H. Will an algal CO2-concentrating mechanism work in higher plants? Curr. Opin. Plant Biol. 31, 181–188 (2016).
Nölke, G. et al. The integration of algal carbon concentration mechanism components into tobacco chloroplasts increases photosynthetic efficiency and biomass. Biotechnol. J. 14, 1800170 (2019).
Hennacy, J. H. & Jonikas, M. C. Prospects for engineering biophysical CO2 concentrating mechanisms into land plants to enhance yields. Annu. Rev. Plant Biol. 71, 461–485 (2020).
Yamamoto, H., Takahashi, S., Badger, M. R. & Shikanai, T. Artificial remodelling of alternative electron flow by flavodiiron proteins in Arabidopsis. Nat. Plants 2, 16012 (2016).
Wada, S. et al. Flavodiiron protein substitutes for cyclic electron flow without competing CO2 assimilation. Plant Physiol. 176, 1509–1518 (2017).
Gómez, R. et al. Faster photosynthetic induction in tobacco by expressing cyanobacterial flavodiiron proteins in chloroplasts. Photosynth. Res. 136, 129–138 (2018).
Vicino, P. et al. Expression of flavodiiron proteins Flv2–Flv4 in chloroplasts of Arabidopsis and tobacco plants provides multiple stress tolerance. Int. J. Mol. Sci. 22, 1178 (2021).
Burlacot, A., Burlacot, F., Li-Beisson, Y. & Peltier, G. Membrane inlet mass spectrometry: a powerful tool for algal research. Front. Plant. Sci. 11, 1302 (2020).
Burlacot, A. et al. Flavodiiron-mediated O2 photoreduction links H2 production with CO2 fixation during the anaerobic induction of photosynthesis. Plant Physiol. 177, 1639–1649 (2018).
Burlacot, A., Richaud, P., Gosset, A., Li-Beisson, Y. & Peltier, G. Algal photosynthesis converts nitric oxide into nitrous oxide. Proc. Natl Acad. Sci. USA 117, 2704–2709 (2020).
Desplats, C. et al. Characterization of Nda2, a plastoquinone-reducing type II NAD(P)H dehydrogenase in Chlamydomonas chloroplasts. J. Biol. Chem. 284, 4148–4157 (2009).
Yamano, T. et al. Light and low-CO2-dependent LCIB–LCIC complex localization in the chloroplast supports the carbon-concentrating mechanism in Chlamydomonas reinhardtii. Plant Cell Physiol. 51, 1453–1468 (2010).
Moroney, J. V. et al. Isolation and characterization of a mutant of Chlamydomonas reinhardtii deficient in the CO2 concentrating mechanism. Plant Physiol. 89, 897–903 (1989).
Gerster, R. An attempt to interpret the kinetics of isotope exchange between C18O2 and the water of a leaf: experiments in the dark. Planta 97, 155–172 (1971).
Silverman, D. N. In Methods in Enzymology Vol. 87 (ed. Purich, D. L.) 732–752 (Academic Press, 1982).
Cruz, J. A., Sacksteder, C. A., Kanazawa, A. & Kramer, D. M. Contribution of electric field (Δψ) to steady-state transthylakoid proton motive force (pmf) in vitro and in vivo. control of pmf parsing into Δψ and ΔpH by ionic strength. Biochem. 40, 1226–1237 (2001).
Douchi, D. et al. Membrane-inlet mass spectrometry enables a quantitative understanding of inorganic carbon uptake flux and carbon concentrating mechanisms in metabolically engineered cyanobacteria. Front. Microbiol. 10, 1356–1356 (2019).
Kramer, D. M. & Evans, J. R. The importance of energy balance in improving photosynthetic productivity. Plant Physiol. 155, 70–78 (2011).
Acknowledgements
This work was supported by the A*MIDEX (ANR-11-IDEX-0001-02) project and by the ANR OTOLHYD. A.B. acknowledges support from the Carnegie Institution for Science. O.D. is the recipient of a PhD grant awarded to Y.L-.B. We thank A. Grossman for use of the JTS-100; G. Peers for stimulating discussion; K. K. Niyogi and M. Iwai for critical reading of the manuscript; J. V. Moroney for the bsti-1 mutant; L. Mackinder for the BST3 antibody; and H. Fukuzawa for the LCIA and LCI1 antibodies. We gratefully acknowledge the contributions of S. Moulin for artistic drawings in Fig. 4, S. Blangy for LCIC antibody preparation, E. Calikanzaros and V. Epting for technical assistance, and A. Gosset for performing genetic crosses of flvB and pgrl1 mutants. The authors acknowledge the European Union Regional Developing Fund, the Region Provence Alpes Côte d’Azur, the French Ministry of Research, and the CEA for funding the HelioBiotec platform.
Author information
Authors and Affiliations
Contributions
A.B. and G.P. designed the research; A.B., O.D., P.A., S.C. and G.P. performed research; A.B. and G.P. contributed new reagents and analytic tools; A.B. and G.P. analysed data; A.B. and G.P. wrote the paper with input from Y.L.-B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Cornelia Spetea Wiklund and the other, anonymous, reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Obtaining and characterizing of pgrl1 flvB double mutants.
a, Upon crossing of pgrl1 and flvB-21 opposite mating types, progenies were screened for FLVB deficiency based on the chlorophyll fluorescence pattern, and for insertion in the PGRL1 locus by PCR. Five independent double mutants and control strains were randomly selected among the pool of double mutants and wild-types for further analysis. b, c, Chlorophyll fluorescence patterns of parental strains (pgrl1 and flvB-21) and of their respective control strains (137AH and CC-4533 respectively) in b, and of progenies of the crossing (pgrl1 flvB-1, 2, 3, 4 and 5; WT-1, 2, 3, 4) in C. Chlorophyll fluorescence measurements were performed using a PAM fluorimeter in the dark and under red actinic light (100 µmol photon m−2 s−1, from t = 22 s to t = 110 s) at a Ci concentration of 1 mM. Data are normalized to initial FM and slightly shifted on the time axis for clarity. d, PCR amplification targeting the PGRL1 locus in the different strains. Shown is representative of n = 3 independent experiments.
Extended Data Fig. 2 Maximum net O2 evolution and K1/2 values measured in pgrl1, flvB, pgrl1 flvB and their respective controls when grown under Low CO2 or High CO2.
Net O2 production rates were measured as described in Fig. 1 in low CO2 (a, b) or high CO2 (c, d) grown cells. a, c, Maximal net O2 production rates. Shown are mean values and replicates (n = 3 biologically independent sample). b, d, K1/2 values determined for each strain from hyperbolic fits. Shown are mean values ±SD of the fit of data shown in Fig. 1a–f. Letters (a–d) above bars in (b) represent significant differences (p < 0.05) between strains based on one-way ANOVA analysis.
Extended Data Fig. 3 Growth of pgrl1, flvB, pgrl1 flvB mutants and their control strains.
Cells were spotted on plates containing minimal medium at pH 7.2 (a, c, e) or pH 8.2 (b, d, f) and grown under continuous high light (100 µmol photon m−2 s−1, a, b), medium light (60 µmol photon m−2 s−1, c, d) or low light (30 µmol photon m−2 s−1, e, f). For each pH and light conditions, cells were grown either at 100 ppm CO2 (Very low CO2), 400 ppm CO2 (Low CO2) or 3% of CO2 (High CO2). Growth was assessed in pgrl1, flvB, and their respective control strains (137AH and CC-4533) (left panels) and on double mutants (pgrl1 flvB-1 to -5) and their control strains (WT-1 to -4) (right panels); the CCM1 mutant cia5 was introduced as a CCM-deficient control together with its reference strain CC-125. Shown are representative spot tests of n = 10 independent experiments.
Extended Data Fig. 4 Photosynthesis and CCM components assessment.
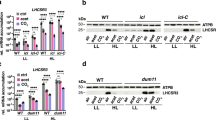
a, Maximum PSII efficiency (FV/FM) of double mutants (pgrl1 flvB-1 to -5) and their control strains (WT-1 to -4), measured using a PAM fluorimeter after 15 min dark adaptation. b, Immunodetection of the PSII subunit PsbD, the PSI subunit PsaC, the cytochrome b6f complex subunit PetB, the large Rubisco subunit RbcL, the type II NADH dehydroagenase NDA2, the mitochondrial cytochrome aa3 oxidase subunit CoxIIB, of the alternative oxidase AOX1, and of the Fe superoxide dismutase FeSOD, in two independent pgrl1 flvB double mutants and controls grown under Low CO2 or High CO2. c, Immunodetection of LCI1, BST3, RbcL and LHCSR3 in pgrl1, flvB, and their respective control strains (137AH and CC-4533). d, Immunodetection of FLVA, FLVB, PGRL1, LCI1, and BST3 in bsti-1 and its control strain (D66) at low density. e, Immunodetection of LHCSR3 in bsti-1 and its control strain (D66) cultivated at low (LD, 6 µg chlorophyll. mL−1) or high (LD, 12 µg chlorophyll. mL−1) cell density. When cultivating algae at different cell densities, which affects light and CO2 availability (two factors known to modulate LHCSR3 accumulation), we observed that bsti-1 accumulates lower LHCSR3 levels as compared to D66 at LD, but similar levels at HD. Data shown in b–e have been reproduced with n = 2 independent experiments. f, Carbonic anhydrase (CA) activity was determined in vivo by following the unlabelling of 18O-enriched CO2 in the same strains and conditions as in Fig. 2b. Shown are mean values and replicates (n = 3 biologically independent sample).
Extended Data Fig. 5 Electrochromic carotenoid absorbance changes measurements and pmf size and partitioning between ΔpH and ΔΨ.
ECS signal was measured upon light to dark transition after 2 min red light illumination (500 µmol photons m2 s−1) in the presence of 1mM NaHCO3. a, Typical ECS traces are shown for single mutants (pgrl1, flvB and bsti-1), their control strains (137AH, CC-4533 and D66 respectively), for double mutants (pgrl1flvB-1 to -5) and their controls (WT-1 to -4). Light was turned off at t = 0. b, relative pmf size was determined for each strain. c. Proportion of ΔpH in the pmf was determined for each strain. Shown are means and replicate values (n = 3 biologically independent sample). pgrl1 flvB double mutants showed a significantly lower pmf and ΔpH as compared to simple mutants and control strains (p < 0.05, one way ANOVA).
Extended Data Fig. 6 Chlorophyll fluorescence, NPQ and Ci measured during dark-light-dark transients in pgrl1, flvB, pgrl1 flvB and their respective controls.
a–f, Chlorophyll fluorescence patterns (upper panels), NPQ and Ci measurements (lower panels). Shown are representative experiments of three biological replicates. g, h, CO2-dependent NPQ was determined by subtracting the NPQ value measured after Ci injection from the NPQ value measured upon the initial Ci depletion as indicated by dotted lines (a–f). Immunodetection of LHCSR3 is shown on top of each corresponding strain. Shown are mean values and replicates (n = 3 biologically independent sample). All strains were grown as in Fig. 1 under low CO2. Red arrows indicate addition of bicarbonate. i, j, pgrl1 flvB double mutants have maintained a full qE potential. Chlorophyll fluorescence and NPQ were measured in pgrl1 flvB-1 and pgrl1 flvB-2 following the addition of acetic acid (13 mM) decreasing the pH to 5.5 and further addition of KOH (13 mM) according to the experimental protocol described in ref. 34 of the main text. Shown are representative experiments of three biological replicates.
Extended Data Fig. 7 Ci-dependent NPQ variations are independent of state transition and linked with trans-thylakoidal ΔpH.
a, b, 77K chlorophyll fluorescence spectra were measured during Ci depletion in the control strain WT3. Samples were taken at 22 µM Ci (blue arrow, a) and 0 µM Ci (orange arrow, a) and 77K chlorophyll fluorescence spectra were recorded (b). Addition of carbonyl cyanide trifluoromethoxy phenylhydrazone (FCCP, 5 µM) was used as a control to induce state 2 (pink curve). Shown are representative experiments of n = 3 biologically independent sample. a, c–g, Chlorophyll fluorescence patterns (upper panels), NPQ and Ci measurements (lower panels) in control strains, the state transition mutant stt7-9 and bsti-1 mutant during Ci depletion in the light followed either by a dark period (a, c) or subsequent addition of the uncoupler nigericin (10 µM, purple arrow) (d–g). Shown are representative experiments of n = 3 biologically independent sample. (H, I) CO2-dependent NPQ was determined as described in Extended Data Fig. 5 (black bar, c–f). Nigericin-dependent NPQ was determined from NPQ values measured before and after nigericin addition (red bars, d–f). All strains were grown as in Fig. 1 under low CO2. Red arrows indicate addition of bicarbonate.
Extended Data Fig. 8 Ci dependency of O2 exchange rates measured by using MIMS in the presence of 18O2-enriched O2 in pgrl1, flvB and their respective controls.
Strains were grown under low CO2 as in Fig. 1. a, b, O2 exchange rates in pgrl1 and its control strain 137AH. c, Difference in O2 uptake rates between 137AH and pgrl1. d, Difference in O2 uptake rates as determined in (c) normalized to the net O2 production of pgrl1 is used to determine the CEF contribution (e, f) O2 exchange rates in flvB and its control strain (CC-4533). g, Difference in O2 uptake rates between CC-4533 and flvB. h, Light-induced O2 uptake in flvB mutant. i, The difference in O2 uptake rates as determined in (g) normalized to the net O2 production in flvB is used to determine the contribution of PCEF. j, the difference in O2 uptake rates as determined in (h) normalized to the net O2 production in flvB is used to determine the contribution of CMEF. k, Effect of the mitochondrial respiration inhibitors myxothiazol and SHAM on the light-induced O2 uptake rate measured in flvB mutant at low Ci after 3 min of illumination. Shown are mean values and replicates (n = 3 biologically independent sample). l, Effect of myxothiazol and SHAM on O2 exchange rates in pgrl1 flvB-1 double mutant. Shown are representative exchange rates from 3 independent replicates. The full inhibition of photosynthetic O2 exchange observed in these conditions reflects the requirement of CO2 fixation for additional ATP supply by CEF, PCEF and CMEF.
Extended Data Fig. 9 Effect of mitochondrial respiration inhibitors on the Ci affinity of net O2 photosynthesis.
Photosynthetic net O2 production was measured as in Fig. 1 in low CO2 (a, b, c, g, h, k) or high CO2 (d, e, f, i, j, l) grown cells in the absence or presence of the two mitochondrial respiration inhibitors myxothiazol (Myxo, 2.5 µM) and salicylhydroxamic acid (SHAM, 400 µM). a–j, For each replicate, net O2 production was measured at four different concentrations of Ci and normalized to the maximum photosynthetic net O2 production. Shown are three replicates for each strain (dots) and hyperbolic fit with variability (plain lines, dotted lines). k, l, Maximum net O2 production rates. Shown are mean values and replicates (n = 3 Shown are mean values and replicates (n = 3 biologically independent sample). m, K1/2 Ci values of high CO2 grown cells in the absence or presence of myxothiazol and SHAM. Shown are mean values ±SD of the fit of data shown in Extended Data Fig. 9 d–f, i, j.
Extended Data Fig. 10 Effects of respiratory inhibitors myxothiazol and SHAM added separately or simultaneously on the Ci affinity of net O2 photosynthesis.
Photosynthetic net O2 production was measured as in Fig. 1 in WT-3 low CO2 grown cells in the absence or in the presence of respiratory inhibitors. a, In the absence of inhibitors. b, in the presence of 400 µM SHAM. c, in the presence of 2.5 µM myxothiazol. d, in the presence of both 2.5 µM myxothiazol and 400 µM SHAM. Data shown on a–d are from n = 3 biologically independent sample (dots) and hyperbolic fit with variability (plain lines, dotted lines). e, K1/2 values were determined from hyperbolic fits. Shown are mean ± SD of the fit of experimental data shown in a–d. Asterisk represent significant difference (p < 0.05, multiple t-test).
Supplementary information
Supplementary Information
This file contains Supplementary Discussion 1, 2; legends for Supplementary Figures 1–23 and Supplementary References.
Supplementary Figures
Supplementary Figures 1–23.
Rights and permissions
About this article
Cite this article
Burlacot, A., Dao, O., Auroy, P. et al. Alternative photosynthesis pathways drive the algal CO2-concentrating mechanism. Nature 605, 366–371 (2022). https://doi.org/10.1038/s41586-022-04662-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04662-9
This article is cited by
-
Crosstalk between photosynthesis and respiration in microbes
Journal of Biosciences (2024)
-
Extra O2 evolution reveals an O2-independent alternative electron sink in photosynthesis of marine diatoms
Photosynthesis Research (2024)
-
Analyzing the Impact of Physicochemical Factors on Chlorella vulgaris Growth Through Design of Experiment (DoE) for Carbon Capture System
Molecular Biotechnology (2024)
-
Enhancing photosynthetic CO2 fixation by assembling metal-organic frameworks on Chlorella pyrenoidosa
Nature Communications (2023)
-
High salt-induced PSI-supercomplex is associated with high CEF and attenuation of state transitions
Photosynthesis Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.